Abstract
The lower respiratory tract (LRT) microbiome impacts human health, especially among critically ill patients. However, comprehensive characterizations of the LRT microbiome remain challenging due to low microbial mass and host contamination. We develop a chelex100-based low-biomass microbial-enrichment method (CMEM) that enables deep metagenomic profiling of LRT samples to recover near-complete microbial genomes. We apply the method to 453 longitudinal LRT samples from 157 intensive care unit (ICU) patients in three geographically distant hospitals. We recover 120 high-quality metagenome-assembled genomes (MAGs) and associated plasmids without culturing. We detect divergent longitudinal microbiome dynamics and hospital-specific dominant opportunistic pathogens and resistomes in pneumonia patients. Diagnosed pneumonia and the ICU stay duration were associated with the abundance of specific antibiotic-resistance genes (ARGs). Moreover, CMEM can serve as a robust tool for genome-resolved analyses. MAG-based analyses reveal strain-specific resistome and virulome among opportunistic pathogen strains. Evolutionary analyses discover increased mobilome in prevailing opportunistic pathogens, highly conserved plasmids, and new recombination hotspots associated with conjugative elements and prophages. Integrative analysis with epidemiological data reveals frequent putative inter-patient strain transmissions in ICUs. In summary, we present a genome-resolved functional, transmission, and evolutionary landscape of the LRT microbiota in critically ill patients.
Similar content being viewed by others
Introduction
Lower respiratory tract infections (LRTIs) are a major global health concern, causing at least three million deaths in 2019 as the world’s deadliest communicable disease1,2,3. Patients in ICUs face 3–10 times increased susceptibility to hospital-acquired LRTIs, substantially increasing mortality rates4,5. Significant differences in the LRT microbiota have been demonstrated between patients and healthy individuals, suggesting that the LRT microbiome is essential in maintaining immune homeostasis as a novel therapeutic target6,7,8,9. However, our understanding of the LRT microbiome is limited, especially in critically ill patients. Few studies examined the longitudinal LRT microbiome dynamics, the interactions between the LRT microbiome and host factors, and the evolutionary adaptations of pathogens in clinical settings10. Experimental challenges included contamination from oral commensals11 and host DNA12, low microbial biomass13, and longitudinal LRT sample collection14.
Consequently, most LRT microbiome studies used targeted amplicon sequencing (e.g., 16S rRNA), with many limitations, such as PCR bias, difficulties in pan-domain species detection, and functional and evolutionary analyses15,16,17. Collecting LRT samples longitudinally is also challenging. The sputum samples are contaminated by oral commensals18, and the bronchoalveolar lavage fluids (BALFs) are too invasive to be frequently collected19.
Researchers have traditionally relied on culture-based methods for diagnosis, whole-genome information, and functional characterization of specific LRT microbial species. However, culture-based approaches require the knowledge of species-specific media and extensive efforts, leading to few species being thoroughly investigated. In contrast, with sufficient sequencing depth, metagenome-based approaches can generate high-quality MAGs that approximate the genomic information of diverse species20,21,22. However, the MAG-based studies of the LRT microbiota were limited due to the aforementioned difficulties.
We present a multi-center longitudinal study of the LRT microbiome in 157 patients (Supplementary Data 1), totaling 453 samples. Utilizing the developed method, we acquired sufficient amounts of microbial DNA for deep metagenomic next-generation sequencing (NGS). With CMEM, we achieved comprehensive species-level profiling of the microbial species in the LRT of critically ill patients. We found hospital-specific and diverging longitudinal patterns of microbiome among pneumonia patients. We reconstructed hundreds of high-quality MAGs without the need for labor-intensive culturing. Further analyses of MAGs revealed the genome-resolved functional, evolutionary, and transmission landscapes in the ICU LRT microbiome.
Results
The development of CMEM enabled the deep-sequencing of LRT microbiomes
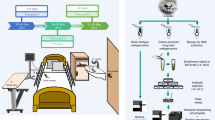
We developed CMEM that improves microbial DNA yield after host DNA removal (Methods; Fig. 1a). This substantially improved detectable microbial DNA recovery (as measured by Qubit) and DNA yield (Supplementary Fig. 1a, b). Briefly, we employed a modified saponin-based differential lysis method to remove the human nucleic acids, optimizing with a reduced saponin concentration to minimize the potential loss of microbial communities23,24. To further address the low microbial biomass challenges in LRT samples, we incorporated ultrasonic beating to improve microbial cell lysis. Next, Chelex100, commonly used in forensic investigations, was leveraged for extracting very low amounts of microbial DNA, ultimately producing high-quality DNA suitable for in-depth metagenomic sequencing (Fig. 1a).
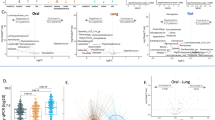
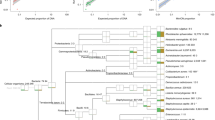
a General schematic representation of the study design and experimental pipeline. The workflow included sample liquefying, human DNA removal, ultrasonic beating to improve microbial cell lysis, DNA extraction using Chelex 100, DNA and protein precipitation, and the final steps of library preparation and sequencing. See the methods for detailed information. The plot was created with BioRender.com (citation: https://biorender.com/j76b222). b Venn diagram showing the total number of species detected in each hospital. c Percentage of detected samples for the top 10 prevalent microbial species present in the LRT microbiome from Hospital A (red), Hospital B (yellow), and Hospital C (blue), respectively. d Phylogenetic tree of 289 annotated metagenome-assembled genomes (MAGs). Bacterial species names are colored according to their phylum: The stars, circles, and squares indicate the collection sites. The bars in the outermost circle indicate the fraction of the query genome that aligns with the reference genome (up to 100%). Prevalent species with more than 3 MAGs are highlighted by colored arcs and labels. e Bar plot indicating the demographic and clinical factors significantly associated with the variation of the LRT microbiome profiles. Source data are provided as a Source Data file. Abbreviation: ARG, antibiotic resistance gene.
Head-to-head comparisons with Qiagen Power Water and Qiagen Allprep DNA/RNA kits demonstrated that CMEM produced significantly higher final DNA yield and markedly increased detectable DNA recovery (as measured by Qubit, Supplementary Fig. 1a, b). Due to CMEM’s high efficiency in DNA recovery, we were able to use an average of 5 amplification cycles for constructing sequencing libraries for LRT samples, substantially minimizing the biases introduced by PCR amplifications when dealing with low-biomass samples in previous studies. Additionally, the CMEM is implemented in a streamlined single-tube protocol to minimize DNA loss and the risk of introducing potential contamination during transfers.
To assess the impact of the host-depletion step on the LRT microbial community, we collected 32 endotracheal aspirate (ETA) samples and divided each sample into two aliquots: one assigned to the treatment group (processed with CMEM) and the other control group (processed with the same steps but without the host depletion). The host-depletion introduced noticeable variances in the relative abundance of individual taxa, and we observed no significant difference in the overall beta diversity between the two groups (PERMANOVA, p > 0.05 in Supplementary Fig. 1c, d and Supplementary Data 2 and 3). We further compared the microbiome alpha diversity and total abundance of ARGs and found no significant difference between the two groups (Supplementary Fig. 1e, f).
To further validate our approach, we observed a 92.2% consistency between microbial species detected via clinical culture and CMEM for samples with available clinical microbiology data (Supplementary Data 4). These data suggest that our approach can be a robust tool for characterizing the LRT microbiota in clinical samples with low microbial biomass and significant human nucleic acid contamination.
Characterization of LRT microbiota in ICU patients
The understanding of LRT microbiomes has been limited, especially in ICUs. We collected 453 LRT samples from 157 intubated patients from three hospitals in different cities in China (Fig. 1a and Supplementary Data 1). We successfully sequenced 442 LRT samples with the CMEM method, yielding an average of 7.09 × 107 high-quality reads per sample. An average of 14.8% microbial reads were obtained in the sequencing data (median 12.1%, interquartile range 9.7%–16.4%; Supplementary Fig. 1g). To minimize the influence of contamination from the ICU environments and medical equipment, we also collected ten environmental sampling controls from each hospital, and the identified species in these controls were highly consistent and excluded from further analysis (Supplementary Data 5 and 6). Consequently, CMEM facilitated a comprehensive species-level characterization of the microbiota in the LRT of critically ill patients. Two hundred and four microbial species were identified, including 196 bacteria, 2 fungi, and 6 viruses (Fig. 1b and Supplementary Data 7). Twenty-five opportunistic pathogen species were observed across hospitals, encompassing well-known opportunistic pathogens such as Klebsiella pneumoniae, Acinetobacter baumannii, and Pseudomonas aeruginosa. Additionally, we uncovered several less well-known opportunistic pathogens, such as Slackia exigua, Delftia acidovorans, and Aggregatibacter segnis (Supplementary Data 7). Our analysis indicated that Corynebacterium striatum, Acinetobacter baumannii, and Klebsiella pneumoniae were highly prevalent across hospitals (Fig. 1c and Supplementary Fig. 1h).
Consistent with previous findings25,26, our results revealed that the microbiome alpha diversity in samples from pneumonia patients was significantly lower than that of those without pneumonia (Wilcoxon, p = 0.00012, Supplementary Fig. 1i, Methods). Furthermore, we noted a significantly higher total abundance of ARGs in samples from pneumonia patients (Wilcoxon, p = 0.0011, Supplementary Fig. 1j). Collectively, these findings confirmed the ability of CMEM to characterize ecological diversity and ARG abundance in LRT samples effectively.
De-novo assembly of microbial reads and binning at the individual level resulted in 433 metagenome-assembled genomes (MAGs), of which 120 MAGs were high-quality based on the MIMAG (Minimum Information about a Metagenome-Assembled Genome, completeness >90% and contamination < 5%) standards27 (Methods; Fig. 1d). Among these genomes, 108 MAGs were not annotated to any references, 289 MAGs were annotated to the species level, and 36 MAGs failed to meet the Genome Taxonomy Database (GTDB) quality standards. The identified MAGs were parsed into four major phyla, including Proteobacteria, Actinobacteria, Firmicutes, and Bacteroidetes. We identified 45 species with more than one MAG, of which the most common species were Corynebacterium striatum (12.5%, 36 MAGs), Acinetobacter baumannii (11.1%, 32 MAGs), and Klebsiella pneumoniae (5.5%, 16 MAGs) (Fig. 1d and Supplementary Data 8). Crucially, the quality of the reconstructed MAGs was comparable to that of sequenced isolates, exhibiting an average nucleotide identity of 99.945% (Supplementary Data 9). This high level of concordance further validates the reliability of these MAGs recovered directly from ETA samples. The diversity of MAGs and the substantial number of unannotated genomes underscore the urgent need to characterize the LRT microbial communities with microbial-enrichment deep metagenomic approaches.
LRT microbiomes of pneumonia patients show significant site-specific signatures
We performed variation decomposition analysis to identify factors contributing to the dynamics of microbiome profiles (microbiota, resistome, and functional profiles; Methods). Among all factors, the sampling site was identified as the most important factor contributing to the variations in microbiome profiles (Fig. 1e), with the number of antibiotic classes administered and duration of ICU stay also playing important roles (Fig. 1e). Moreover, microbiome profiles showed greater variation between ICUs than within the same ICU (Supplementary Fig. 1k), indicating that ICU-specific microbial factors influenced patient microbiome profiles.
We further analyzed microbial profiles in patients with diagnosed pneumonia across three hospitals at the species level. Principal coordinate analysis (PCoA) revealed site-specific heterogeneity (Adonis p = 0.001; Fig. 2a, b). Specifically, we observed a significantly higher abundance of Klebsiella pneumoniae in Hospital A; Hospital B had a higher abundance of Corynebacterium striatum; Acinetobacter baumannii was overrepresented in Hospital C (Fig. 2b and Supplementary Fig. 2a). Variations in significantly enriched microbial pathways among hospitals existed (Methods; Supplementary Fig. 2b), including the fatty acid and lipid biosynthesis and degradation pathways, highlighting the diverse metabolic capacities upon distinct environmental pressures among hospitals.
a Principal coordinate analysis (PCoA) demonstrating hospital-specific variation in microbial profiles. Individual profiles for pneumonia patients in each hospital are illustrated in red (Hospital A; n = 33), green (Hospital B; n = 182), and blue (Hospital C; n = 124), respectively. Arrows represent contributions of the four most abundant species (arrow length is proportional to mean abundance). The F-ratio and associated p-value calculated with Adonis analysis are shown (p= 0.001, p-value was calculated by one-way PERMANOVA with 999 permutations). b PCoA highlighting the relative abundance of four species in (a). c, d Panels represent an example of temporal patterns in patients with one consistent dominant species and multiple dominant species during intubation. e The circos plot showing the mean relative abundance of the top 10 ARG types in pneumonia patients in each hospital. The colored arcs in the outermost circle indicate the percentage of mean ARG type (left half) or the percentage of each hospital contributed to each ARG type (right half). f Panels showing the total ARG abundance of the top 3 ARG types (multidrug, beta−lactam, and aminoglycoside) in samples (n = 342) collected from pneumonia patients in each hospital. Significance was determined using the two-sided Wilcoxon rank sum test. The exact p-values are directly annotated on the plot. g, h Box plots show the abundance of (g) tetA and (h) mexB genes in samples collected from patients without pneumonia (n = 66) and with pneumonia diagnosis (n = 356). Significance was determined using the two-sided Wilcoxon rank sum test. The exact p-values are directly annotated on the plot. i, j Box plots showing the abundance of (i) mexE and (j) smeB genes with days in the ICU (n = 297). Time points are indicated in the brackets. Significance was determined using the two-sided Wilcoxon rank sum test. The exact p-values are directly annotated on the plot. Figures (f–j) include a center line (median), box limits (upper and lower quartiles), and whiskers (1.5x interquartile range). Source data are provided as a Source Data file. Abbreviation: MDS multidimensional scaling, ICU intensive care unit, MLS Macrolide-Lincosamide-Streptogramin.
Temporal dynamics of LRT microbiomes
Next, we explored the temporal dynamics of the LRT microbiome. For patients with over one week of ICU stay and diagnosed pneumonia, we found that the dominant species could remain consistent during intubation (n = 12), in line with previous studies8,9, or be replaced by other dominant species over time (n = 31). For example, the dominant taxon remained stable for P68, with Pseudomonas aeruginosa persisting as the most abundant species throughout nearly a one-month ICU stay. However, for P23, the dominant taxon changed from Acinetobacter baumannii to Prevotella bivia after approximately two weeks (Fig. 2c, d). We further compared the oxygenation index (OI, Methods) and the percentage of neutrophils between patients with a consistent dominant species and dynamically changing dominant species over time. Our findings showed that patients with changing dominant species exhibited a significantly lower OI (p = 0.016) and a significantly higher percentage of neutrophils (p = 0.00014) compared to those with a single stable dominant species (Supplementary Fig. 2c). This highlights the possibility that dynamically changing dominant species could be associated with more severe pulmonary dysfunction and stronger inflammatory responses for pneumonia patients in the ICU.
Associations between clinical variables and the LRT resistomes
The high abundance of ARGs in opportunistic pathogens significantly contributes to the complexity and severity of LRTIs28,29. Our deep sequencing data extensively characterized the resistome profiles in patients with diagnosed pneumonia. A total of 20 ARG types and 716 ARG subtypes were observed in pneumonia patients across three hospitals (Supplementary Data 10). The most abundant ARG types included Multidrug, Beta-lactam, and Aminoglycoside, followed by Macrolide-Lincosamide-Streptogramin (MLS), Tetracycline, and Sulfonamide (Fig. 2e). Furthermore, specific ARGs were associated with the most prevalent ARG types in each hospital. For example, within the Beta-lactam ARGs, cfxa2, oxa-225, and shv-39 were identified as the most predominant in each hospital, respectively (Supplementary Fig. 3a). We next evaluated the resistome variation among hospitals. Except for Macrolide-Lincosamide-Streptogramin (MLS), the most predominant ARG types were significantly more abundant in Hospital C than in Hospital B (Fig. 2f and Supplementary Fig. 3b). We applied nonmetric multidimensional scaling (NMDS) to normalized ARG abundance, again showing significant site-specific differences of resistomes (Supplementary Fig. 3c; Adonis p = 0.001 and NMDS2; Methods). Key resistance-contributing genes were identified with sparse partial-least-squares discriminant analysis (sPLS-DA; Supplementary Fig. 3d, e).
Using the multivariate linear mixed models, we observed 67 significant associations between 58 ARG subtypes and 3 clinical variables (FDR < 0.05; Supplementary Fig. 3f). In addition to the observed higher total abundance of ARGs in pneumonia patient samples (Supplementary Fig. 1j), the abundances of Tetracycline genes such as tetA, tetB, Multidrug genes (mexB), as well as aminoglycoside gene (aph(3’)-IIb), were significantly higher in pneumonia patient samples (Fig. 2g, h and Supplementary Fig. 3g). The increased abundance of a variety of ARGs in pneumonia patient samples underscores the broad-spectrum resistance in pneumonia-associated pathogens in ICU settings. Notably, the abundances of multidrug genes such as mexE, smeB, and smeC, as well as the aminoglycoside gene aph(3’)-IIb, were significantly higher in patients with over 28 days of stay (Fig. 2i,j and Supplementary Fig. 3h). The significant association between the abundance of multidrug-resistant genes and the duration of ICU stay highlights that colonization by multidrug-resistant opportunistic pathogen species may be associated with prolonged ICU stay.
The strain-resolved dynamic resistome and virulome
Strain-level antibiotic resistance and virulence are critical in studying pathogens. Conventionally, this type of analysis would require culture-dependent genome sequencing approaches. We dissected 91 high-quality MAGs of five prominent opportunistic pathogen species, each MAG derived from an individual patient. The number of ARGs observed in Acinetobacter baumannii, Klebsiella pneumoniae, and Pseudomonas aeruginosa was drastically higher than in Corynebacterium striatum and Stenotrophomonas maltophilia (Fig. 3a), which may lead to the enhanced capacity to withstand antibiotic treatments. Intriguingly, the ARG profiles of the five species showed little overlap (Fig. 3b).
a Box plot showing the number of ARGs identified in the five predominant species (n = 91). The number of ARGs in each species was normalized by its genome length. Significance was determined using the two-sided Wilcoxon rank sum test. The exact p-values are annotated on the plot. The plot includes a center line (median), box limits (upper and lower quartiles), and whiskers (1.5x interquartile range). Individual data points are shown using overlaid dot plots. b Heatmap depicting the distribution of antibiotic resistance genes across strains for the five predominant species. c Panels showing the variations of ARG content among strains in Acinetobacter baumannii, Corynebacterium striatum, and Klebsiella pneumoniae. The F-ratio and associated p-value calculated with Adonis analysis are shown (p = 0.001, p-value was calculated by one-way PERMANOVA with 999 permutations). Ellipses represent 90% confidence regions. d Box plot showing the number of virulence genes identified in the five predominant species (n = 91). The number of virulence genes in each species was normalized by its genome length. Significance was determined using the two-sided Wilcoxon rank sum test. The exact p-values are annotated on the plot. The plot includes a center line (median), box limits (upper and lower quartiles), and whiskers (1.5x interquartile range). Individual data points are shown using overlaid dot plots. e Heatmap depicting the distribution of virulence genes across strains for the five predominant species. f Panels showing the variations of virulence gene content among strains in Acinetobacter baumannii, Klebsiella pneumoniae, and Pseudomonas aeruginosa. The F-ratio and associated p-value calculated with Adonis analysis are shown (p = 0.001, p-value was calculated by one-way PERMANOVA with 999 permutations). Ellipses represent 90% confidence regions. Source data are provided as a Source Data file. Abbreviation: PCoA, principal coordinate analysis.
Notably, we observed intra-species significant variations in resistome, especially in Acinetobacter baumannii, Corynebacterium striatum, and Klebsiella pneumoniae (Fig. 3c). At the higher pathway level, Corynebacterium striatum strains displayed diverging resistance, with several strains conferring resistance against tetracycline and aminoglycoside antibiotics, while others don’t (Supplementary Fig. 4a). Some Stenotrophomonas maltophilia strains exhibited strong resistance against tetracycline and fluoroquinolone antibiotics, while others demonstrated resistance against aminoglycoside antibiotics (Supplementary Fig. 4a).
Next, we profiled the virulence genes (VFs) of these strains. Consistent with previous findings30,31, Pseudomonas aeruginosa harbored an overwhelming number of VFs (Fig. 3d). Conversely, a significantly smaller number of VFs were identified in Corynebacterium striatum and Stenotrophomonas maltophilia (Fig. 3d). Overall, the number of ARGs and VFs showed strong positive correlations, indicating that highly antibiotic-resistant strains are more virulent (Supplementary Fig. 4b). Interestingly, the virulome also showed little overlap (Fig. 3e). The distribution of VFs among strains in Acinetobacter baumannii, Klebsiella pneumoniae, and Pseudomonas aeruginosa also exhibited strain-specific signatures (Fig. 3f).
Importantly, we found no substantial hospital-specific impact on the strain-level resistome, virulome, and average nucleotide identity (ANI) (Supplementary Fig. 4c-e). This suggests that these strains are widely distributed across distinct hospital environments. Our genome-resolved analysis highlights the feasibility of directly profiling resistome and virulome without culturing isolates.
Increased mobilome and highly conserved plasmids in prevalent opportunistic pathogen species
ARGs and VFs are frequently transmitted among microbes due to mobile genetic elements (MGEs). We classified MGEs into seven categories based on the criteria of MobileElementFinder (Methods), including insertion sequence (88.96%), composite transposon (5.79%), unit transposon (2.83%), and others (2.42%). The three most abundant MGEs were IS26, ISAba1, and ISCx1, which were reported to be associated with antibiotic resistance (Supplementary Fig. 5a)32,33,34. Additionally, we found that OXA-23, tet-M, and LpsB were the three most abundant ARGs closely related to the MGEs (Supplementary Fig. 5b). We next characterized the mobilome in the prominent opportunistic pathogen species using MAGs. We observed that the more prevalent opportunistic pathogen species displayed a substantially higher number of MGEs (Fig. 4a, b and Supplementary Fig. 6c–e). The length distribution of their MGEs also showed significant differences (Fig. 4c). ARGs associated with MGEs primarily confer resistance to aminoglycoside, macrolide, and cephalosporin antibiotics (Fig. 4d). The increased mobilome may contribute to the elevated prevalence of these species.
a Bar plot showing the prevalence of the top six most abundant mobile genetic elements (MGEs) identified in the LRT microbiome (n = 105). The plot includes vertical bars representing the mean values. Whiskers extend upward from the top of each bar, with the upper end of the whisker representing the mean plus one standard error (SD). b Panels indicating the average number of each type of MGEs for each species identified in our MAGs (n = 105). EHMOS represents a group of less prevalent opportunistic pathogens, including Elizabethkingia anophelis (E), Haemophilus influenzae (H), Prevotella melaninogenica (M), Prevotella oris (O), and Neisseria subflava (S). The plot includes a center line (median), box limits (upper and lower quartiles), and whiskers (1.5x interquartile range). Individual data points are shown using overlaid dot plots. Significance was determined using the two-sided Wilcoxon rank sum test. The exact p-values are directly annotated on the plot. c An analysis of the length distribution of the identified MGEs in each species. The x-axis indicates the length of MGEs, and the y-axis indicates the count. Significance was determined using the two-sided Kolmogorov-Smirnov test. The exact p-values are annotated on the plot for significant comparisons. d Sankey diagram showing the distribution of different types of MGEs (middle) identified in the five predominant species (left) linking to the antibiotic resistance genes (ARGs; right). e The schematic map for the representative plasmid identified in our samples. The gene names are labeled based on the annotation using eggNOG. f Pairwise average nucleotide identity (ANI) analysis of a highly conserved plasmid in Acinetobacter baumannii. Additional metadata, including the NCBI RefSeq ID, sampling site, and collection time of each plasmid sequence, are annotated on the plot. g The alignment map showing the whole-genome comparisons for two Acinetobacter baumannii plasmids discovered in 1982 and 2021 (the latter identified in our samples), respectively. The e-value for the two major regions was calculated using Blast. Source data are provided as a Source Data file. Abbreviation: IS, insertion sequences; MITE, miniature inverted repeats; ICE, integrative conjugative elements; ANI, average nucleotide identity.
Besides short MGEs, plasmids are the primary vehicle for horizontally transferring genetic materials. Our deep sequencing data allowed us to directly identify 68 plasmids using the Plasmid Database (PLSDB). The plasmids were primarily circular, with lengths up to tens of kilobases (Fig. 4e as an example; Supplementary Data 11). Upon comparing some of our abundant plasmids with references, we noted remarkable conservation in the plasmid sequence. Weisberg et al. and Wein et al. reported unexpected conservation and stability during plasmid transmission and evolution in plants or under non-selective conditions35,36. We focused on a plasmid commonly isolated from Acinetobacter baumannii in clinical specimens and observed a striking similarity among the analyzed plasmids spanning 40 years from different countries. The ANI between the plasmid in 2021 (our sample) and 1982 was 99.992%, or less than 1 SNP (Fig. 4f, g and Supplementary Fig. 5f). Similarly, a plasmid frequently isolated from Klebsiella pneumonia remained stable over 8 years, with a 99.955% ANI, or 3 SNPs, observed between 2022 and 2014 (Supplementary Fig. 5g, h). Interestingly, it appears that isolates of plasmids underwent extensive rearrangement events (Fig. 4g and Supplementary Fig. 5f, g). These observations starkly contrast the idea that plasmid generally mutates faster than host genomes37,38. Their long-term evolutionary stability in clinical specimens worldwide suggests a crucial role in bacterial fitness under restrictive and clinical conditions.
Identification of SNP and recombination hotspots in LRT opportunistic pathogen species
We further utilized our high-depth sequencing data and MAGs to investigate the evolution of bacterial lineages. Phylogenetic trees were constructed by integrating newly constructed MAGs with all available references, and the genetic variants were identified using the references within the same cluster (Methods). The SNPs were found to be homogeneously distributed in the genomes of some species, such as Corynebacterium striatum, Pseudomonas aeruginosa, and Stenotrophomonas maltophilia (Supplementary Fig. 6a). However, SNP-dense regions were observed in strains of Acinetobacter baumannii and Klebsiella pneumoniae (Fig. 5a, b). We were intrigued by the underlying causes of these regions.
a, b Bar plots showing the distribution of synonymous (blue) and nonsynonymous (red) single-nucleotide polymorphisms (SNPs) in coding regions of Acinetobacter baumannii (a) and Klebsiella pneumoniae (b). Shared SNP hotspot regions are shaded in light blue. c, d Nucleotide diversity and detected recombination blocks across the genome of Acinetobacter baumannii are shown in (c) and (d), respectively. e A combination of the phylogenetic tree and balloon plot showing the distribution of missense SNPs in the shared dense SNP region (near 3,000,000 bp) for Acinetobacter baumannii strains in the study. Each specific node in the phylogenetic tree is labeled with the patient ID, indicating the host of the corresponding Acinetobacter baumannii strain. The color of the patient ID indicates the hospital. Each locus tag represents an identified gene. The size and color of each circle indicate the percentage of the missense SNPs. The red dashed box highlights the region with dense missense SNPs, as described in the main text. f Schematic representation of the putative prophage identified in the SNP hotspot of Klebsiella pneumoniae, showing gene features, nucleotide diversity, and SNP distribution. The boxes indicate the presence (filled) or absence (open) of identified SNPs within the corresponding genes for each patient’s MAG. Source data are provided as a Source Data file.
Phylogenetic analysis based on 1,917 conserved proteins shows that all our Acinetobacter baumannii strains belonged to a single cluster dominated by clinical isolates (Supplementary Fig. 6b). Three SNP-dense regions, especially a conserved region near 3,000,000 bp, were present (Fig. 5a). We calculated the genome-wide nucleotide diversity (π), which unveiled elevated genetic diversity in the three regions (Fig. 5c). SNP-dense regions are a well-known signature for recombination39,40, a major driving force behind the evolution of bacteria41. We calculated the recombination rate (r/m) and identified recombination hotspots in these three SNP-dense regions (Fig. 5c, d and Supplementary Fig. 6d). Similarly, we also observed three SNP-dense regions for Klebsiella pneumoniae strains within the same cluster (Fig. 5b). However, the recombination hotspots did not strictly correspond to the SNP hotspots (Supplementary Fig. 6e, f), suggesting that Klebsiella pneumoniae may acquire SNPs through non-recombining mechanisms, such as mutations.
Exploring the functional impact of the recombination hotspots
The recombination hotspot in bacteria is likely driven by selective pressure, such as host immunity and clinical practices. We observed consistently elevated missense SNPs in the SNP-dense regions, indicating that the functions of genes may be directly impacted (Fig. 5a, b). We analyzed the associated functions of the genes in the recombination hotspots of Acinetobacter baumannii and Klebsiella pneumoniae. Most genes were associated with critical metabolic pathways, such as carbohydrate metabolism, glycan biosynthesis and metabolism, and amino acid metabolism (Supplementary Fig. 6g, h). Other pathways were mainly related to cellular processes and environmental information processing, indicating the roles of these proteins in environmental adaptations.
Specifically, in one of the shared SNP-dense regions of Acinetobacter baumannii (Locus 25-34, Fig. 5e), we identified genes associated with bacterial conjugation, the toxin-antitoxin (TA) system, and recombination (Supplementary Data 12). For example, we found the gene with the highest SNPs (Locus 33) to be virB5, a protein involved in the type IV secretion systems (T4SSs), and the adjacent gene is associated with the ParE toxin of type II toxin-antitoxin system, both essential to bacterial survival and bacterial interactions with the environmental biotic and abiotic stresses in ICU settings42,43.
Chaguza et al. reported that SNP hotspots in Streptococcus pneumoniae could be associated with phage DNA44. Interestingly, we found a near-complete putative prophage in the shared SNP-dense region of Klebsiella pneumoniae (Fig. 5f). These genes encode phage attachment proteins, including tail tape measure, tail, and head-tail adapter proteins. In total, our results demonstrated frequent regional recombinations as a driving force behind genetic variation in A. baumannii and K. pneumoniae, with associated genes involved in key metabolic, stress response, and survival pathways, suggesting their role in adaptation to clinical environments.
The whole-genome comparisons reveal putative strain transmission events within a single ward and between wards
Based on analysis of clinical cultured isolates, it has been previously demonstrated that putative patient strain transmission events were associated with specific room types and environmental microbiomes45,46,47. Here, using whole-genome SNP comparisons and longitudinal sampling, we provide robust evidence for 12 putative strain transmission events among three centers for Acinetobacter baumannii and Klebsiella pneumoniae, utilizing high-quality MAGs directly recovered from LRT samples (Methods; Fig. 6a; Supplementary Data 13 and 14). Additionally, we used in vitro cultured isolates recovered from the corresponding samples to validate the genome-wide average nucleotide identity (ANI) for 8 out of 12 putative strain transmission events (Supplementary Data 14). We observed strikingly high 99.9995% to 100% average nucleotide identity (ANI) within 0–16 SNPs differences between strains from patients in single- or multi-occupancy ICU wards. Moreover, of the 12 putative transmission events involving 15 patients, longitudinal sampling data were available for 10 patients. To further substantiate the putative transmission events, we investigated the temporal changes in the relative abundance of transmitted species. For at least 5 patients, the species of interest was undetected upon ICU admission but identified on subsequent sampling days (Supplementary Fig. S7). The remaining patients acquired the transmitted species before the first sampling point, indicating they were possibly infected earlier. This temporal pattern provides additional evidence supporting the putative transmission events. Notably, the frequent putative transmission events observed between patients in two separate single-occupancy rooms indicate that these opportunistic pathogens could be widespread in ICU environments, possibly disseminating through the staff or ventilation systems (Supplementary Data 13). For instance, P25 and P77 were hospitalized in two single-occupancy ICU rooms with a ten-day overlapping hospitalization period; however, their Acinetobacter baumannii strains shared a 100% ANI with 0 SNPs (Supplementary Data 13). Interestingly, in five putative transmission events, there were gaps up to dozens of days between hospitalization periods (Supplementary Data 13), suggesting indirect transmission events through a potential microbial reservoir, such as bed rails, door handles, and sink traps45,48. Importantly, in direct and indirect transmission events, ANI and SNPs showed little difference (Fig. 6b, c).
a The chart represents putative strain transmission events in the ICUs of three hospitals. Each row represents an individual patient, and the length of the segment represents the length of stay (from admission to discharge). The segment color indicates the hospital. A line connecting two patients represents a transmission event. A solid line denotes a potential direct transmission event with an overlapping hospitalization period between two patients (direct transmission); A dashed line denotes transmission events that cannot be explained by overlapping hospitalization periods (indirect transmission), which may result from a potential microbial reservoir in the hospital. b, c Violin plots showing the ANI (b) and SNP number (c) comparisons for the strains in patients with direct (n = 7) or indirect transmissions (n = 5). The plot includes a center line (median), box limits (upper and lower quartiles), and whiskers (1.5x interquartile range). Significance was determined using the two-sided Wilcoxon rank sum test. The exact p-values are directly annotated on the plot. Source data are provided as a Source Data file.
These results demonstrated the feasibility of directly tracing between-patient transmission events with our approach, indicating that patients in the ICU are subjected to acquiring pathogens from other patients and potential environmental microbial reservoirs. A systematic investigation of microbial transmission events and environmental microbiomes is critical for evaluating the biomedical safety of clinical setups.
Discussion
Due to the challenges associated with LRT sampling and processing, the LRT microbiome has yet to be markedly underexplored compared with the gut, skin, and oral microbiomes49. Most recent LRT studies used targeted amplicon sequencing approaches, such as 16S rRNA, which have inherent limitations in genomic, functional, and evolutionary analyses50. Therefore, there is a desperate need for an efficient microbial-enrichment processing method.
With the introduction of Chelex100, we developed a highly efficient experimental method to enrich low microbial biomass from host-rich samples. Using this method, we systematically profiled the longitudinal LRT microbial community, resistome, and strain-based functional, genomic, and evolutionary dynamics in critically ill patients. The abundance of several opportunistic pathogen species among pneumonia patients significantly varied across ICUs (Fig. 2a, b). Furthermore, the total abundance of each ARG in these pneumonia patients also showed notable variation (Fig. 2f and Supplementary Fig. 3b). The distinct site-specific signatures of microbiomes and resistomes in pneumonia patients across ICUs may be associated with their disparate environmental prevalence in the respective ICU environments, supporting the critical need for comprehensive profiling of environmental microbiomes in ICU settings. We also observed that specific multidrug resistance genes accumulated in patients with ICU stays exceeding 28 days (Fig. 2i,j), indicating that the increased abundance of multidrug genes may contribute to antibiotics treatment failure and prolonged ICU stay.
We acquired 433 MAGs (including bacteria, viruses, and fungi) directly from the metagenomic sequencing data at the individual level, 108 of which were unknown. Compared to conventional culture-based methods, CMEM enables the simultaneous characterizations of different species without laborious culturing. Additionally, the successful recovery of MAGs for several rare opportunistic pathogenic species (such as Aggregatibacter segnis and Anaeroglobus geminatus; Supplementary Data 8) demonstrates CMEM’s potential in recovering genomes of undescribed or difficult-to-culture microbes directly from critically ill patients.
Importantly, high-quality MAGs, in lieu of isolates’ genomes, can be directly exploited to investigate the strain-level resistome, virulome, mobilomes, inter-patient transmissions, and evolutionary dynamics. For example, based on the MAGs, we parsed strain-level resistome and virulome variations associated with individual patients, which may be critical for tailored treatments. Interestingly, we also recovered plasmids that showed remarkable SNP-level conservation over decades across countries. The frequent chromosomal rearrangements instead of mutations in these plasmids may have profound implications for plasmid evolution, calling for more systematic investigations into the evolution of pathogen-related plasmids in clinical settings.
Previous studies utilizing culture-based isolates have demonstrated that frequent recombination is the major evolutionary driver for opportunistic pathogen species, allowing them to evolve rapidly in response to selective pressures such as antibiotic treatment39,40,41,44,51,52. In this study, we used MAGs directly recovered from LTR samples to identify frequent recombinations at previously unknown genomic regions as one of the major evolutionary driving forces for A. baumannii and K. pneumoniae strains, which are widespread in the ICU environment. Finally, we detected frequent putative patient-to-patient strain transmission events by genome-wide comparisons (Fig. 6a). Some showed a gap period of at least a few days, suggesting a hospital-specific reservoir may exist. There is a pressing need to understand better how microbial reservoirs can affect patients’ LRT microbiomes in clinical settings, again calling for investigating how the environmental microbial species transmit to and eventually inflict harm on patients53.
Our study has several limitations. Firstly, our approach is limited by its potentially reduced ability to characterize viruses from LRT samples with the host-removal step. The host-removal step also led to noticeable changes in the relative abundance of microbial taxa. While the most abundant species in the samples maintained their dominance, measurements of very low-abundance taxa (average relative abundance < 0.5%) may be inaccurate. The performance of CMEM in handling other types of respiratory specimens, such as sputum or BALF, may differ and warrants further evaluation. Secondly, the complexity of our cohorts, such as the heterogeneity of the health conditions of these critically ill patients, the diverse treatment they received, and differences in commodities may have contributed to additional variances in their respiratory microbiomes. Thirdly, our study only included ETA samples. The utilization of alternative respiratory specimen types, such as bronchoalveolar lavage fluid (BALF) or bronchial brushings, may yield varying findings. Finally, our primary findings were predominantly derived from the metagenomic sequencing data and MAGs, with several findings validated with the cultured isolate approach. However, some results, such as the observed associations between various temporal dynamics patterns of LRT microbiomes and pulmonary functions, require more detailed mechanistic experiments to verify the potential causal relationships.
To conclude, we developed a highly efficient microbial-enrichment experimental pipeline to deep sequence and directly reconstruct high-quality MAGs from LRT samples without culturing, enabling further sophisticated analyses. We provided longitudinal profiling of the microbiome and resistome, as well as a rare genome-resolved functional, evolutionary, and transmission perspective of the LRT microbiota in ICU pneumonia patients. We believe the CMEM method will be a powerful tool for in-depth LRT microbial surveillant, functional, evolutionary, and clinical research.
Methods
Study cohort
This multicenter, retrospective study was conducted at the ICUs of the First Affiliated Hospital of Zhejiang University of School of Medicine (general ICU; Hangzhou), Peking University Third Hospital (surgical ICU; Beijing), and the Second Xiangya Hospital of Central South University (respiratory ICU; Changsha) in China. Samples were collected longitudinally from 157 intubated patients in the ICU between February 2021 and July 2022. The study was approved by the Clinical Research Ethics Committee of the First Affiliated Hospital (reference: 1T20221227A), Peking University Third Hospital Medical Science Research Ethics Committee (reference: IRB00006761-M2022419), and the Medical Ethics Committee of the Second Xiangya Hospital (reference: LYG2020093). The written informed consent was obtained from all the participants for the collection of respiratory samples and the use of their relevant clinical data. Patients admitted to the ICU were screened based on the following criteria: (1) adult (>18 years of age) with an index admission to ICU; (2) requirement for mechanical ventilation, with an anticipated need for continuous mechanical ventilation (>72 h) as judged by the treating ICU specialist; and (3) absence of any pre-existing and transmissible diseases.
From the 157 enrolled patients, 453 endotracheal aspirates (ETA) were initially collected for downstream analysis. Of these, 11 samples were excluded due to insufficient DNA yielding, making the final sample size 442. Sixty-three patients were sampled more than three times, 32 were sampled twice, and 62 were sampled once. Patient demographic and clinical data were recorded at the time of admission to the ICU and are detailed in Supplementary Data 1.
The status of LRTI was adjudicated by the clinicians referenced on the previously reported criteria25,54, classifying the patients into (1) Diagnosed pneumonia, defined as clinically documented pneumonia in the medical record system, with clinical positive microbiological culture results or computed tomography (CT) images (2) Non-pneumonia, defined as no clinically documented pneumonia in the medical records and the lack of microbiological or CT evidence of respiratory infection. (3) Undefined LRTI status, assigned to patients without clinically relevant records to determine LRTI status. To mitigate the potential influence of antibiotic usage, all patients included in the comparisons between the pneumonia and non-pneumonia groups were treated with antibiotics during the ICU stay. The oxygenation index (OI) was calculated by dividing the partial pressure of arterial oxygen (PaO2) by the fraction of inspired oxygen (FiO2).
Optimized LRT microbial-enrichment DNA extraction method
We first attempted reported host DNA removal methods for 26 of our samples23. However, the low microbial biomass resulted in undetectable DNA concentrations (measured by Qubit) for approximately half of the samples, making it impractical for metagenomic sequencing. Hence, we developed an experimental method with the following procedures:
Host DNA depletion
Respiratory samples (400 µL), including ETAs and BALF, were first treated with a 1:1 mixture of sputasol and incubated for a minimum of 15 min at 37 °C. Sputasol-treated samples were then subjected to the procedure for host DNA depletion, modified from a previously reported saponin-based differential lysis method23. Briefly, the mixture of respiratory samples and sputasol was centrifuged at 15,000 g for 5 min, and the supernatant was removed while not disturbing the pellet. The pellet was resuspended in 250 µL of PBS supplemented with saponin (Tokyo Chemical Industry, Tokyo, Japan) to a final concentration of 2% and incubated at room temperature for 10 min. After the incubation, 350 µL of nuclease-free water was added and incubated for 30 s, after which 12 µL of 5 M NaCl was added. Samples were next centrifuged at 10,000 g for 5 min, with the supernatant removed and the pellet resuspended in 100 µL of PBS. Subsequently, 100 µL of HL-SAN buffer (5.5 M NaCl and 100 mM MgCl2 in nuclease-free water) and 12 µL of HL-SAN DNase (ArcticZymes, Tromsø, Norway) were added and incubated for 15 min at 37 °C with shaking at 800 rpm. After the incubation, samples were washed twice with PBS (800 µL and 1 mL) and centrifuged at 8,000 g for 5 min after each wash. Finally, the supernatant was discarded, and the pellet was resuspended in 100 µL of PBS.
DNA extraction
Following the host DNA depletion, the resuspended pellet underwent an additional 30 min sonication at 65 °C (Scientz Biotechnology, Ningbo, China) to promote microbial cell lysis55. A 5% Chelex 100 solution (Bio-Rad, Hercules, USA) was subsequently added in a volume of 120 µL. The mixture was thoroughly mixed and then incubated at 65 °C for 20 min in a heating block. The mixture was vortexed for 30 s and heated at 95 °C for 10 minutes. After the heating process, the tube was vortexed again for 30 s and centrifuged at 15,000 g for 2 min. The supernatant was pipetted into a new tube.
Precipitation of proteins and DNA
After the DNA extraction with Chelex 100, ammonium acetate was introduced to precipitate proteins, after which sodium acetate was added to concentrate the final DNA. The supernatant was then treated with a 7.5 M stock solution of ammonium acetate to achieve a final working concentration of 2.5 M and then promptly incubated for 5 minutes on ice. The sample was vortexed for 5 s and centrifuged at 15,000 g for 10 min at room temperature, after which the supernatant was carefully transferred to a new tube. Subsequently, 10% of the 3 M sodium acetate (pH 5.5) and two volumes of 100% ethanol were added to the supernatant and incubated at −20 °C for 3 h. The solution was centrifuged at 15,000 g for 30 min at 4 °C, and the supernatant was pipetted out. The pellet was then washed with 200 µL of 70% ethanol and stored at −20 °C. The solution was vortexed for 15 s and centrifuged at 15,000 g for 15 min at 4 °C, after which the supernatant was carefully removed. Finally, the pellet was allowed to air dry for 5 min and resuspended in 100 µL of nuclease-free water, and the clear supernatant containing genomic DNA was collected. The quantity of DNA was measured using the Qubit dsDNA HS Assay Kit (Vazyme #EQ121, Nanjing, China) on the Qubit 3.0 Fluorometer (Thermo Fisher Scientific, Waltham, USA).
Metagenomic sequencing library preparation and sequencing data pre-processing
According to the manufacturer’s protocol, metagenomic libraries with an insert size of 350 bp were prepared using a VAHTS Universal Plus DNA Library Prep Kit (Vazyme #ND617, Nanjing, China). The quantity and quality of libraries were assessed using the Qubit dsDNA HS Assay Kit (Vazyme #EQ121, Nanjing, China) and Qsep100 (BiOptic, New Taipei City, China). The final libraries were sequenced for 2 × 150 bp paired-end sequencing on the Illumina NovaSeq 6000 platform. The raw sequencing reads were processed with Fastp56 to remove adapter sequences and trim low-quality bases. Human reads were removed using a high-performance two-stage bioinformatics approach57, in which reads were first aligned to human genome GRCh38 using Bowtie258 and underwent a secondary alignment using HISAT259.
Negative controls
For each hospital, ten negative environmental sampling control samples were collected. Sterile saline solution (Hospital A: Kelun Pharmaceutical Co., Ltd, H43020456, China; Hospital B: Otsuka Pharmaceutical Co., Ltd, H12020024, China; Hospital C: Kelun Pharmaceutical Co., Ltd, H43020456, China) was collected by aspiration through the sputum aspirator as a negative sampling control. Ten reagent controls (DNA extraction blanks) were collected at the laboratory that handles study samples. All thirty negative sampling controls and ten reagent controls were included for library preparation and sequencing. Bacterial species identified in more than two negative controls with relative abundance >0.001 were considered potential contaminants and excluded from downstream analyses (Supplementary Data 5, 6, 15, and 16). The sequencing data from our samples were remapped against the MAGs recovered from the negative controls using Bowtie2. The average proportion of mapped reads across our samples is 0.31% (median 0.12%, interquartile range 0.06%-0.48%).
Bacterial culture and isolation
ETA samples were diluted and homogenized in phosphate-buffered saline (PBS), followed by inoculation onto Columbia Blood Agar (Hopebio, Qingdao, China) and Blood Agar Plates (Hopebio, Qingdao, China). For samples with a high abundance of Pseudomonas aeruginosa, the CN Agar Plates (Hopebio, Qingdao, China) were used for selective isolation to isolate Pseudomonas aeruginosa. Plates were incubated at 37 °C for 48 h under ambient air conditions. Single colonies were picked and streaked onto fresh plates under the same culture conditions to obtain pure isolates. Isolate DNA was extracted and purified using Chelex-100 (Bio-Rad, Hercules, USA), following the same procedure as for ETA samples. Library construction and sequencing of isolates were performed identically to the ETA sample processing.
Microbial community profiling and metrics
Taxonomic profiling and relative abundance of species were determined based on the exact alignment of unique clade-specific markers with MetaPhlAn3 v.3.0.1360 on human-removed sequences. Functional profiling was performed using HUMAnN3 v.3.760, with the abundance of functional pathways classified against UniRef9061 and ChocoPhlAn databases. We normalized raw count values for sequencing depth and merged multiple tables using the ‘humannrenormtable’ and ‘humannjointables’ utility scripts. Diversity metrics and Bray-Curtis (BC) distances were calculated based on MetaPhlAn3 profiling outputs using the phyloseq v.1.40.062 and vegan v.2.6.4 (https://github.com/vegandevs/vegan) package.
Analysis of antibiotic resistance genes
For the read-based identification of ARGs, all processed reads were classified and annotated against the SARG database using ARG-OAP v.2.563 with default parameters. ARGs were categorized into ARG types (the class of antibiotics targeted by the gene) and subtypes (functional gene annotation). ARG abundance was normalized by the number of 16S reads, and richness was calculated as the total number of ARGs. The dissimilarity matrix based on the Bray-Curtis distance was created from the normalized ARG abundance using the vegan package. For the contig-based profiling of ARGs in metagenomic assemblies, contigs were annotated against CARD v.3.2.564 using Resistance Gene Identifier (RGI) v.6.0.1 available as a command-line tool downloaded from the CARD website with default parameters.
Linear mixed effect (LME) modeling
We explored the relationship between microbial features and demographic and clinical variables using LME models. The LME models treated the subjects (participants) and time points (sampling date) as the random effect. The multivariate LME model was established between the normalized microbiome features and demographic and clinical variables using the ‘lmer()’ function of the R package lmer4 v1.1.32, with age, sex, BMI, and sampling site adjusted as covariates: dependent variable ~(intercept) + independent variable + age + sex + BMI + sampling site + (1| time point) + (1| subject). The continuous variables were scaled before LME modeling to calculate the standardized beta coefficients. The Benjamini-Hochberg procedure was applied to control FDR.
De novo genome assembly, quality assessment, binning, and annotation
The pre-processed reads were assembled into draft genomes using MEGAHIT v.1.2.965 at the individual level, and the quality of genome assemblies was assessed with CheckM v.1.1.566. All assemblies were initially binned using MaxBin v.2.2.767, with a secondary binning process using MetaWRAP v.1.3.068 employed if the contamination rate of the desired bins exceeded 5%. The species-level taxonomic classification of assemblies was performed using GTDB-Tk v.2.0.069 against GTDB70. Genomic bins that met the criteria (completeness > 90% and contamination < 5%) were classified as high quality and were subjected to further strain-level analysis. Genes were identified using Prodigal71 with ‘meta’ mode and annotated using KofamKOALA72 to search against the Kyoto Encyclopedia of Genes and Genomes (KEGG)73. Virulence genes were screened against VFDB74 using ABRicate v.1.0.1 (https://github.com/tseemann/abricate).
Detection of putative strain-transmission events
To investigate the possible strain-transmission events, we integrated both genetic and epidemiological information and referenced the previously reported methodologies46,75,76. Raw reads corresponding to genomes for potential outbreak species were jointly analyzed with their respective high-quality draft genomes using inStrain v.1.5.776 to detect the SNPs and ANI through whole-genome pairwise alignment. Specifically, a potential transmission event was confirmed when meeting the criteria: (1) a highly stringent cutoff of 99.9995% ANI between two compared genomes; (2) the fraction of compared bases higher than 90%; and (3) an overlapping hospitalization period recorded in clinical data for patient-to-patient transmission.
Phylogenetic analysis and SNPs calling
First, to estimate the genetic distance and overall population structure for these predominant opportunistic pathogens detected in our samples, we downloaded all available reference genomes for these species from NCBI. Second, a set of core marker genes belonging to each species, downloaded from Uniref90, were used to construct a high-resolution phylogenetic tree for each species. The alignment and concatenation of core marker genes were performed using PhyloPhlAn3 v.3.0.677, and the final phylogenetic tree was inferred using the maximum-likelihood method with RAxML v.8.2.1278, incorporating our draft genome, publicly available reference genomes, and outgroups. All trees were visualized using the ggtree package79. Pair-end reads for each species derived from the individual patient were aligned to a closely related reference genome, selected based on the phylogeny result demonstrating it was within the same cluster of our draft genomes and its geographical proximity to our sampling site. Variant calling was performed using Snippy v.4.6.0 (https://github.com/tseemann/snippy) with default parameters, requiring a minimum of 10-fold read depth for calling alleles. Samples with a breadth of coverage for the reference below 90% were additionally excluded.
Recombination hotspot detection and population genetics metrics
The homologous recombination regions were detected using Gubbins v.3.2.180 based on the whole genome sequence alignments generated by Snippy. We calculated evolutionary parameters, including the relative substitution rate due to recombination versus mutation (r/m) and the number of recombination blocks relative to the number of SNPs outside the recombination blocks (ρ/θ) based on the Gubbins output files. Population genetic metrics, such as nucleotide diversity (π), were calculated using VCFtools v.0.1.1681 based on the VCF files generated with samtools v.1.1182 and bcftools v.1.15.183. Genes annotation was performed using eggNOG-mapper v.2.084 to search against the eggNOG 5.0 database85. Recombination blocks and reference genomes were visualized using phandango86, and gene features were generated using DNA Features Viewer package87.
Mobile genetic element identification
All the assembled contigs from our samples were analyzed to explore the full spectrum of MGEs using MobileElementFinder v.1.0.388, which categorized MGEs into Miniature Inverted Repeats (MITEs), Insertion Sequences (ISs), Composite Transposons (ComTns), Unit Transposons (Tns), Integrative Conjugative Elements (ICEs), Integrative Mobilizable Elements (IMEs) and Cis-Mobilizable Elements (CIMEs). Species-level characterization of MGEs was performed using high-quality genomic bins. Only high-quality MGEs were retained if the identity > 90% and coverage > 95%. An antibiotic resistance gene was considered associated with MGEs if located within an MGE or with an interval < 5 kb89.
Plasmid identification
Complete plasmid sequences were retrieved from the PLSDB v.20210623_v290, and Mash v.2.391 sketches were generated following the instructions on the PLSDB website. Plasmid sequences were screened from our assembled contigs using Mash dist, with the Mash distance < 0.001. Plasmids were annotated using Prokka v.1.14.692, and ANI comparisons were performed using FastANI v.1.3393 with a smaller fragment length (500 bp). Pairwise genome alignments of similar plasmids were conducted using Mauve v.2.4.094, and comparative graphics were visualized using Kablammo95 and genoPlotR package96. The angularplasmids package (https://github.com/vixis/angularplasmid) was used to visualize representative plasmid sequences.
Statistics & Reproducibility
Microbiome and resistome alpha diversity metrics were analyzed using the phyloseq v.1.40.0 and vegan v.2.6.4 package. Microbiota β-diversity was calculated on the Bray–Curtis dissimilarity by PERMANOVA using the ‘adonis’ function in the vegan v.2.6.4 package. Statistical analyses of NMDS of total ARG abundance among three hospitals were performed using multivariable PERMANOVA in the vegan v.2.6.4 package. A two-tailed Wilcoxon (paired) or Mann–Whitney U-test (unpaired) test was used for two-group comparisons. The ‘splsda’ function of the mixomics v.6.20 package was used to identify ARGs positively or negatively associated with resistome profile variations among three hospitals. Correlations were assessed using the ‘cor.test’ function of the stats v.4.2.3 package. Kolmogorov–Smirnov test was used for the MGE length distribution comparisons. All statistical tests and associated diagrams generation were conducted in R v.4.2.0. No data were excluded from analyses and no statistical method was used to predetermine sample size.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All human-removed metagenomic sequencing data and cultured isolate sequencing data generated in this study have been deposited in the European Nucleotide Archive (https://www.ebi.ac.uk/ena/browser/home) under the project accession number PRJEB64676. The processed data and analytical code are available at the Github repository: https://github.com/processbys/LRT_microbiomes_critical_illness [https://zenodo.org/doi/10.5281/zenodo.13627161]97. The source data generated in this study for the figures are provided in the Source Data file. Source data are provided with this paper.
Code availability
The analytical code is available at https://github.com/processbys/LRT_microbiomes_critical_illness [https://zenodo.org/doi/10.5281/zenodo.13627161]97.
References
World Health Organization. The Top 10 Causes of Death. https://www.who.int/en/news-room/fact-sheets/detail/the-top-10-causes-of-death (2017).
el Bcheraoui, C. et al. Trends and patterns of differences in infectious disease mortality among US counties, 1980-2014. JAMA 319, 1248–1260 (2018).
Ruan, Z. et al. Disease burden and attributable risk factors of respiratory infections in China from 1990 to 2019. Lancet Reg. Health West. Pac. 11, 100153 (2021).
Vincent, J.-L. et al. Prevalence and outcomes of infection among patients in intensive care units in 2017. JAMA 323, 1478–1487 (2020).
Laupland, K. B. et al. Population-based assessment of intensive care unit-acquired bloodstream infections in adults: Incidence, risk factors, and associated mortality rate. Crit. Care Med. 30, 2462 (2002).
Dickson, R. P. et al. Lung microbiota predict clinical outcomes in critically Ill patients. Am. J. Respir. Crit. Care Med 201, 555–563 (2020).
Kitsios, G. D. et al. Respiratory tract dysbiosis is associated with worse outcomes in mechanically ventilated patients. Am. J. Respir. Crit. Care Med. 202, 1666–1677 (2020).
Kelly, B. J. et al. Composition and dynamics of the respiratory tract microbiome in intubated patients. Microbiome 4, 7 (2016).
Qi, X. et al. Lower respiratory tract microbial composition was diversified in Pseudomonas aeruginosa ventilator-associated pneumonia patients. Respir. Res. 19, 139 (2018).
Wei, X. et al. MASS cohort: Multicenter, longitudinal, and prospective study of the role of microbiome in severe pneumonia and host susceptibility. Imeta. 3, e218 (2024).
Berton, D. C., Kalil, A. C. & Teixeira, P. J. Z. Quantitative versus qualitative cultures of respiratory secretions for clinical outcomes in patients with ventilator‐associated pneumonia. Cochrane Database Syst. Rev. 10, CD006482 (2014).
Pereira-Marques, J. et al. Impact of host DNA and sequencing depth on the taxonomic resolution of whole metagenome sequencing for microbiome analysis. Front. Microbiol. 10, 1277 (2019).
Segal, L. N. et al. Enrichment of the lung microbiome with oral taxa is associated with lung inflammation of a Th17 phenotype. Nat. Microbiol 1, 1–11 (2016).
Peng, Z., Zhou, J. & Tian, L. Pathogenic characteristics of sputum and bronchoalveolar lavage fluid samples from patients with lower respiratory tract infection in a large teaching hospital in China: a retrospective study. BMC Pulm. Med. 20, 233 (2020).
Chiu, C. Y. & Miller, S. A. Clinical metagenomics. Nat. Rev. Genet. 20, 341–355 (2019).
Hong, S., Bunge, J., Leslin, C., Jeon, S. & Epstein, S. S. Polymerase chain reaction primers miss half of rRNA microbial diversity. ISME J. 3, 1365–1373 (2009).
Angly, F. E. et al. CopyRighter: a rapid tool for improving the accuracy of microbial community profiles through lineage-specific gene copy number correction. Microbiome 2, 11 (2014).
Sulaiman, I. et al. Evaluation of the airway microbiome in nontuberculous mycobacteria disease. Eur. Respir. J. 52, 1800810 (2018).
Li, C. et al. Diagnostic value of bronchoalveolar lavage fluid galactomannan assay for invasive pulmonary aspergillosis in adults: a meta-analysis. J. Clin. Pharm. Ther. 47, 1913–1922 (2022).
Lloyd-Price, J. et al. Strains, functions and dynamics in the expanded human microbiome project. Nature 550, 61–66 (2017).
Yan, Y., Nguyen, L. H., Franzosa, E. A. & Huttenhower, C. Strain-level epidemiology of microbial communities and the human microbiome. Genome Med. 12, 71 (2020).
Zou, Y. et al. 1,520 reference genomes from cultivated human gut bacteria enable functional microbiome analyses. Nat. Biotechnol. 37, 179–185 (2019).
Charalampous, T. et al. Nanopore metagenomics enables rapid clinical diagnosis of bacterial lower respiratory infection. Nat. Biotechnol. 37, 783–792 (2019).
Yang, L. et al. Metagenomic identification of severe pneumonia pathogens in mechanically-ventilated patients: a feasibility and clinical validity study. Respir. Res. 20, 265 (2019).
Langelier, C. et al. Integrating host response and unbiased microbe detection for lower respiratory tract infection diagnosis in critically ill adults. Proc. Natl Acad. Sci. USA 115, E12353–E12362 (2018).
Pérez-Cobas, A. E., Ginevra, C., Rusniok, C., Jarraud, S. & Buchrieser, C. The respiratory tract microbiome, the pathogen load, and clinical interventions define severity of bacterial pneumonia. CR Med. 4, 101167 (2023).
Bowers, R. M. et al. Minimum information about a single amplified genome (MISAG) and a metagenome-assembled genome (MIMAG) of bacteria and archaea. Nat. Biotechnol. 35, 725–731 (2017).
Santella, B. et al. Lower respiratory tract pathogens and their antimicrobial susceptibility pattern: a 5 year study. Antibiotics 10, 851 (2021).
Wu, Y. et al. Resident bacteria contribute to opportunistic infections of the respiratory tract. PLOS Pathog. 17, e1009436 (2021).
Qin, S. et al. Pseudomonas aeruginosa: pathogenesis, virulence factors, antibiotic resistance, interaction with host, technology advances and emerging therapeutics. Sig Transduct. Target Ther. 7, 1–27 (2022).
Gomila, M. et al. Genetic diversity of clinical Pseudomonas aeruginosa isolates in a public hospital in Spain. BMC Microbiol. 13, 138 (2013).
Harmer, C. J. & Hall, R. M. IS26-mediated formation of transposons carrying antibiotic resistance genes. mSphere 1, e00038-16 (2016).
Noel, H. R., Petrey, J. R. & Palmer, L. D. Mobile genetic elements in Acinetobacter antibiotic-resistance acquisition and dissemination. Ann. N. Y. Acad. Sci. 1518, 166–182 (2022).
Schröder, J. et al. Complete genome sequence, lifestyle, and multi-drug resistance of the human pathogen Corynebacterium resistens DSM 45100 isolated from blood samples of a leukemia patient. BMC Genom. 13, 141 (2012).
Weisberg, A. J. et al. Unexpected conservation and global transmission of agrobacterial virulence plasmids. Science 368, eaba5256 (2020).
Wein, T., Hülter, N. F., Mizrahi, I. & Dagan, T. Emergence of plasmid stability under non-selective conditions maintains antibiotic resistance. Nat. Commun. 10, 2595 (2019).
Rodríguez-Beltrán, J. et al. High recombinant frequency in extraintestinal pathogenic Escherichia coli strains. Mol. Biol. Evol. 32, 1708–1716 (2015).
Niaudet, B., Jannière, L. & Ehrlich, S. D. Recombination between repeated DNA sequences occurs more often in plasmids than in the chromosome of Bacillus subtilis. Mol. Gen. Genet. 197, 46–54 (1984).
Chewapreecha, C. et al. Dense genomic sampling identifies highways of pneumococcal recombination. Nat. Genet. 46, 305–309 (2014).
Snitkin, E. S. et al. Genome-wide recombination drives diversification of epidemic strains of Acinetobacter baumannii. Proc. Natl Acad. Sci. USA 108, 13758–13763 (2011).
Spratt, B. G., Hanage, W. P. & Feil, E. J. The relative contributions of recombination and point mutation to the diversification of bacterial clones. Curr. Opin. Microbiol. 4, 602–606 (2001).
Fronzes, R., Christie, P. J. & Waksman, G. The structural biology of type IV secretion systems. Nat. Rev. Microbiol 7, 703–714 (2009).
Jurėnas, D., Fraikin, N., Goormaghtigh, F. & Van Melderen, L. Biology and evolution of bacterial toxin–antitoxin systems. Nat. Rev. Microbiol. 20, 335–350 (2022).
Chaguza, C. et al. Within-host microevolution of Streptococcus pneumoniae is rapid and adaptive during natural colonisation. Nat. Commun. 11, 3442 (2020).
Chng, K. R. et al. Cartography of opportunistic pathogens and antibiotic resistance genes in a tertiary hospital environment. Nat. Med. 26, 941–951 (2020).
Snitkin, E. S. et al. Tracking a hospital outbreak of carbapenem-resistant Klebsiella pneumoniae with whole-genome sequencing. Sci. Transl. Med. 4, 148ra116–148ra116 (2012).
Wei, X. et al. Charting the landscape of the environmental exposome. Imeta. 1, e50 (2022).
Jiang, C. et al. Dynamic Human Environmental Exposome Revealed by Longitudinal Personal Monitoring. Cell 175, 277–291.e31 (2018).
Di Simone, S. K., Rudloff, I., Nold-Petry, C. A., Forster, S. C. & Nold, M. F. Understanding respiratory microbiome–immune system interactions in health and disease. Sci. Transl. Med. 15, eabq5126 (2023).
Wei, X. et al. Vaginal microbiomes show ethnic evolutionary dynamics and positive selection of Lactobacillus adhesins driven by a long-term niche-specific process. Cell Rep. 43, 114078 (2024).
Croucher, N. J. et al. Rapid pneumococcal evolution in response to clinical interventions. Science 331, 430–434 (2011).
Shaidullina, E. R. et al. Genomic analysis of the international high-risk clonal lineage Klebsiella pneumoniae sequence type 395. Genome Med. 15, 9 (2023).
Wei, X. et al. Charting the landscape of the environmental exposome. iMeta 1, e50 (2022).
Chu, V. T. et al. The antibiotic resistance reservoir of the lung microbiome expands with age in a population of critically ill patients. Nat. Commun. 15, 92 (2024).
Luhung, I. et al. Protocol Improvements for low concentration DNA-based bioaerosol sampling and analysis. PLoS ONE 10, e0141158 (2015).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Bush, S. J., Connor, T. R., Peto, T. E. A., Crook, D. W. & Walker, A. S. Evaluation of methods for detecting human reads in microbial sequencing datasets. Microb. Genom. 6, e000393 (2020).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Beghini, F. et al. Integrating taxonomic, functional, and strain-level profiling of diverse microbial communities with bioBakery 3. eLife 10, e65088 (2021).
Suzek, B. E. et al. UniRef clusters: a comprehensive and scalable alternative for improving sequence similarity searches. Bioinformatics 31, 926–932 (2015).
McMurdie, P. J. & Holmes, S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8, e61217 (2013).
Yin, X. et al. ARGs-OAP v2.0 with an expanded SARG database and hidden Markov models for enhancement characterization and quantification of antibiotic resistance genes in environmental metagenomes. Bioinformatics 34, 2263–2270 (2018).
Alcock, B. P. et al. CARD 2020: antibiotic resistome surveillance with the comprehensive antibiotic resistance database. Nucleic Acids Res. 48, D517–D525 (2020).
Li, D., Liu, C.-M., Luo, R., Sadakane, K. & Lam, T.-W. MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 31, 1674–1676 (2015).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Wu, Y.-W., Simmons, B. A. & Singer, S. W. MaxBin 2.0: an automated binning algorithm to recover genomes from multiple metagenomic datasets. Bioinformatics 32, 605–607 (2016).
Uritskiy, G. V., DiRuggiero, J. & Taylor, J. MetaWRAP—a flexible pipeline for genome-resolved metagenomic data analysis. Microbiome 6, 158 (2018).
Chaumeil, P.-A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. G. T. D. B.- Tk: a toolkit to classify genomes with the genome taxonomy database. Bioinformatics 36, 1925–1927 (2020).
Parks, D. H. et al. GTDB: an ongoing census of bacterial and archaeal diversity through a phylogenetically consistent, rank normalized and complete genome-based taxonomy. Nucleic Acids Res. 50, D785–D794 (2022).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinforma. 11, 119 (2010).
Aramaki, T. et al. KofamKOALA: KEGG ortholog assignment based on profile HMM and adaptive score threshold. Bioinformatics 36, 2251–2252 (2020).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51, D587–D592 (2023).
Liu, B., Zheng, D., Jin, Q., Chen, L. & Yang, J. VFDB 2019: a comparative pathogenomic platform with an interactive web interface. Nucleic Acids Res. 47, D687–D692 (2019).
Brooks, B. et al. Strain-resolved analysis of hospital rooms and infants reveals overlap between the human and room microbiome. Nat. Commun. 8, 1814 (2017).
Olm, M. R. et al. inStrain profiles population microdiversity from metagenomic data and sensitively detects shared microbial strains. Nat. Biotechnol. 39, 727–736 (2021).
Asnicar, F. et al. Precise phylogenetic analysis of microbial isolates and genomes from metagenomes using PhyloPhlAn 3.0. Nat. Commun. 11, 2500 (2020).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Yu, G., Smith, D. K., Zhu, H., Guan, Y. & Lam, T. T.-Y. ggtree: an r package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol. Evol. 8, 28–36 (2017).
Croucher, N. J. et al. Rapid phylogenetic analysis of large samples of recombinant bacterial whole genome sequences using Gubbins. Nucleic Acids Res. 43, e15 (2015).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993 (2011).
Cantalapiedra, C. P., Hernández-Plaza, A., Letunic, I., Bork, P. & Huerta-Cepas, J. eggNOG-mapper v2: functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol. Biol. Evol. 38, 5825–5829 (2021).
Huerta-Cepas, J. et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 47, D309–D314 (2019).
Hadfield, J. et al. Phandango: an interactive viewer for bacterial population genomics. Bioinformatics 34, 292–293 (2018).
Zulkower, V. & Rosser, S. DNA features viewer: a sequence annotation formatting and plotting library for python. Bioinformatics 36, 4350–4352 (2020).
Johansson, M. H. K. et al. Detection of mobile genetic elements associated with antibiotic resistance in Salmonella enterica using a newly developed web tool: MobileElementFinder. J. Antimicrob. Chemother. 76, 101–109 (2021).
Wang, M. et al. VRprofile2: detection of antibiotic resistance-associated mobilome in bacterial pathogens. Nucleic Acids Res. 50, W768–W773 (2022).
Schmartz, G. P. et al. PLSDB: advancing a comprehensive database of bacterial plasmids. Nucleic Acids Res. 50, D273–D278 (2022).
Ondov, B. D. et al. Mash: fast genome and metagenome distance estimation using MinHash. Genome Biol. 17, 132 (2016).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Jain, C., Rodriguez-R, L. M., Phillippy, A. M., Konstantinidis, K. T. & Aluru, S. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat. Commun. 9, 5114 (2018).
Darling, A. C. E., Mau, B., Blattner, F. R. & Perna, N. T. Mauve: Multiple alignment of conserved genomic sequence with rearrangements. Genome Res. 14, 1394–1403 (2004).
Wintersinger, J. A. & Wasmuth, J. D. Kablammo: an interactive, web-based BLAST results visualizer. Bioinformatics 31, 1305–1306 (2015).
Guy, L., Roat Kultima, J. & Andersson, S. G. E. genoPlotR: comparative gene and genome visualization in R. Bioinformatics 26, 2334–2335 (2010).
Cheng, M. et al. Deep longitudinal lower respiratory tract microbiome profiling reveals genome-resolved functional and evolutionary dynamics in critical illness, github repository. Zenodo https://doi.org/10.5281/zenodo.13627162 (2024).
Acknowledgements
We thank all patients and clinicians who participated in this study, all our collaborators and their staff for their invaluable contributions to this work, and Kaiyue Tang for her suggestions on using Chelex100. We are grateful to the NECHO computing cluster at the core facility of the Life Sciences Institute. This work was supported by NSFC 82173645 and 82341109, the Fundamental Research Funds for the Central Universities, Capital’s Funds for Health Improvement and Research (No. 2022-2G-40911), and the special fund of the Beijing Clinical Key Specialty Construction Program, P. R. China (2021).
Author information
Authors and Affiliations
Contributions
C.J. conceived the study, secured funding, and provided overall supervision. Y.L. and C.J. designed the strategy to collect clinical data and samples. Y.L., H.L., J.X, and M.Z. supervised each center’s sample collection and clinical data gathering. C.L., Y.X., X.C., Y.C., L.X., J.X., and Y.Z. collected clinical samples and data. M.C. performed method development and sample processing for sequencing. M.C. and X.W. performed statistical and bioinformatics analyses. M.C. and C.J. drafted the paper. L.H., W.Z., H.L., and Y.L. provided suggestions to revise the manuscript. All authors have critically reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Leopoldo Segal, and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cheng, M., Xu, Y., Cui, X. et al. Deep longitudinal lower respiratory tract microbiome profiling reveals genome-resolved functional and evolutionary dynamics in critical illness. Nat Commun 15, 8361 (2024). https://doi.org/10.1038/s41467-024-52713-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-52713-8
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.