Abstract
Metastatic urothelial carcinoma (mUC) is a lethal cancer, with limited therapeutic options. Large-scale studies in early settings provided critical insights into the genomic and transcriptomic characteristics of non-metastatic UC. The genomic landscape of mUC remains however unclear. Using Whole Exome (WES) and mRNA sequencing (RNA-seq) performed on metastatic biopsies from 111 patients, we show that driver genomic alterations from mUC were comparable to primary UC (TCGA data). APOBEC, platin, and HRD mutational signatures are the most prevalent in mUC, identified in 56%, 14%, and 9% of mUC samples, respectively. Molecular subtyping using consensus transcriptomic classification in mUC shows enrichment in neuroendocrine subtype. Paired samples analysis reveals subtype heterogeneity and temporal evolution. We identify potential therapeutic targets in 73% of mUC patients, of which FGFR3 (26%), ERBB2 (7%), TSC1 (7%), and PIK3CA (13%) are the most common. NECTIN4 and TACSTD2 are highly expressed regardless of molecular subtypes, FGFR3 alterations and sites of metastases.
Similar content being viewed by others
Introduction
Although recent significant improvements have been made in the management of patients with metastatic urothelial cancers (mUC)1,2,3,4,5,6,7,8, mUC remain incurable and are a major cause of cancer-related deaths. Patients with mUC are managed with platinum-based chemotherapy9 and/or PD(L)1 inhibitors1,2,3,4. Recently, the therapeutic armamentarium has been enriched with antibody-drug conjugates, enfortumab vedotin6, and sacituzumab govitecan7, as well as the pan FGFR inhibitor erdafitinib8,9. Unfortunately, the prognosis remains dismal, and all patients eventually succumb to mUC.
In recent years, several large-scale whole exome sequencing (WES) and transcriptomic analyses have provided critical insights into molecular alterations and pathways that drive early-stage urothelial carcinoma, paving the way for precision-medicine care in UC10,11,12. These studies have shown that non-metastatic UC can be subgrouped into several classes with basal- and luminal-like characteristics and different clinical outcomes13. Additionally, these comprehensive analyses identified a number of significantly mutated genes and highlighted APOBEC-signature mutagenesis as a critical oncogenic process in early-stage UC. However, most analyses were performed in early settings, i.e., non-muscle invasive bladder cancer (NMIBC) and non-metastatic muscle-invasive bladder (MIBC) cancer samples, while metastatic cancers, which cause nearly all cancer deaths in this disease, have been less thoroughly investigated, mainly due to the difficulty in obtaining tumor samples in this fast-growing disease.
As cancer molecular features evolve over time, especially with drug exposure11, the genomic landscape of early cancers might not be representative of that of lethal cancers. A better understanding of mUC will be highly valuable to improve medical strategies for advanced mUC in the context of precision-medicine. There is thus an urgent clinical need to compare the genomic characteristics of primary and metastatic bladder cancers for potentially actionable genomic alterations.
In this work, we took advantage of three prospective institution-wide tumor sequencing studies14,15,16 to analyze the genomic landscape of metastases from 97 patients with mUC (both mMIBC and mUTUC) by performing both whole exome sequencing (WES) and whole-transcriptome sequencing (RNA-seq) with the following aims: 1- to decipher both the genome and transcriptome landscape of mUC, 2- to compare metastases and primary tumors (i.e., muscle-invasive bladder cancer, [MIBC] from TCGA), and 3- to explore the clinical relevance of genomic alterations in the context of druggable alterations.
Results
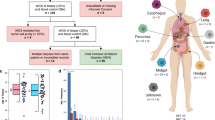
In order to study the genomic landscape of mUC, we collected demographic, clinical and treatment data from urothelial cancer patients who underwent prospective tumor DNA and RNA profiling in the context of three clinical trials between 2012 and 2020: SHIVA01 [NCT01771458]14, MOSCATO-01 [NCT02613962]15, MATCH-R [NCT02517892]16. The sample distribution over clinical characteristics and primary tumor type ([UBC] vs [UTUC]) broadly reflects the disease characteristics (Supplementary Data 1, Supplementary Data 2A, B, Fig. 1A). Overall, 111 patients with mUC were enrolled in the three studies, five patients were enrolled both MOSCATO and MATCH-R trials and thus 116 samples were obtained mainly from lymph nodes (n = 40), liver (n = 29) and lung (n = 24) (Fig. 1B). Median age at biopsy was 67 years and median time between diagnosis of metastasis and biopsy was 13 months (IQR: 7–23).
Genomic landscape of metastatic UC
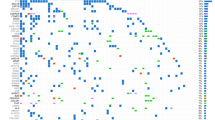
We first sought to identify the genomic alterations driving metastases of UC. WES was performed on 97 samples with matched germline DNA from 97 patients (mean coverage 105X, with 86% of target bases covered >30X). Median inferred cellularity was 0.6 (IQR = 0.44–0.78). Figure 2A describes somatic single nucleotide variants (SNVs), shorts indels and copy number alterations (CNVs) along with tumor mutational burden (TMB), mutational signatures, fusions, transcriptomic subtypes and clinical data. We identified 21,124 somatic mutations (including 20,258 SNVs and 866 indels), with median somatic mutation rate of 5.17 mutations (IQR: 4.93; 8.31) per megabase (Mb). Using a custom algorithm (see supplementary methods) to assess frequent alterations, the most frequently altered (≥10%) oncogenes were FGFR3 (27%), CCND1 (14%), PI3KCA (14%), ERBB2 (11%) and MDM2 (11%) (Fig. 2A). The most frequently altered (≥10%) tumor suppressor genes (TSG) were TP53 (38%), KMT2D (27%), CDKN2A (23%), CDKN2B (21%), ARID1A (19%), RB1 (18%) KDM6A (16%). Analysis of TERT promoter reported alterations in 77.5% of the cases. Validated fusions in mUC samples were reported in FGFR3 (n = 4 and 1 patient not included in the WES analysis also showed a FGFR3 fusion; Supplementary Data 3), ARID1A (n = 2), PIK3R1 (n = 1) and CREBBP (n = 1) genes. Additional fusion transcripts were identified involving mismatch-repair (MMR)-related partner genes (EEA1-RAD50, TAF3-GATA3 and COLQ-PPARG). Interestingly, four fusion transcripts involving SWI/SNF related partners were detected: ARID1A-ZDHHC18 (n = 2), ARID4B-POLR2D (n = 1) and CABIN1-SMARCB1 (n = 1) (Supplementary Data 3). Known cancer-susceptibility genes (e.g, MSH2, MSH6, MLH1, BRCA2) were not frequent in mUC. One sample (M2125) exhibit somatic MSH2/MSH6 mutation. Five genes were significantly mutated (q < 0.05) across the cohort using MutsigCV. All were reported to be drivers in non-metastatic UC: TP53, CDKN1A, RB1 and KMT2D. RB1 and FGFR3 were the most mutually exclusive gene pair in mUCs (exclusivity score Φ = 0.014). Pathway-based analysis (Supplementary Data 4) showed alterations in chromatin organization (59%), cell cycle (55%), genome integrity (52%), RTK-MAP (48%), PI3K-AKT (36%), transcription factors and regulators (20%), Hippo (16%) and apoptosis (13%) pathways (Fig. 2B).
A Driver gene mutations in mUC. Clinical features, immunohistochemical subgroups, and somatic genetic alterations identified in 97 mUC subjected to whole-exome sequencing. The effects of the somatic alterations are color-coded according to the legend. B Main altered pathways in mUC. Frequently altered genes were regrouped into specific signaling pathways detailed in Supplementary Table 4. Clinical features, immunohistochemical subgroups, and somatic genetic alterations identified in 97 mUC subjected to whole-exome sequencing. ICI Immune Checkpoint Inhibitor. Differential analysis between groups was performed using Fisher’s exact test, and Benjamini–Hochberg correction, two-sided. Source data are provided as a Source Data file.
A recent study suggested that UC patients harbor a high prevalence of putative deleterious germline variants that truncate tumor suppressor proteins17. We thus analyzed germline DNA in order to detect small germline variations in 66 tumor suppressor genes (TSG) of interest18. Five validated germline variants in RAD51D, FANCI, ATM, TP53 and FANCC were identified in 4 cancer predispositions genes (4% of all patients) (Supplementary Data 5).
Comparison with early UBC (eMIBC) from TCGA
To gain insight into overall genomic differences between primary and mUC, we restricted our analysis to the mMIBC subgroup (n = 68) of our metastatic cohort and compared the genomic landscape of mMIBC cohort with the TCGA dataset10 (thereafter defined as eMIBC). We reanalyzed the TCGA raw data with the same pipeline used for mUC samples and showed that our pipeline provides similar data to that of TGCA (correlation = 0.9, p = 2e−16, Wilcoxon–Mann–Whitney test) (Supplementary Fig. 2). In our metastatic cohort, the most frequent mutational signatures were APOBEC (62%), Platin (14%), HRD (6%), Deamin-5MC (5%), MMR (5%) and Aflatoxin (5%) (Supplementary Fig. 3A,B) consistent with the prior exposure to platinum-based chemotherapy. Following our quality controls (see methods), the treatment-naïve cystectomy data set included 219 samples eMIBC. The mMIBC most frequently altered (≥10%) oncogenes were FGFR3 (22%), CCND1 (13%), ERBB2 (13%), PIK3CA (12%), MDM2 (12%) and E2F3 (10%). The mMIBC most frequently altered (≥10%) TSG were TP53 (43%), KMT2D (24%), ARID1A (21%), RB1 (21%), CDKN2A (19%), CDKN2B (18%) and KDM6A (18%) (Fig. 3A right panel, Supplementary Data 6). The frequency of altered genes is similar to that reported in eUBC by the TCGA10 (Fig. 3A left panel, Supplementary Data 7). Pathway-based analysis (Supplementary Data 4) in eMIBC found alterations in chromatin organization (60%), cell cycle (55%), Genome integrity (49%), RTK-MAP (33%), PI3K-AKT (31%), transcription factors and regulators (16%) and adhesion/migration (11%) pathways (Fig. 3B). At single-gene level, FGFR3 DNA alterations were enriched in mMIBC as compared to eMIBC (22% vs 11% respectively; p = 0.043, Fisher’s exact test and Benjamini–Hochberg correction). ARID2 and BCL2L1 alteration had a tendency to be enriched in mMIBC as compared to eMIBC (7% vs 2% p = 0.060 and 6% vs 1% p = 0.057; respectively, Fisher’s exact test and Benjamini–Hochberg correction) (Fig. 3C, Supplementary Data 8). At pathways level, alterations in the following pathways were enriched in mMIBC as compared to eMIBC: RTK-MAP (47% vs 33%; p = 0.043, Fisher’s exact test and Benjamini–Hochberg correction) and apoptosis (10% vs 3%; p = 0.026, Fisher’s exact test and Benjamini–Hochberg correction) (Fig. 3B, Supplementary Data 9).
A Driver gene mutations in eMIBC (right panel; n = 219) and mMIBC (left panel; n = 68). Clinical features, immunohistochemical subgroups, and somatic genetic alterations identified in 219 eMIBC and 68 mMIBC subjected to whole-exome sequencing. The effects of the somatic alterations are color-coded according to the legend. Differential analysis between groups was performed using Fisher’s exact test, and Benjamini–Hochberg correction, two-sided. B Main altered pathways in eMIBC (right panel; n = 219) and mMIBC (left panel; n = 68). Frequently altered genes were regrouped into specific signaling pathways detailed in Sup Table 4. Clinical features, immune histochemical subgroups, and somatic genetic alterations identified 219 eMIBC and 68 mMIBC subjected to whole-exome sequencing. The effects of the somatic alterations are color-coded according to the legend. Differential analysis between groups was performed using Fisher’s exact test, and Benjamini–Hochberg correction, two-sided. C Comparison with eMIBC from the TCGA cohort revealed significantly mutated genes in mMIBC. Scatter plots depicting the mutational frequencies (percentage of patients) between the overall cohorts of mMIBC (n = 68) and eMIBC (n = 219) (FGFR3 - p = 0.043/ARID2 – p = 0.060/BCL2L1 – p = 0.057, Fisher’s exact test and Benjamini–Hochberg correction, two-sided). D Evolution of the clonal composition from the eMIBC to the matched metastases in four patients. The intersections represent clonal and subclonal variants common to both eMIBC and mMIBC. The blue circles represent variants (clonal or subclonal) detected only in eMIBC or variants that are contradictory between the two categories (i.e. subclonal in eMIBC and clonal in mMIBC). The orange circles represent variants (clonal or subclonal) detected only in mMIBC or variants that are contradictory between the two categories. ICI Immune Checkpoint Inhibitor. Source data are provided as a Source Data file.
TMB was not significantly different (median 5.45 mut/Mb in mMIBC vs median 4.95 mut/Mb in eMIBC, p = 0.32, Wilcoxon–Mann–Whitney test) (Supplementary Fig. 4).
To obtain insight into ongoing tumor evolution dynamics, we examined the clonality of all variants. Notably, most mMIBC samples exhibit clonal mutations (mean fraction of clonal mutation = 74%, IQR = 0.26; 0.89–0.63). Although the cohort contains some samples with high fraction of clonal variants, most metastatic tumor samples are heterogeneous. When we compared the data to that of eMIBC, we found that the clonal diversity as assessed by the mean percentage of subclonal mutations was significantly higher mMIBC (p = 0.002, Wilcoxon–Mann–Whitney test) indicating that mutational diversity occurs over time in UC (Supplementary Fig. 5). We studied the evolution of the clonal composition from the eMIBC to the matched metastases in four patients (Fig. 3D, Supplementary Data 10). While mutational burden was not significantly different (p = 0.24, Fisher’s exact test and Benjamini–Hochberg correction), the average mutational concordance rate was low at 24% for all mutations (ranged between 9 and 47%).
Given the genomic heterogeneity observed at an individual level, we compared mutational signatures in mMIBC vs eMIBC to further understand mutational process operative in mUC. The most frequent signatures in eMIBC were APOBEC (SBS2/SBS13), Tobacco (SBS3/SBS29) and MMR (SBS15/SBS21/SBS26/SBS44) at 72%, 9% and 8% respectively, reflective of different mutagenic mechanisms operative in MIBC. Compared to eMIBC, Platin (SBS31/SBS35) signature was significantly increased in mMIBC subgroup (14% vs 0.46%, p = 0.0001, Fisher’s exact test and Benjamini–Hochberg correction) (Supplementary Fig. 3A, B, Supplementary Data 11). Platin signature was predominant in 9 samples, of which 8 had low TMB and one moderate TMB in mMIBC. These observations probably reflect the mutagenic effects of the previous treatments received, especially platinum-based chemotherapies. Other major mutational signatures including APOBEC signatures were not different between eMIBC and mMIBC (Supplementary Data 11).
Prior studies suggested that UBC and UTUC might have different biological characteristics given the disease evolution and aggressiveness19,20. We assessed overall survival (OS) in mUC from the time of biopsy, by primary tumor site (UBC vs UTUC). We found that the 1-year OS rate for patients with mUTUC tumors was 46.1 % compared to 28.6 % for patients with mMIBC tumors (p = 0.11, Fisher’s exact test and Benjamini–Hochberg correction). To evaluate whether a genetic basis exists, we analyzed TMB and gene alterations by primary tumor site (mMIBC n = 68; mUTUC n = 28). There were few genomic differences between mUC and mUTUC. The TMB was not significantly different between mMIBC and mUTUC (median 5.445 vs 4.125 mut/Mb, p-value = 0.08). Conversely, mMIBC tumors showed a higher proportion of ARID1A alterations, although not significant (p = 0.08, Fisher’s exact test and Benjamini–Hochberg correction). (Supplementary Data 12). Visceral metastases and especially liver metastases are well-known prognosis factors in mUC. A higher proportion of tumors in patients with visceral metastases harbored alterations in RB1 gene (p = 0.02, Fisher’s exact test and Benjamini–Hochberg correction), consistent with an increased tumor cell proliferation (as verified by GSEA analysis from RNAseq analysis; data not shown) and poorer prognosis associated with visceral metastases (Supplementary Data 13).
Finally, frequently altered genes and pathways, TMB, mutational signatures, breakpoints number, Percentage of Genome Altered (PGA) and clonality were correlated with overall survival (OS). TMB, breakpoints number, Percentage of Genome Altered (PGA) and clonality were not associated with outcome. Cell cycle pathway alterations were significantly associated with a poorer OS (p = 0.03, log-rank test), which is also illustrated by the negative prognostic impact of CDKN2B alterations (p = 0.04, log-rank test) and CDKN2A (p = 0.06, log-rank test) (Supplementary Data 14)
Transcriptomic molecular subtypes of metastatic UC
Recent studies reported that non-metastatic bladder cancer was a molecularly diverse disease at a transcriptomic level with heterogeneous clinical outcomes13. An important question arose as to whether or not transcriptomic features along with microenvironment characteristics evolve over time in UC. To investigate the phenotypic landscape of mUC, we first analyzed the transcriptomes of 104 mUC samples from 98 unique patients. We first sought to evaluate the distribution of the consensus molecular subtypes13 compared to those reported earlier in non-metastatic setting by TCGA study. Overall, the proportion of each subtype was similar between eMIBC and mUC, with the exception of the Neuroendocrine-like subtype which was more frequent in the metastatic population (Figs. 4A, 8.16% vs 2.2% in TCGA eMIBC, p = 0.007, Fisher’s exact test), suggesting a higher metastatic potential. Given that metastatic biopsies may derive from different tumor clones, we then estimated the composition of metastatic samples in each of the six consensus subtypes (i.e. the proportion of each subtype in each sample) rather than focusing solely on the major component. This approach showed that some metastatic sites were preferentially associated to specific subtype composition (Fig. 4B), in particular lung metastases exhibited higher proportions of luminal papillary molecular components. Consistently, FGFR3 gene alterations were enriched in lung metastases (p = 0.028, Fisher’s exact test). We next evaluated the phenotypic evolution in 9 patients from which paired samples were obtained. Figure 4C shows the proportion of the molecular subtypes in both primary tumor and metastatic lesions. In most cases (7 out of 9), the discrepancies between the predominant subtypes of the two lesions resulted from a major stroma-rich subtype composition in metastases. To avoid potential misclassification of samples ought to non-urothelial contamination in metastases compared to paired primary tumors, we focused on the epithelial phenotypes by estimating the proportion of the five epithelial specific molecular subtypes. In six patients, the predominant epithelial molecular constituent in the primary sample remained the same in the metastatic lesion. Interestingly, in two patients (M092 and M128), the co-existence of significant basal/squamous and luminal papillary epithelial components were observed in both primary and metastatic lesions. In parallel, we also evaluated class concordance and subtype-scores with a classifier designed to work independently of stromal-content (Lund classifier https://github.com/LundBladderCancerGroup/LundTaxonomy2023Classifier). The Lund classifier demonstrated the same observation with only one clear subtype change and the reiterated demonstration of tumors combining a luminal and basal-squamous subtype (Supplementary Fig. 6).
N = 98 samples. A Proportion of molecular subtypes in eMIBC vs mUC. Side by side proportions of each subtype in the mMIBC (large bars) cohort and in the eMIBC from TCGA (narrow bars). P-value shown is Fisher’s exact test for NE-like vs others. B Subtype proximity in mMIBC depending on metastatic site. Intra-tumor proportion by subtype deconvolution from whole-transcriptomic profiles are shown grouped by metastatic site. Kruskal–Wallis (K-W) test results are shown. C Evolution of subtype composition in paired primary-metastasis. Pie chart of intra-tumor proportions are shown for the 6-subtype consensus classification and for the same classification after removing the ‘stroma-rich’ subtype. Adapted from Servier Medical Art under the licence Creative Commons Attribution 3.0 France. D Overall survival Kaplan–Meier in mUC stratified by molecular subtype. Box boundaries show the first and third quartiles, with the median line shown. Whiskers spread at 1.5 times the interquartile range. Basal/Squamous (Ba.Sq): n = 25, Luminal Non Specified (LumNS): n = 1, Luminal Papillary (LumP): n = 30, Luminal Unstable (LumU): n = 21, Neuroendocrine-like (NE.like): n = 10 and Stroma rich (Stroma.rich): n = 11. Source data are provided as a Source Data file.
Finally, unlike in primary tumors13, molecular subtypes were not associated with survival in our mUC cohort (Fig. 4D). Overall, the data suggest a high level of transcriptomic heterogeneity in metastatic with coexistence of several subtypes within the same metastatic samples.
Transcriptomic-based immune landscape of metastatic UC
To interrogate the relationship between cancer genomics and the immune landscape, we next applied different bioinformatics methods to deconvolute immune cell populations from bulk transcriptome data using immune cell–specific signatures (CIBERSORT21, XCELL22, MCPcounter23, Epic24 and microenvironment composition signatures (ESTIMATE25). From these data, we inferred overall immune infiltrate and relative immune cell populations in mUC biopsies, and observed substantial heterogeneity in overall immune infiltrate–related transcripts among tumor biopsy sites, as well as heterogeneity in inferred immune cell populations. The overall level of inferred immune infiltration was not associated with TMB (Fig. 5A), prior exposure to FGFR3 inhibitors (Fig. 5B) and immune checkpoint inhibitors (Fig. 5C). The level of inferred immune infiltration was associated with the site of biopsy with lymph-node metastasis exhibiting overall higher immune infiltration (Fig. 5D). Liver metastases have been associated with lower response to ICI. There was no statistical significance in terms of immune score between the different metastatic sites. Liver metastases have a lower immune score (mean Estimate immune score: 1.9e4) compared to lymph nodes (mean: 2.11e4, Student’s test p-value: 4.71e−4) which is expected given that the purity of the liver metastasis samples is not 100% and that lymph-node metastases are surrounded entirely by lymphocytes. We did then an exploratory analysis of 762 immune-related genes in order to discover transcripts differentially overexpressed or underexpressed in liver metastases. We hypothesized that such transcripts might have a key role to play in anticancer inflammation and, if rigorously validated, could yield not only a better understanding of these processes, but novel anticancer therapeutic strategies. Overall, 36 genes were upregulated in liver metastases after adjustment for multiple hypothesis testing (FDR = 5%), of which 31 were also found to be overexpressed in normal liver (Supplementary Data 15). Of the remaining 5 genes specifically upregulated in liver metastasis (MST1R, TGFB2, DMBT1, CDKN1A, TOLLIP), TGFB2 has a known role in immunosuppression26.
Different bioinformatics methods were applied to deconvolute immune cell populations from bulk transcriptome data using immune cell–specific signatures (CIBERSORT, XCELL, MCPcounter, Epic) and microenvironment composition signatures (ESTIMATE). N = 98 samples. Source data are provided as a Source Data file. Global immune infiltration (estimated from transcriptomic profiles using the ESTIMATE algorithm) was associated to TMB (a), prior FGFR inhibitor therapy (N = 88 naive and N = 10 treated patients) (b), prior immunotherapy (N = 80 naive and N = 18 treated patients) (c), and biopsy sampling site (N = 20 lung; N = 34 lymph nodes; N = 25 liver and N = 19 other) (d). Whiskers box plot represent interquartiles with 1.5x IQR, and outliers dots. e Subtype-specific immune enrichment in eMIBC vs mUC. The level of immune and stromal infiltration (estimated by MCPcounter in upper panel and ESTIMATE in lower panel) was compared in one subtype versus the others, or in FGFR3-mutated tumors vs wild-type, in either mUC after removing lymph-node biopsies (left panel) or in eMIBC from the TCGA (right panel). f Forest plot of univariate survival models in mUC of stromal and immune estimates, stratified by presence of visceral metastasis and lymph node biopsy site. The squares represent HR with the confidence intervals in gray lines. Two-tailed Mann–Whitney test was applied g Forest plot of univariate survival models in eMIBC of stromal and immune estimates. The squares represent HR with the confidence intervals in gray lines. Two-tailed Mann–Whitney test was applied. TMB Tumor Mutational Burden, LN lymph node, MCP Microenvironment Cell Population, HR Hazard Ratio. N = 98 mUC versus N = 408 eMIBC from TCGA.
We then assessed the association between molecular subtypes with tumor immune composition. We showed that in metastatic samples unlike in primary tumors, molecular subtypes have virtually no association with the tumor immune landscape (Fig. 5E). For example, metastatic FGFR3 mutated tumors have similar immune infiltrates than FGFR3-WT tumors. The composition of the microenvironment of metastatic tumors, in particular its enrichment in B-cells, CD8 cytotoxic T-cells and endothelial cells (Fig. 5F) is significantly predictive of an improved prognostic (after correction for biopsy site and the presence of visceral metastases at the time of biopsy) while these have no impact on the survival of patients with localized diseases (Fig. 5G).
Clinical actionability of genomic alterations
Finally, we evaluated how often genomic analysis provides potentially clinically actionable therapeutic information in mUC and compared the spectrum of targetable genomic alterations by primary tumor site. Using the OncoKB classification27, we stratified frequent genomic alterations (in the oncoprint) by highest level of clinical actionability (Fig. 6, Supplementary Data 16). OncoKB assigns levels of actionability based on evidence for the genomic alteration to serve as a biomarker either in that cancer type or in other cancer types. The only Level 1 alterations in mUC are hotspot FGFR3 mutations and FGFR2/3 fusions, which are markers for anti FGFR erdafitinib8. In our cohort of mUC, 25 (26%) patients had an FGFR2/3 alteration, 13 (13%) PIK3CA mutations, 7 (7%) ERBB2 mutations and 7 (7%) TSC1 mutations. Overall, 71 (73%) patients harbored at least one predicted actionable target event. We identified 8 patients (8%) with HRD signature and 6 patients (6%) that did harbor loss-of-function alterations in HRD genes (Fig. 2B and Supplementary Data 1). Only 1 patient was reported to have HRD signature and LOF alterations in HRD genes. Based on their HRD-phenotype, these patients might benefit from PARP-inhibitors and/or chemotherapeutics that induce double strand DNA breaks. Finally, we sought to investigate NECTIN-4 and TROP-2 expression as enfortumab vedotin and sacituzumab govitecan might be approved in the management of patients with mUC. We found that both targets showed similar expression levels in all metastatic sites, suggesting that metastatic localization should not impact treatment. TROP-2/TACSTD2 and NECTIN-4 also displayed high levels of expression in all molecular subtypes except for the tumors classified as Neuroendocrine-like which had lower expression, suggesting that these might not be suitable for such therapies (Supplementary Fig. 7A). In addition, both NECTIN 4 and TACSTD2 were expressed in FGFR3-wild-type and FGFR3 altered mUC (Supplementary Fig. 7B), in ICI-naïve and ICI-exposed mUC (Supplementary Fig. 7C) and independently from metastatic site (Supplementary Fig. 7D).
Frequently altered genes were regrouped according to druggable alterations detailed in Supplementary Data 16. Clinical features, immunohistochemical subgroups, and somatic druggable genetic alterations identified in n = 97 mUC were analyzed by whole-exome sequencing. The effects of the somatic alterations are color-coded according to the legend. Source data are provided as a Source Data file.
Discussion
Although some progress has been achieved in mUC, the prognosis for these patients remains dismal. To define and identify targets and new strategies for therapy, it is of utmost importance to better understand the biology of mUC. We reported a large cohort of fresh-frozen metastatic samples analyzed by whole exome and RNA sequencing. Our results showed that at the cohort level, the molecular landscapes at the DNA level are globally similar to that reported for primary MIBC, consistent with other pan genomic analyses28. However, we found that FGFR3 alterations were enriched in mUC (22% in mUC vs 11% in eMIBC). Of note, reanalysing a subset of the TCGA data, FGFR3 alterations were found to be 11% rather than 14% as previously reported in the literature10. This could be explained by our restrictive analysis and samples selection that are more coherent with our metastatic dataset. Recent reports suggest a similar frequency of FGFR3 alteration based on metastases samples. A recent study29 analyzed whole-genome DNA and RNA sequencing data for fresh-frozen metastatic tumor biopsies from 116 mUC patients who were scheduled for palliative systemic treatment within the context of a clinical trial (NCT01855477 and NCT02925234). 21% of patients had FGFR3 alterations (mainly mutations) which seems very similar to our findings.
Additionally, we identified the APOBEC mutational process as predominant in mUC, similar to eUC, but we found that acquired mutational signatures resulted from prior exposure to chemotherapies, especially platinum-containing regimens.
A recent study29 of WGS found two major genomic subtypes GenS1 in metastatic samples: the predominant (found in 67%) was APOBEC-driven with a large contribution from APOBEC-associated SBS2 and SBS13 signatures, and the second GenS2 (24%) predominantly comprised tumors with low APOBEC mutagenesis and was characterized by COSMIC signatures of unknown etiology. In this study, other signatures were the platinum treatment signature, the defective DNA mismatch repair signature, the microsatellite instability signature, and the reactive oxygen species signature. The same study also reported that driver genes resembled those found in eUC29. Another study of 32 chemotherapy-treated patients showed that the APOBEC mutational process was enriched in samples collected after platinum-based chemotherapy11. Such differences may reflect the timing of sample collection and differences in prior therapies.
Importantly, we analyzed several paired samples, and we showed that at an individual level, a high degree of heterogeneity exists between the primary and the metastatic samples. These results are consistent with prior studies that suggest that branched evolution may occur during disease progression11. In the context of precision medicine with the approval of FGFR3 inhibitors8,9 (i.e., erdafitinib) and the development of HER2 and PARP inhibitors30, these data raise the question of what type of samples should be tested in daily practice, as the majority of patients with metastases were found to have actionable alterations. Our data suggest that the analysis of a fresh biopsy or cell-free DNA should be recommended, at least for the Level 1 targets (e.g., FGFR3 mutation), as the concordance between cfDNA and metastasis samples was acceptable.
The important finding of our study was that molecular subtyping based on gene expression is highly heterogeneous. We found the six different subtypes previously identified in non-metastatic MIBC13, suggesting that all transcriptomic subtypes have metastatic potential. However, the NE-like subtype was enriched in mUC compared to eMIBC. This may reflect, again, the therapeutic impact of prior therapies, including chemotherapies, immunotherapy, and targeted therapies as recently reported in the literature31. Interestingly, we analyzed several paired samples and were able to find different subtypes coexisting within individual samples. For the majority of the patients, the predominant subtype remains despite the use of several lines of systemic therapies.
We also showed that in metastatic samples, unlike in primary tumors, molecular subtypes have virtually no association with the tumor immune landscape. Metastatic FGFR3 mutated samples exhibited similar immune infiltrates to FGFR3-wild-type samples, which is not consistent with prior reports in non-metastatic MIBC13. Chemotherapy has been associated with the induction of inflammation, and the apparent discrepancy might be explained by the prior exposure to chemotherapy in the patients included in our analysis. The composition of the microenvironment of metastatic tumors, particularly its enrichment in B-cells, CD8 T-cells, and endothelial cells, was associated with prognosis. B-cells and T-cell signatures have been associated with better outcomes in patients treated with ICI32,33. Another level of complexity lies in the differences between metastasis sites, as the proportion of each subtype may vary across tumor sites. A previous study applied hierarchical consensus clustering to organ-corrected paired RNA-seq data for 90 mUC samples and revealed five transcriptomic subtypes with two luminal subtypes (40% of the cohort), stroma-rich tumors (24% of samples), a basal/squamous subtype (23% of samples), and a nonspecified subtype (12% of samples). Studies with larger numbers of paired biopsies (obtained before and after treatment) would be needed to confirm these data in mUC. In a recent study, molecular subtypes of urothelial bladder cancer show significant associations with particular metastatic sites, indicating that subtype-specific molecular factors may play roles at various stages of the metastatic process34. Overall, these data suggest that patient stratification based on RNA analysis for therapy selection may be difficult, at least in the metastatic setting.
Overall, the majority of patients with mUC have actionable targets, making precision medicine relevant in urothelial cancer. In the context of antibody-drug conjugates, we found that all molecular subtypes express both NECTIN-4 and TROP-2 except the NE-like subtype. Currently, there is no need to test tumor samples for administering enfortumab and sacituzumab govitecan. This is not consistent with a previous study that reported low NECTIN-4 expression in the basal/squamous subtype35.
Limitations of the study include the heterogeneity of the study population and the lack of pathological data. However, the study provided insights into the molecular subtypes of mUC based on whole-exome and transcriptome analyses of metastatic biopsies from almost 100 mUC patients. Also, we analyzed several paired samples, which added data to the context of tumor heterogeneity in urothelial cancer. The collection of metastatic samples from larger cohorts is needed, especially in the context of international collaboration, given the difficulty to perform biopsies of metastases in urothelial cancer. The next step will be to focus at the single-cell level to better decipher tumor heterogeneity and the tumor microenvironment. We also acknowledge that the current study enrolled patients who had received multiple and different lines of therapies before sample collection, which may interfere with the understanding of the metastatic process and preclude any reliable association with outcomes. Nevertheless, our study provides a large set of genomic and transcriptomic data in a currently under-explored field in urothelial cancer.
Methods
Patient eligibility
Biopsies and data were collected in the context of three prospective IRB-approved clinical trials: SHIVA0114 [NCT01771458], MOSCATO-0115 [NCT02613962], and MATCH-R16 [NCT02517892]. All patients were included after written informed consent. The three trials were approved by the Ethics Committee of the National Institute of Pharmacy and Nutrition, and carried out in accordance with the Declaration of Helsinki, the Good Clinical Practice guidelines of the International Conference on Harmonization, and relevant French and European laws and directives.
Demographic and clinical characteristics, as well as genomic data, were collected from patients enrolled at Gustave Roussy (MOSCATO-01 and MATCH-R) and Institut Curie (SHIVA01) between 2012 and 2020. Blood was collected along with fresh tumors at the time of trial enrollment. Clinical characteristics, including demographics, prior therapies, response, and survival outcomes, were extracted from electronic medical databases.
In our study, we did not conduct analyses based on gender, sex, or ethnicity. This decision aligns with common practices in France, where such distinctions are generally not made unless there is a specific scientific question that necessitates it. French law tends to protect against making distinctions based on these categories except in cases where it is explicitly justified by the research objectives, which was not applicable in our case.
Whole exome and RNA sequencing
WES and RNA-seq were performed using newly extracted DNA and RNA. Concerning the 97 tumoral DNA and matched normal DNA samples:
i) The 85 matched samples from MOSCATO-01 and MATCH-R were sequenced in paired-end 75 bp reads on Illumina sequencers (NextSeq500 or Hiseq 2000/2500/4000) using Sureselect Human All Exon V5 (Agilent #5190-6213) or Sureselect Clinical Research Exome captures (Agilent #G9496A-012).
ii) The 12 matched samples from SHIVA01 were sequenced in paired-end 100 bp reads on NovaSeq (Illumina) instruments, using Sureselect Clinical Research Exome v2 capture (Agilent #5190-9492). The intersection of these three different captures was used in all analyses (Supplementary Fig. 1).
Throughout the text, the “n” refers to the number of samples analyzed in each section.
TCGA analysis
The TCGA-BLCA dataset, including 410 tumor and matched-normal samples in BAM file format, and associated clinical data were downloaded from the GDC data portal (https://portal.gdc.cancer.gov). BAM files were converted into FASTQ format using BEDTools and reanalyzed applying the same analysis pipeline used for the mUC cohort, with Agilent SureSelect Human All Exon 50 Mb exome capture. To maintain homogeneity between primary tumors from TCGA and our metastases samples, only samples with an average depth of coverage ≥65X (219 samples) were kept for downstream analyses.
Exome and RNA analysis
The bioinformatics pipeline and tools are described in the supplementary methods.
Statistical analysis
Differential mutation analysis between groups was performed using an oncoprint, Fisher’s exact test, and Benjamini–Hochberg correction. For statistical analysis of mutational signatures, Fisher’s exact test and Benjamini–Hochberg correction were also applied. For continuous data, the Wilcoxon–Mann–Whitney test was applied. Survival time was measured from the biopsy date to the time of the most recent follow-up (in months). Survival analyses were performed using the R package Survival (v2.44) on clinical and molecular features, employing the Kaplan–Meier method.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The original files and raw NGS data generated in this study have been deposited in the EGA database under accession code EGAD50000000550 [https://ega-archive.org/studies/EGAS50000000373]. Data on EGA is under controlled access. Sequencing data will be made available upon request through EGA, and additional clinical information can be made available upon institutional approval. Requests should be addressed to Drs Nicolas Servant (Nicolas.servant@curie.fr) and Marc Deloger (marc.deloger@gustaveroussy.fr). The estimated timeframe for access to be granted is 2 months, and the duration will be determined according to the request needs. The source data generated in this study are provided in the different Source Data files. All relevant clinical trial data used in this study are accessible in the Supplementary Data files and de-identified. Source data are provided with this paper.
Code availability
All code packages were previously published and detailed hereafter. Vegan v1.2.0 [https://doi.org/10.5281/zenodo.12167638]; WISP [https://doi.org/10.1038/s41467-019-09307-6] (https://github.com/cit-bioinfo/WISP); ImmuneDeconv R package [https://doi.org/10.1007/978-1-0716-0327-7_16]; Limma-Voom [https://doi.org/10.1186/gb-2014-15-2-r29] (https://ucdavis-bioinformatics-training.github.io/2018-June-RNA-Seq-Workshop/thursday/DE.html); pyTMB script (https://github.com/bioinfo-pf-curie/TMB); Complex Heatmap package [https://doi.org/10.18129/B9.bioc.ComplexHeatmap]; CRAN package https://doi.org/10.32614/CRAN.package.ggpubr]. The software used in this study comprise: BWA-MEM algorithm (v0.7.15); SAMtools v0.1.19, BEDtools v2.21.0; MarkDuplicates tool (v2.6.0); Picard Haplotype Caller (GATK v4.0.2.1); Annovar (2017/07/16 version); 1000Genomes (2015/08 version), ESP6500 and ExAC databases; MuTect2 (GATK v4.0.2.1); Facets (v0.5.11); R (v3.6.0) package Complex Heatmap (v2.2.0); MSIsensor2 (commit 0d6528b6ff, niu-lab); Palimpsest (v2.0.0); COSMIC database – version 3, release v89; CoMEt (commit 20423da, raphael-group); MutSigCV tool (v1.41); Ensembl’s human genome and transcriptome (GRCh38, version 91); MIBC R package (https://github.com/cit-bioinfo/consensusMIBC); GTEX (v7); Arriba (v1.2.0), FusionCatcher (v1.20); StarFusion (v1.8.1) and FusionReport (v2.0.1).
References
Bellmunt, J. et al. Pembrolizumab as Second-Line Therapy for Advanced Urothelial Carcinoma. N. Engl. J. Med. 376, 1015–1026 (2017).
Balar, A. V. et al. First-line pembrolizumab in cisplatin-ineligible patients with locally advanced and unresectable or metastatic urothelial cancer (KEYNOTE-052): a multicentre, single-arm, phase 2 study. Lancet Oncol. 18, 1483–1492 (2017).
Balar, A. V. et al. Atezolizumab as first-line treatment in cisplatin-ineligible patients with locally advanced and metastatic urothelial carcinoma: a single-arm, multicentre, phase 2 trial. Lancet 389, 67–76 (2017).
Powles, T. et al. Avelumab Maintenance Therapy for Advanced or Metastatic Urothelial Carcinoma. N. Engl. J. Med. 383, 1218–1230 (2020).
Powles, T. et al. Enfortumab Vedotin in Previously Treated Advanced Urothelial Carcinoma. N. Engl. J. Med. 384, 1125–1135 (2021).
Tagawa, S. T. et al. TROPHY-U-01: A Phase II Open-Label Study of Sacituzumab Govitecan in Patients With Metastatic Urothelial Carcinoma Progressing After Platinum-Based Chemotherapy and Checkpoint Inhibitors. J. Clin. Oncol. 39, 2474–2485 (2021).
Powles, T. et al. Electronic address: clinicalguidelines@esmo.org. Bladder cancer: ESMO Clinical Practice Guideline for diagnosis, treatment and follow-up. Ann. Oncol. 33, 244–258 (2022).
Loriot, Y. et al. THOR Cohort 1 Investigators. Erdafitinib or Chemotherapy in Advanced or Metastatic Urothelial Carcinoma. N. Engl. J. Med. 389, 1961–1971 (2023).
Siefker-Radtke, A. O. et al. Y; BLC2001 Study Group. Efficacy and safety of erdafitinib in patients with locally advanced or metastatic urothelial carcinoma: long-term follow-up of a phase 2 study. Lancet Oncol. 23, 248–258 (2022).
Robertson et al. Comprehensive Molecular Characterization of Muscle-Invasive Bladder Cancer. Cell 171, 540–556.e25 (2017).
Faltas, B. M. et al. Clonal evolution of chemotherapy-resistant urothelial carcinoma. Nat. Genet 48, 1490–1499 (2016).
Hurst, C. D. et al. Genomic Subtypes of Non-invasive Bladder Cancer with Distinct Metabolic Profile and Female Gender Bias in KDM6A Mutation Frequency. Cancer Cell 32, 701–715.e7 (2017).
Kamoun, A. et al. A Consensus Molecular Classification of Muscle-invasive Bladder Cancer. Eur. Urol. 77, 420–433 (2020).
Le Tourneau, C. et al. Molecularly targeted therapy based on tumour molecular profiling versus conventional therapy for advanced cancer (SHIVA): a multicentre, open-label, proof-of-concept, randomised, controlled phase 2 trial. Lancet Oncol. 16, 1324–1334 (2015).
Massard, C. et al. High-Throughput Genomics and Clinical Outcome in Hard-to-Treat Advanced Cancers: Results of the MOSCATO 01 Trial. Cancer Discov. 7, 586–595 (2017).
Recondo, G. et al. Feasibility and first reports of the MATCH-R repeated biopsy trial at Gustave Roussy. NPJ Precis. Oncol 4, 27 (2020).
Vosoughi, A. et al. Common germline-somatic variant interactions in advanced urothelial cancer. Nat. Commun. 11, 6195 (2020).
Huang, K. L. et al. Pathogenic Germline Variants in 10,389 Adult Cancers. Cell 173, 355–370.e14 (2018).
Moss, T. J. et al. Comprehensive Genomic Characterization of Upper Tract Urothelial Carcinoma. Eur. Urol. 72, 641–649 (2017).
Winters, B. R. et al. Genomic distinctions between metastatic lower and upper tract urothelial carcinoma revealed through rapid autopsy. JCI Insight 5, e128728 (2019).
Chen, B., Khodadoust, M. S., Liu, C. L., Newman, A. M. & Alizadeh, A. A. Profiling Tumor Infiltrating Immune Cells with CIBERSORT. Methods Mol. Biol. 1711, 243–259 (2018).
Aran, D., Hu, Z. & Butte, A. J. xCell: digitally portraying the tissue cellular heterogeneity landscape. Genome Biol. 18, 220 (2017).
Becht, E. et al. Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol. 17, 218 (2016).
Racle, J. & Gfeller, D. EPIC: A Tool to Estimate the Proportions of Different Cell Types from Bulk Gene Expression Data. Methods Mol. Biol. 2120, 233–248 (2020).
Liu, W. et al. Transcriptome-derived stromal and immune scores infer clinical outcomes of patients with cancer. Oncol. Lett. 15, 4351–4357 (2018).
Khanolkar, R. C. et al. TGFβ2 Induces the Soluble Isoform of CTLA-4 - Implications for CTLA-4 Based Checkpoint Inhibitor Antibodies in Malignant Melanoma. Front. Immunol. 12, 763877 (2022).
Chakravarty, D., Gao, J. & Phillips, S. M. OncoKB: a precision oncology knowledge base. JCO Precis. Oncol. 1, 1–16 (2017).
Nguyen, B. et al. Genomic characterization of metastatic patterns from prospective clinical sequencing of 25,000 patients. Cell 185, 563–575.e11 (2022).
Nakauma-González, J. A. et al. Comprehensive Molecular Characterization Reveals Genomic and Transcriptomic Subtypes of Metastatic Urothelial Carcinoma. Eur. Urol. 81, 331–336 (2022).
Grivas, P. et al. Efficacy and safety of rucaparib in previously treated, locally advanced or metastatic urothelial carcinoma from a phase 2, open-label trial (ATLAS). BMC Cancer 21, 593 (2021).
Erlmeier, F. et al. Spatial Immunephenotypes of Distant Metastases but not Matched Primary Urothelial Carcinomas Predict Response to Immune Checkpoint Inhibition. Eur. Urol. 83, 133–142 (2023).
Fridman, W. H. et al. B cells and tertiary lymphoid structures as determinants of tumour immune contexture and clinical outcome. Nat. Rev. Clin. Oncol. 19, 441–457 (2022).
Vanhersecke, L. et al. Mature tertiary lymphoid structures predict immune checkpoint inhibitor efficacy in solid tumors independently of PD-L1 expression. Nat. Cancer 2, 794–802 (2021).
Sjödahl, G. et al. Metastasis and recurrence patterns in the molecular subtypes of urothelial bladder cancer. Int J. Cancer 154, 180–190 (2024).
Chu, C. E. et al. Heterogeneity in NECTIN4 Expression Across Molecular Subtypes of Urothelial Cancer Mediates Sensitivity to Enfortumab Vedotin. Clin. Cancer Res. 27, 5123–5130 (2021).
Acknowledgements
The authors acknowledge the global clinical team for its involvement in the enrollment of patients in MOSCATO trial, as well as all relevant personnel from Gustave Roussy Direction de la Recherche who made possible this academic-sponsored trial (Aurélie Abou-Lovergne, Lisa Lambert, Thibaud Motreff, and Delphine Vuillier). Maud Ngo-Camus, Aljosa Celebic, and Katty Malekzadeh are thanked for Clinical Data Management/Data Management. Dorota Gajda, Dienabou Sylla, and Adrien Allorant are thanked for their statistical assistance. The authors acknowledge laboratory and bioinformatic teams, including Amélie Boichard, Mélanie Laporte, Isabelle Miran, Nelly Motté, Ludovic Bigot, Stéphanie Coulon, Marie Breckler, Catherine Richon, Aurélie Honoré, Magali Kernaleguen, Glawdys Faucher, Zsofia Balogh, Jonathan Sabio, Lionel Fougeat, Marie Xiberras, Leslie Girard, Lucie Herard, Catherine Lapage, Guillaume Meurice, Romy Chen-Min-Tao, Yannick Boursin, Marc Deloger, Celine Lefebvre, and Marion Pedrero, for their support in the preanalytic processing of samples, storage, and for tumor sample analysis (wet lab and bioinformatic) of the patients included in the MOSCATO 01 trial. The authors especially acknowledge Yuki Takahashi for her editing support. We thank the patients who volunteered to participate in the SHIVA01 study for their dedication and the study-site staff who cared for them. We thank the Agence Nationale de le Recherche (Investissements d’avenir) and Site de Recherche Intégré contre le Cancer (SiRIC). High-throughput sequencing was done by the NGS platform of the Institut Curie, supported by grants ANR-10-EQPX-03 and ANR-10-INBS-09-08 from the Agence Nationale de la Recherche (investissements d’avenir) and the Canceropôle Ile-de-France. The results shown here are in whole or part based upon data generated by the TCGA Research Network: https://www.cancer.gov/tcga.
Author information
Authors and Affiliations
Contributions
Conceptualization: Y.L., M.K., C.L.T., C.M., I.B. Data curation: Y.L., M.K., L.S., R.N., C.D., N.M. Formal Analysis: Y.L., M.K., L.S., R.N., C.D., N.M., N.S. Methodology: Y.L., M.K. L.S., R.N., C.D., N.S., C.L.T., C.M., I.B. Project administration: Y.L., M.K., C.D., C.L.T., I.B. Resources: Y.L., M.K., L.S., R.N., C.D., N.M., I.D., P.L., C.N., M.N., L.L., L.T., G.C., L.F., Z.C.A., Y.N., E.B., P.B., G.M., T.G., J.W., F.R., S.D., J.Y.S., N.S., Y.A., B.B., F.A., C.L.T., C.M., I.B. Supervision: Y.L., M.K., N.S., C.L.T., C.M., I.B. Validation: Y.L., M.K., N.S., F.A., C.L.T., C.M., I.B. Writing – original draft: Y.L., M.K., L.S., R.N., C.D. Writing – review & editing: all authors. L.S., R.N. and C.D. contributed equally. C.L.T., C.M. and I.B. contributed equally. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
C.L.T.: Roche, Seattle Genetics, Rakuten, Nanobiotix, MSD, BMS, Merck Serono, AstraZeneca, GlaxoSmithKline, Novartis, Celgene, Exscientia, ALX Oncology, Seattle Genetics. M.K.: Roche, AstraZeneca. Pernelle Lavaud: Payment or honoraria for lectures, presentations, speakers bureaus, manuscript writing or educational events from BMS, Astra Zeneca, Sanofi, Astellas; Support for attending meetings and/or travail from Daichi, Ipsen, Astellas, Sanofi, Jannssen, Pfizer; Advisory board from Daichi. L.L.: Hospitalities, Travel, advice board, research grant last 5 years with: Abbott; Adept Field Solutions; Amgen; AstraZeneca; Beckman Coulter; Bayer Boeringer; BMS; Icomed; Illumina, Genomic Health; Guardant health; Lilly; Medimmune; Myriad; Novartis; Pfizer; QualWorld1; Roche; Siemens Healthineer; Taiho Oncology, Thermofisher Sc; VelaDx. E.R.: Travel, advice board, research grant: AstraZeneca, Roche, BMS, GSK, and Clovis. L.F.: Research funding from Debiopharm, Incyte, Relay Therapeutics and Nuvalent and Non financial support from Illumina Inc and Guardant Health. F.F.: no disclosures. Benjamin Besse: grants from 4D Pharma, AbbVie, Amgen, AstraZeneca, BeiGene, Blueprint Medicines, Celgene, Cergentis, Chugai Pharmaceutical, Da Voltera, Daiichi Sankyo, Eli Lilly, Ellipse Pharma, Eisai, F-Star, Genmab, Genzyme Corporation, GSK, Hedera Dx, Inivata, Ipsen, Janssen, MSD, Onxeo, OSE Immunotherapeutics, Pfizer, Pharmamar, Roche/Genentech, Sanofi, Socar Research, Taiho Oncology, Takeda, Tolero Pharmaceuticals, and Turning Point Therapeutics during the conduct of the study. C.M.: Consultant/Advisory fees from Amgen, Astellas, Astra Zeneca, Bayer, BeiGene, BMS, Celgene, Debiopharm, Genentech, Ipsen, Janssen, Lilly, MedImmune, MSD, Novartis, Pfizer, Roche, Sanofi, Orion. Principal/sub-Investigator of Clinical Trials for Abbvie, Aduro, Agios, Amgen, Argen-x, Astex, AstraZeneca, Aveo pharmaceuticals, Bayer, Beigene, Blueprint, BMS, Boeringer Ingelheim, Celgene, Chugai, Clovis, Daiichi Sankyo, Debiopharm, Eisai, Eos, Exelixis, Forma, Gamamabs, Genentech, Gortec, GSK, H3 biomedecine, Incyte, Innate Pharma, Janssen, Kura Oncology, Kyowa, Lilly, Loxo, Lysarc, Lytix Biopharma, Medimmune, Menarini, Merus, MSD, Nanobiotix, Nektar Therapeutics, Novartis, Octimet, Oncoethix, Oncopeptides AB, Orion, Pfizer, Pharmamar, Pierre Fabre, Roche, Sanofi, Servier, Sierra Oncology, Taiho, Takeda, Tesaro, Xencor. F.A.: grants from Roche, Lilly, Novartis, Pfizer, Daiichi Sankyo, and AstraZeneca outside the submitted work. Y.L.: personal fees, and non-financial support from Sanofi, Janssen, Astellas, Seattle Genetics, Gilead, AstraZeneca, BMS, MSD, Pfizer, Merck KGaA, Pfizer, and Tahio; grants, grants from Celsius, Sanofi, Roche, MSD. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Roland Seiler, Gottfrid Sjödahl and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Loriot, Y., Kamal, M., Syx, L. et al. The genomic and transcriptomic landscape of metastastic urothelial cancer. Nat Commun 15, 8603 (2024). https://doi.org/10.1038/s41467-024-52915-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-52915-0
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.