Abstract
Arboviruses can be paternally transmitted by male insects to offspring for long-term persistence, but the mechanism remains largely unknown. Here, we use a model system of a destructive rice reovirus and its leafhopper vector to find that insect ribosome-rescuer Pelo-Hbs1 complex expressed on the sperm surface mediates paternal arbovirus transmission. This occurs through targeting virus-containing tubules constituted by viral nonstructural protein Pns11 to sperm surface via Pns11-Pelo interaction. Tubule assembly is dependent on Hsp70 activity, while Pelo-Hbs1 complex inhibits tubule assembly via suppressing Hsp70 activity. However, virus-activated ubiquitin ligase E3 mediates Pelo ubiquitinated degradation, synergistically causing Hbs1 degradation. Importantly, Pns11 effectively competes with Pelo for binding to E3, thus antagonizing E3-mediated Pelo-Hbs1 degradation. These processes cause a slight reduction of Pelo-Hbs1 complex in infected testes, promoting effective tubule assembly. Our findings provide insight into how insect sperm-specific Pelo-Hbs1 complex is modulated to promote paternal virus transmission without disrupting sperm function.
Similar content being viewed by others
Introduction
Arthropod-borne viruses (arboviruses), which pose significant global health and agricultural challenges, can be vertically transmitted to the offspring of insects, ensuring the long-term persistence of the viruses in nature1,2. Apart from maternal transmission through female insects, arboviruses also can be paternally transmitted through male insects1,2. For example, the two flaviviruses, La Crosse virus and Zika virus, and the plant reovirus, rice gall dwarf virus (RGDV), are paternally transmitted by infected male mosquitoes and leafhoppers, respectively3,4,5. Transovarial transmission between an infected female and its offspring has been extensively explored6,7,8,9,10,11,12. However, to date, the mechanisms involved in sperm-mediated paternal transmission remain largely unknown.
We previously discovered that RGDV and an insect-specific virga virus in rice green leafhopper (Recilia dorsalis) cooperatively hijack the surface proteins of insect sperms for paternal transmission without significantly disturbing sperm functions5,13. Currently, the mechanisms of insect sperm-specific protein-mediated paternal arbovirus transmission are not fully understood. The serpin protein HongrES1 in mammals is specifically deposited on the sperm surfaces and is involved in sperm maturation14,15. We have shown that the interaction of viral capsid proteins of RGDV and a virga virus with the sperm-specific HongrES1 synergistically facilitates paternal transmission of both viruses13. Pelo (Protein pelota homolog) forms a complex with GTPase Hbs1 (Hsp70 subfamily B suppressor1) to rescue the stalled ribosomes in the terminal step of protein synthesis, and thus Pelo-Hbs1 complex is a well-known surveillance factor in ribosome-associated quality control machinery16,17. Pelo is an evolutionarily conserved gene found in diverse species from archaebacteria to humans18,19 and contains an RNA-binding domain similar to eukaryotic release factor I (eRF1)20,21,22. Hbs1 is originally identified for its ability to rescue the stalled ribosomes by suppressing Hsp70 activity23,24,25. In mammals, the Pelo-Hbs1 complex is crucial for recognising and rescuing the stalled ribosomes during protein synthesis23,24,25. In Drosophila, the Pelo-Hbs1 complex is essential for spermatogenesis and germline stem cell maintenance16,26. Mutant Drosophila lacking Pelo or Hbs1 is male-sterile due to the defect in meiosis and spermatid individualisation16,26and shows resistance to Drosophila C virus (DCV), cricket paralysis virus, Drosophila X virus, and invertebrate iridescent virus27. The presence of bacterial endosymbiont Wolbachia in mosquitoes causes the reduced accumulation of Pelo, which contributes to the inhibition of dengue virus replication28. Furthermore, miRNA Bta-miR-2411 induces the downregulation of Pelo mRNA in Madin-Darby bovine kidney cells, restricting the replication of bovine viral diarrhoea virus29. Potentially, the absence of Pelo may impair the recycling of stalled ribosomes, which reduces the availability of free ribosomes and inhibits viral protein production, thus restricting viral replication. However, whether Pelo-Hbs1 complex functions as a defender for insect vectors against virus infection remains to be determined. Importantly, how viruses modulate the expression of Pelo-Hbs1 complex to promote sperm-mediated paternal transmission also remains unknown.
RGDV is insect-borne plant reovirus transmitted by R. dorsalis in a persistent-propagative manner30,31. During the early stage of viral infection in insect vector cells, the nonstructural proteins Pns7, Pns9, and Pns12 of RGDV associate to form the initial viroplasm matrix, where viral replication and assembly of progeny virions occur32,33,34. Subsequently, tubular structures are constructed by the nonstructural protein Pns11 of RGDV package virions to facilitate efficient virus spread among insect vector cells35. The heat shock proteins function as molecular chaperones to ensure the proper folding and translocation of nascent membrane proteins36. Tubular proteins of plant reoviruses are retained by endoplasmic reticulum (ER) membrane proteins like DNAJB12 and BAP31 to the ER for protein synthesis and then transport to the cytosolic Hsp70 for tubule assembly37,38. Here, we identify that the Pelo-Hbs1 complex is highly expressed in the male reproductive organ of R. dorsalis, and is prominently present on the sperms. RGDV infection slightly reduces the accumulation of Pelo-Hbs1 complex in the male reproductive organ; however, Pns11-Pelo interaction mediates the spread of virus-containing Pns11 tubules for targeting the sperms. Pelo-Hbs1 complex suppresses Hsp70 activity to inhibit Pns11-induced tubule assembly; however, Pns11 antagonises ubiquitin ligase E3-mediated ubiquitinated degradation of Pelo, ensuring a slight reduction in the accumulation of Pelo-Hbs1 complex. These mechanisms result in a slightly reduced level of Pelo-Hbs1 complex in infected testes, ultimately aiding in effective tubule assembly. Our findings shed light on how insect sperm-specific Pelo-Hbs1 complex is regulated to support sperm-mediated paternal transmission of viruses without significantly affecting sperm function.
Results
Identification of leafhopper sperm-specific protein complex Pelo-Hbs1
In Drosophila, the Pelo-Hbs1 complex is essential for the progression of meiosis during spermatogenesis and germline stem cell maintenance16. R. dorsalis Pelo and Hbs1 displayed the conserved amino acid sequence homology with Drosophila and Aedes aegypti (Supplementary Figs. S1, 2). Yeast two-hybrid (Y2H) and glutathione S-transferase (GST) pull-down assays demonstrated the specific interaction of Pelo with Hbs1, thereby forming Pelo-Hbs1 complex in R. dorsalis (Fig. 1a, b). RT-qPCR and western blot assays showed that the Pelo-Hbs1 complex accumulated at higher levels in the male reproductive organ compared to other insect organs (Fig. 1c, d). Immunofluorescence microscopy demonstrated the co-localisation of Pelo and Hbs1 in the bundle of filamentous structures within the testes (Fig. 1e). Immunoelectron microscopy further confirmed that Pelo and Hbs1 antibodies specifically reacted with the sperms, particularly with the surfaces of the sperms within the sperm cavity in the male testes (Fig. 1f). Thus, we identified a leafhopper sperm-specific protein complex Pelo-Hbs1.
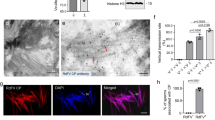
a Interaction between Pelo and Hbs1, as detected by Y2H assay. Transformants were plated on QDO. QDO, SD/-Trp-Leu-His-Ade. b Interaction between Pelo and Hbs1, as detected by GST Pull-down assay. Pelo-GST was incubated with glutathione-Sepharose beads and Hbs1-His was then added to the beads, followed by western blot assay to detect the Hbs1-His bound to Pelo-GST. c RT-qPCR assay showing the relative transcript levels of Pelo and Hbs1 in different organs. Means (± SD) represents three replicates. Different letters in the same column in c indicate a significant difference (P < 0.05, Tukey’s HSD multiple test). d The accumulation levels of Pelo and Hbs1 in different organs, as determined by western blot assay. The relative intensities of protein bands are shown below using ImageJ. H3 is used as the loading control. Data represent three replicates and each replicate contains 30 different organs. e Distribution of Pelo and Hbs1 in testes, as determined by immunofluorescence microscopy. The testes dissected from 30 males were immunostained with Pelo-FITC, Hbs1-FITC, or Hbs1-rhodamine. Bars, 10 μm. f Immunogold labelling of Pelo or Hbs1 in testes of nonviruliferous male R. dorsalis. Red arrows indicate gold particles labelled by Pelo or Hbs1 antibody on the surface of sperms. Bars, 100 nm. g, h The relative transcript (g) and accumulation (h) levels of Pelo and Hbs1 in testes treated with dsPelo or dsGFP. i, j The relative transcript (i) and accumulation (j) levels of Pelo and Hbs1 in testes treated with dsHbs1 or dsGFP. k, l The relative transcript (k) and accumulation (l) levels of Pelo and Hbs1 in testes of nonviruliferous and viruliferous leafhoppers, as determined by RT-qPCR and western blot assays. Means (± SD) in (g, i) and (k) represent three replicates and are analysed using a two-tailed t test. Relative protein levels in (h, j), and (l) are quantified using ImageJ with H3 as a reference protein, and the data present three replicates, with each replicate containing 30 testes. Wb, whole body; Sg, salivary gland; Mg, midgut; Ov. Ovary; Te, testis; S, sperm; V−, nonviruliferous; V+, viruliferous.
We then investigated the effects of the knockdown of Pelo or Hbs1 expression on sperm morphology and the fitness of male leafhoppers and their offspring. Initially, we microinjected adult male R. dorsalis individuals with the in vitro synthesised dsRNAs targeting Pelo, Hbs1, or GFP (dsPelo, dsHbs1, or dsGFP). The knockdown of Pelo expression did not significantly affect the Hbs1 transcript level but decreased the Hbs1 protein accumulation level in virus-free male reproductive organs (Fig. 1g, h). Similarly, the knockdown of Hbs1 expression did not significantly affect Pelo transcription but decreased Pelo protein accumulation in virus-free male reproductive organs (Fig. 1i, j). Immunofluorescence microscopy showed that Pelo- or Hbs1-decorated sperm bundles displayed abnormal morphology, appearing poorly and loosely arranged in dsPelo- or dsHbs1-treated testes (Supplementary Fig. S3a, b). Electron microscopy further demonstrated that the sperms in dsPelo-treated testes exhibited accelerated degradation, displaying a poor and vacuolated appearance (Supplementary Fig. S3c, d). Moreover, it was found that dsPelo- or dsHbs1-treated males showed no significant differences in mortality rates compared to dsGFP-treated control (Supplementary Fig. S3e). However, the eggs collected from the mating of dsPelo- or dsHbs1-treated males with females did not develop normally (Supplementary Fig. S3f). It was found that the egg number decreased, the mean egg development duration became longer, and the egg hatching rate was lower after dsPelo or dsHbs1 treatment (Supplementary Fig. S3g, h). Collectively, these results indicate that the leafhopper Pelo-Hbs1 complex potentially is involved in spermatogenesis, as described for Drosophila16, and thus, the reduced accumulation of Pelo-Hbs1 complex adversely affects sperm development and offspring fitness.
We then determined whether viral infection affected the expression of the Pelo-Hbs1 complex in the leafhopper male reproductive organ. RT-qPCR assay showed that RGDV infection did not significantly affect the transcript levels of Pelo or Hbs1 in the male reproductive organ (Fig. 1k). However, western blot assay showed that RGDV infection led to a slight reduction in the accumulation of Pelo and Hbs1 in virus-infected male reproductive system compared to virus-free control (Fig. 1l). Thus, viral infection negatively influences the accumulation level of Pelo-Hbs1 complex in leafhopper male reproductive organ.
Direct interaction of Pelo with RGDV Pns11 mediates the spread of virus-induced tubules to target sperms
We then investigated whether the Pelo-Hbs1 complex facilitated sperm-mediated paternal virus transmission. Electron microscopy showed the association of RGDV icosahedral virions or virus-containing tubules with the plasma membranes of sperms in RGDV-infected testes (Fig. 2a–d). It appeared that virus-containing tubules were released from the cytoplasm of the testis epithelium into the sperm lumen via the apical plasmalemma, and finally targeted the sperm surfaces (Fig. 2b–d). Notably, this association of virus-containing tubules with the sperms did not affect sperm morphology (Fig. 2c, d). Immunoelectron microscopy confirmed that the Pns11 antibody specifically reacted with virus-containing tubules (Fig. 2b–d). Thus, RGDV Pns11-induced tubules potentially mediate viral spread from the testis epithelium into the sperm lumen by overcoming the member barrier of apical plasmalemma, consistent with previous findings that Pns11-induced tubules could facilitate viral spread among insect vector cells35. Immunofluorescence and immunoelectron microscopy also showed the presence of Pns11 tubule-associated sperms in the dissected spermatheca of uninfected females five days after mating with infected males (Supplementary Fig. S4), confirming the transfer of tubule-associated sperms from infected males to females. Thus, RGDV hijacks virus-induced tubules to target the sperms, which subsequently spread to the female spermatheca during mating, ultimately passing on to the offspring. This proves tubule-mediated paternal transmission of RGDV by R. dorsalis.
a–d Electron microscopy showing the sperms associated with RGDV virions or virus-containing tubular structures in virus-infected testes., Testes in (b–d) were immunolabelled with Pns11-specific IgG as the primary antibody, followed by treatment with 15- nm gold particle-conjugated IgG as the secondary antibody. Panel (b–i) shows the enlarged image of the boxed area in panel (b). Red arrows indicate gold particles. Bars, 100 nm. e–h Immunoelectron microscopy showing the association of Pelo (e, f) or Hbs1 (g, h) with virus-induced tubules and sperms. Testes were immunolabelled with Pelo- or Hbs1-specific IgG as the primary antibody and treated with 15- nm gold particle-conjugated IgG as the secondary antibody. Red arrows indicate gold particles. Bars, 100 nm. i–m Immunofluorescence microscopy showing the association of RGDV Pns11 with Pelo or Hbs1 in infected male vas deferens (i–k) and testes (l and m). Dissected male reproductive organs from virus-free or positive males were immunostained with Pelo-FITC, Hbs1-FITC, or Pns11-rhodamine. Panels (j) and (k) are the enlarged images of the boxed areas in panel (i). Arrows in (j) and (k) indicate the colocalization of Pns11 and Pelo to the puncta structures. Ec, epithelial cells; Te, testes; Vd, vas deferens; S, sperm; AP, apical plasmalemma; Vi, virions; Ts, tubular structure. V−, nonviruliferous; V+, viruliferous. Bars, 10 μm. n Y2H assay showing the interactions between Pelo and Pns11 or P8, and between Hbs1 and Pns11 or P8. Transformants were plated on QDO. QDO, SD/-Trp-Leu-His-Ade medium. o Interaction between Pelo and Pns11 was detected by GST Pull-down assay. Pns11-GST was incubated with glutathione-Sepharose beads, and Pelo-His was then added to the beads, followed by a western blot assay to detect the Pelo-His bound to Pns11-GST.
Immunoelectron microscopy showed that Pelo and Hbs1 antibodies specifically reacted with the sperms and virus-containing Pns11 tubules, which were distributed in the cytoplasm of testis epithelium, in close proximity to the sperms, and even on the sperm surfaces (Fig. 2e–h). Immunofluorescence microscopy showed that Pns11 formed obvious punctate and filamentous structures, which were extensively distributed in the epithelial cells of the testes or vas deferens (Fig. 2i–m). Some Pns11 tubules were observed to colocalize with Pelo in the epithelial cells and appeared to migrate towards the sperm lumen to target Pelo-Hbs1 complex-decorated sperms (Fig. 2i–m). Obviously, the viral infection did not significantly alter the localization or accumulation of Pelo-Hbs1 complex on sperms during viral infection, confirming that RGDV infection just slightly decreased the accumulation of Pelo-Hbs1 complex in the male reproductive organ, as revealed by western blot assay (Fig. 1l). Y2H assay showed that Pns11, but not P8, specifically interacted with Pelo, and neither P8 nor Pns11 interacted with Hbs1 (Fig. 2n). GST pull-down assay confirmed that Pelo specifically interacted with Pns11 in vitro (Fig. 2o). Potentially, the interaction between Pns11 and Pelo mediates the spread of virus-induced tubules to target the sperms.
Pelo-Hbs1 complex promotes Pns11 tubule-mediated RGDV paternal transmission
We then investigated how Pns11 tubules hijacked the Pelo-Hbs1 complex for paternal RGDV transmission. In the neutralising Pns11-sperm binding experiment, in vitro binding of purified Pns11 proteins to live sperms was observed after a 1 hour (h) incubation, and the efficiency for sperm-Pns11 binding was decreased by ~ 55% following pretreatment with Pelo antibody (Fig. 3a, b). This suggests that Pelo mediates the direct binding of Pns11 tubules to the sperms. Subsequently, we microinjected Pelo antibody or pre-immune antibody into RGDV-infected male adult leafhoppers and mated them with uninfected virgin female leafhoppers for two days, and observed ~ 34% or 85% positive RGDV infection rates of the offspring, respectively (Fig. 3c). Thus, Pelo antibody effectively interferences with Pelo-Pns11 interaction, thereby suppressing efficient parental transmission of RGDV in R. dorsalis population.
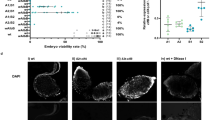
a Reduced Pns11-sperm binding by pre-treatment with Pelo antibody. Live sperms dissected from nonviruliferous testes were incubated with purified Pns11 proteins and pre-immune antibody or Pelo antibody, then stained with Pns11-rhodamine (red) and DAPI (blue). Bars, 10 μm. b The number of sperms associated with Pns11 was calculated as 100 sperms. c Paternal transmission rates of RGDV determined by mating nonviruliferous females with Pelo or pre-immune antibody-treated viruliferous males. Ten pairs of mating combinations were performed for three biological replicates. d, e The effects of dsPelo on the expression of Pelo, Hbs1, Pns11 and P8 in viruliferous males, as determined by RT-qPCR and western blot assays. f, g The effects of dsHbs1 treatment on the expression of Hbs1, Pelo, Pns11 and P8 in viruliferous males, as determined by RT-qPCR and western blot assays. Relative protein levels in (e) and (g) are quantified using ImageJ with H3 as a reference protein, and the data represent three replicates, with each replicate containing 30 testes. h The effects of dsPelo or dsHbs1 on the expression of Pelo, Hbs1, and Pns11 in viruliferous males, as determined by immunofluorescence microscopy. The dissected testes were immunostained with Pelo-Alexa Fluor 647 (blue), Hbs1-FITC (green), and Pns11-rhodamine (red). The right three panels are the enlarged images of the boxed areas in the left panels. Bars, 10 μm. i The mean percentages of sperms (n = 100) associated with Pns11 in testes. j Proposed mechanism of Pelo-Hbs1 complex-mediating paternal transmission of RGDV in testis. Pns11-Pelo interaction mediates the spread of virus-induced tubules from the testes epithelium into the sperm lumen for targeting sperm surfaces. Means (± SD) in (b, c, d, f), and (i) represent three replicates and are analysed using a two-tailed t test. Te testes, AP apical plasmalemma, BL basal lamina, EC epithelial cell, N nucleus, S sperm.
We next knocked down Pelo or Hbs1 expression by microinjecting males with synthesised dsPelo or dsHbs1. The knockdown of Pelo expression effectively decreased the accumulation of Hbs1, P8, and Pns11 in virus-infected male reproductive organs (Fig. 3d, e). Similarly, the knockdown of Hbs1 expression effectively decreased the accumulation of Pelo, P8, and Pns11 in virus-infected male reproductive organs (Fig. 3f, g). Immunofluorescence microscopy confirmed that dsPelo and dsHbs1 treatment significantly decreased the accumulation of Pelo-Hbs1 complex and Pns11 structures on the sperms (Fig. 3h, i). It appears that the less Pelo-Hbs1 accumulates on the sperms, causing the less Pns11 structures to be targeted on the sperms. Potentially, the Pelo-Hbs1 complex directly binds to and mediates the spread of virus-induced tubules from the testes epithelium into the sperm lumen to target the sperms, thereby facilitating paternal virus transmission (Fig. 3j).
Pelo-Hbs1 complex suppresses Hsp70 activity to inhibit Pns11-induced tubule assembly
The expression of Pns11 of RGDV alone induced the formation of tubules in Spodoptera frugiperda (Sf9) cells35. Immunofluorescence microscopy revealed that Pns11 was initially distributed in the cytoplasm at 24 h post-infection (hpi) and then aggregated to form tubular structures at 36 hpi (Fig. 4a). Subsequently, Sf9 cells were singly infected or co-infected with recombinant baculoviruses containing Pns11, Pelo, or Hbs1. When expressed alone, Pelo and Hbs1 appeared to be distributed in the cytosol (Fig. 4b). Co-expression of Pns11 with Pelo or Hbs1 resulted in the formation of tubular structures at 48 hpi (Fig. 4c, d). Interestingly, in Sf9 cells triply expressed with Pns11, Pelo, and Hbs1, Pns11 initially formed tubular structures at 60 hpi (Fig. 4e). Thus, overexpression of Pelo-Hbs1 complex inhibited the assembly of Pns11 tubules in Sf9 cells.
a Sf9 cells singly expressing Pns11 were immunolabelled with Pns11-rhodamine. b Sf9 cells expressing Pelo-Flag or Hbs1-His was immunolabelled with Flag-FITC or His-FITC. c, d Sf9 cells co-expressing Pns11-Myc and Pelo-Flag or Hbs1-His were immunolabelled with Myc-rhodamine, Flag- FITC, or His-FITC. e Sf9 cells triply expressed with Pns11-Myc, Pelo-Flag and Hbs1-His were immunolabelled with Myc-rhodamine, Flag-Alexa Fluor 647, or His-FITC. f After incubation with VER155008 or DMSO, Sf9 cells expressing Pns11-Myc were immunolabelled with Myc-rhodamine. g The mean percentages of tubule-forming or non-tubule-forming Sf9 cells after treatment with VER155008 or DMSO were counted for 100 cells. Means (± SD) represent three replicates. h Sf9 cells expressing Hsp70-His were fixed at 48 hpi and immunolabelled with His-FITC. i Sf9 cells co-expressing Pns11-Myc and Hsp70-His were fixed at 18 or 24 hpi and immunolabelled with Myc-rhodamine and His-FITC. j Y2H assay showing the interaction between Hsp70 and Pns11. Transformants were plated on QDO. QDO, SD/-Trp-Leu-His-Ade medium. k Pull-down assay showing the interaction between Hsp70 and RGDV Pns11. The proteins were detected by GST or His antibodies. l Sf9 cells co-expressing Hsp70-His and Hbs1-Flag were collected at 48 hpi for western blot assay with His and Flag antibodies. Relative protein levels are quantified using ImageJ with GAPDH as a reference protein, and the data represent three replicates. m Sf9 cells triply expressing Pns11-Myc, Hsp70-His, and Hbs1-Flag were immunolabelled with Myc-rhodamine, Flag-Alexa Fluor 647, or His-FITC. n RT-qPCR and western blot assay showing the expression of Hsp70 in nonviruliferous or viruliferous male testes. V− nonviruliferous, V+ viruliferous. o Western blot assay shows the accumulation levels of Hbs1, Hsp70, and Pns11 in RGDV-infected insects after microinjection of purified Hbs1 or GFP proteins. Means (± SD) in (g) and (n) represent three replicates and are analysed using a two-tailed t test. Relative protein levels in (n) and (o) are quantified using ImageJ with H3 as a reference protein, and the data represent three replicates, with each replicate containing 30 testes. Bars (a–f, h, i, and m), 10 μm.
The assembly of virus-containing tubules induced by plant reoviruses is dependent on the ATPase activity of the Hsp70 chaperone37. To examine whether the assembly of Pns11 tubules is dependent on the ATPase activity of Hsp70, an allosteric inhibitor, VER155008, which competes with ATP for Hsp70 binding39, was used to inhibit the folding function of Hsp70 in Sf9 cells. Treatment with 20 μM VER155008 displayed low toxicity to Sf9 cells but effectively inhibited Pns11 tubule formation (Fig. 4f, g). Immunofluorescence microscopy showed that Pns11 was co-localised to Hsp70 in the cytoplasm at 18 hpi, and initially formed tubular structures at 24 hpi in Sf9 cells co-expressed with Hsp70 and Pns11, suggesting that Hsp70 promotes Pns11 tubule formation (Fig. 4h, i). Hence, overexpression of Hsp70 enhances the assembly of Pns11 tubules. Additionally, Y2H and GST pull-down assays verified the interaction between Hsp70 and Pns11 (Fig. 4j, k). Therefore, the capacity of Pns11 tubules to assemble and fold correctly is dependent on the ATPase activity of Hsp70.
Hbs1 was originally identified for its ability to rescue the stalled ribosomes by suppressing Hsp70 activity23. Co-expression of Hbs1 and Hsp70 in Sf9 cells led to the reduced expression of Hsp70, confirming that Hbs1 functions as a suppressor of the Hsp70 subfamily (Fig. 4l). In Sf9 cells triply expressed with Pns11, Hbs1, and Hsp70, Pns11 initially formed tubular structures at 36 hpi (Fig. 4m), suggesting that Hbs1 neutralises the ability for Hsp70 to mediate Pns11 tubules assembly. These findings reveal that Hbs1 delays the formation of Pns11 tubules by inhibiting the ATPase activity of Hsp70 and that Pelo-Hbs1 complex acts as a suppressor of virus-induced tubule assembly by inhibiting Hsp70 activity.
RT-qPCR assay showed that RGDV infection significantly increased the expression of Hsp70 in the male reproductive organ (Fig. 4n). Microinjection of purified Hbs1 proteins decreased Hsp70 expression, resulting in the reduced accumulation of Pns11 in the male reproductive organs (Fig. 4o). Thus, Pelo-Hbs1 complex serves as a viral restriction factor by suppressing ATPase activity of Hsp70, ultimately inhibiting Pns11-induced tubule assembly. However, viral infection causes a slight decrease in the accumulation of the Pelo-Hbs1 complex in the male reproductive organs, thus antagonising its viral restriction function.
Identification of the leafhopper sperm protein ubiquitin ligase E3 that interacts with and negatively regulates Pelo expression
RGDV infection did not significantly influence the transcript levels of Pelo and Hbs1 but slightly decreased Pelo and Hbs1 protein accumulation levels (Fig. 1k, l). Furthermore, the knockdown of Pelo or Hbs1 expression caused the reduced accumulation of viral proteins (Fig. 3d–g). Conversely, RT-qPCR and western blot assays showed that the knockdown of Pelo expression did not significantly change the transcript and protein accumulation levels of β-actin and β-tubulin of R. dorsalis in virus-infected male reproductive organs (Supplementary Fig. S5). It appears that RGDV infection may specifically trigger a post-translational degradation of Pelo and Hbs1. To further verify the proteasomal degradation of Pelo and Hbs1, viruliferous male R. dorsalis adults were microinjected with the proteasome inhibitor MG132 (10 μm), and the male reproductive organs were dissected and collected a day later. Western blot assay showed that the protein accumulation levels of Pelo and Hbs1 were higher in the MG132 treated group compared to the DMSO control (Fig. 5a), suggesting that Pelo and Hbs1 may undergo the ubiquitinated degradation in the male reproductive organ.
a The accumulation levels of Pelo, Hbs1 and Pns11 in the testes of viruliferous male adults treated with MG132, as determined by western blot assay. b Y2H assays showing the interaction relationship for E3, Pelo, Hbs1, P8 or Pns11. Transformants were plated on QDO. QDO, SD/-Trp-Leu-His-Ade medium. c Pull-down assay showing the interaction between E3 and Pelo. Proteins were detected using GST or His antibody. d, e RT-qPCR and western blot assays showing the relative transcript (d) and accumulation (e) levels of E3 in different leafhopper organs. Means (± SD) in d represent three biological replicates. Different letters indicate a significant difference (P < 0.05, Tukey’s HSD multiple test). f, g The relative transcript (f) and accumulation (g) levels of E3 in nonviruliferous or viruliferous leafhopper testes, as determined by RT-qPCR and western blot assays. h, i Distribution of E3, Pelo, or Pns11 in the testes of nonviruliferous or viruliferous leafhoppers, as determined by immunofluorescence microscopy. The testes were immunolabelled with E3-FITC, E3-rhodamine, Pelo-FITC, Pelo-Alexa Fluor 647, or Pns11-rhodamine. Bars, 10 μm. j, k Localization of E3 on the sperms in the testes from nonviruliferous (j) or viruliferous (k) males, as determined by immunoelectron microscopy. Leafhopper testes were immunolabelled with E3-specific IgG as the primary antibody, followed by treatment with 15- nm gold particle-conjugated IgG as the secondary antibody. Red arrows indicate gold particles. Bars, 100 nm. l, m RT-qPCR and western blot assays showing the relative transcript and accumulation levels of E3, Pelo and Hbs1 in the testes from nonviruliferous (l) or viruliferous (m) leafhoppers treated with dsE3 or dsGFP. Relative protein levels in (a, e, g, l), and (m) are quantified using ImageJ with H3 as a reference protein, and the data present three replicates, with each replicate containing 30 testes. Means (± SD) in (f, l), and (m) represent three replicates and are analysed using a two-tailed t test. Wb whole body, Sg salivary gland, Mg midgut, Ov ovary, S sperm, Te testis, Ts tubular structures, V− nonviruliferous, V+ viruliferous.
The ubiquitination-26S proteasome system is a key pathway for protein degradation, involving a cascade reaction catalysed by specific enzymes: ubiquitin-activating enzyme (E1), ubiquitin-binding enzyme (E2), and ubiquitin ligase (E3), with the substrate primarily dictated by ubiquitin ligase40,41,42. Y2H and GST pull-down assays showed that Pelo interacted with an E3 ubiquitin ligase, encoding a 658 amino acid protein that contained a C3HC4-type RING domain at its N-terminus (Fig. 5b, c and Supplementary Fig. S6). In contrast, Hbs1 did not interact with this E3 (Fig. 5b), suggesting that E3 potentially mediates the ubiquitinated degradation of Pelo but not Hbs1. RT-qPCR and western blot assays showed that E3 was highly expressed in the male reproductive organs (Fig. 5d, e), while RGDV infection further activated E3 expression (Fig. 5f, g). It appears that virus-induced elevation of E3 levels promotes the ubiquitinated degradation of Pelo.
Immunofluorescence and immunoelectron microscopy showed that E3 was co-localised with Pelo on the sperms from nonviruliferous insects (Fig. 5h, j). Furthermore, the Y2H assay showed that E3 did not interact with Pns11 or P8 of RGDV (Fig. 5b). However, Immunofluorescence and immunoelectron microscopy showed that E3 antibody reacted with virus-containing tubules in the male reproductive organs (Fig. 5i, k). The knockdown of E3 expression by microinjection of the synthesised dsRNAs targeting E3 (dsE3) into the male adults increased Pelo and Hbs1 expression both in nonviruliferous and viruliferous insects, resulting in the reduced expression of viral proteins P8 and Pns11 in the male reproductive organs (Fig. 5l, m). Thus, virus-activated E3 promotes viral infection in vector male reproductive organs, potentially via negative regulation of the Pelo-Hbs1 complex.
RGDV Pns11 antagonises E3-mediated ubiquitinated degradation of Pelo
We then investigated whether E3 mediated Pelo ubiquitinated degradation. When Sf9 cells co-expressing E3 and Pelo, we observed a gradual decrease in Pelo accumulation levels with the increasing E3 expression levels (Fig. 6a), indicating a negative correlation between E3 and Pelo. However, Pelo expression levels showed a noticeable recovery trend after treatment with the proteasome inhibitor MG132 (Fig. 6b), suggesting that E3 induces Pelo degradation via the ubiquitin-proteasome pathway. Furthermore, the co-immunoprecipitation (Co-IP) assay revealed a significant increase in Pelo ubiquitination levels in Sf9 cells co-expressing E3, Pelo, and ubiquitin (Fig. 6c), suggesting that E3 plays a critical role in Pelo ubiquitination. In the presence of E1, E2, E3, and ubiquitin, we detected the ubiquitinated form of Pelo using a Flag-tag or ubiquitin antibody (Fig. 6d), confirming that Pelo is a substrate of E3 for degradation via the 26 S proteasome pathway.
a The accumulation levels of Pelo-Flag in Sf9 cells co-expressed with Pelo-Flag and E3-Myc, as determined by western blot assay. b Western blot assay showing the effects of MG132 on the accumulation of Pelo in Sf9 cells singly or co-expressing Pelo-Flag and E3-Myc. The proteins in (a) and (b) were detected using Flag, Myc, or GAPDH antibody. c Co-IP assay showing the ubiquitination of Pelo-Flag by E3-Myc in Sf9 cells co-expressing Pelo-Flag, E3-Myc, or ubiquitin-WT-HA. The IP proteins were detected by HA antibody, and the cell lysate was detected by Flag, Myc, or GAPDH antibody. TCL, total cell lysate. d In vitro ubiquitination assay showing the ubiquitination of Pelo-Flag by E3-HA, as determined by western blot assay using ubiquitin or Flag antibody. e Co-IP assay showing the ubiquitination of Pelo by E3 in Sf9 cells co-expressing Pelo-Flag, E3-Myc, ubiquitin-HA, or ubiquitin-K48-HA. f Co-IP assay showing the ubiquitination of Pelo by E3 in Sf9 cells co-expressing Pelo-Flag, E3-Myc, Pns11-His, ubiquitin-WT-HA or ubiquitin-K48-HA. The IP proteins in (e) and (f) were detected by HA antibody, and the cell lysate was detected by Flag, Myc, His, or GAPDH antibody. TCL, total cell lysate. g, h The competitive interactions between Pns11-Pelo and E3-Pelo were detected by pull-down assay. His-Pns11 and GST-Pelo were incubated with Glutathione-Sepharose agarose beads, then His-E3 was added to the beads; when increased the amounts of Pelo-E3, the binding between was not affected (g). His-E3 and GST-Pelo were incubated with Glutathione-Sepharose agarose beads, then His-Pns11 was added to the beads; when increased the amounts of Pelo-Pns11, the binding between was decreased (h). The proteins were detected using E3, Pns11, or GST antibody. i Y2H assay showing the interaction between Pelo-N, Pelo-M, Pelo-C and E3-N, E3-C or Pns11. Transformants were plated on QDO. QDO, SD/-Trp-Leu-His-Ade medium. Relative protein levels in (a, b, g) and (h) are quantified using ImageJ, and the data represent three replicates. Ub ubiquitin.
Ubiquitin is a small protein with seven lysine sites (K6, K11, K27, K29, K33, K48, and K63), and K48-linked polyubiquitination mediates the degradation of proteins through the proteasome43,44. We sought to determine whether E3 could induce K48-linked polyubiquitination of Pelo. Co-IP assay showed that overexpression of E3 significantly enhanced the presence of K48-linked polyubiquitination bands on Pelo, consistent with wild type ubiquitin-linked polyubiquitination of Pelo in Sf9 cells (Fig. 6e), suggesting that E3 mediates K48-linked ubiquitination of Pelo. However, the Co-IP assay showed that co-expression of Pns11 notably decreased the accumulation of ubiquitin or ubiquitin-K48 polyubiquitinated chains on Pelo (Fig. 6f), suggesting that Pns11 potentially inhibits E3-mediated ubiquitinated degradation of Pelo.
We then explored the mechanism behind why Pns11 inhibited E3-mediated ubiquitinated degradation of Pelo. Since Pns11 is specifically bound to Pelo, we hypothesised that RGDV infection triggered the formation of the Pns11-Pelo complex, which competitively inhibited E3-Pelo interaction. We thus compared the binding affinities of Pns11-Pelo and E3-Pelo. When the amount of Pns11 remained constant, the binding of Pns11 remained steady with an increase in E3 (Fig. 6g); however, when the amount of E3 remained constant, the binding of E3 decreased with an increase in Pns11 (Fig. 6h), suggesting that the binding ability of Pns11 to Pelo is stronger than that of E3 to Pelo. Furthermore, we investigated the functional domains involved in the interactions among E3, Pelo, and Pns11. E3 was divided into two segments, E3-N (1–200 aa, the RING domain) and E3-C (201–650 aa). Pelo contained three similar conserved functional domains, eRF1-1, eRF1-2 and eRF1-3, and was divided into three segments, Pelo-N (1–130 aa, eRF1-1), Pelo-M (131–268 aa, eRF1-2), and Pelo-C (269–385 aa, eRF1-3) (Fig. 6i). Y2H assay showed that both E3-N and Pns11 interacted with Pelo-N (Fig. 6i), underscoring the crucial roles of the E3 RING domain and Pelo eRF1-1 domain. Overall, these results suggest that Pns11 potentially antagonises E3-mediated ubiquitinated degradation of Pelo by competitively inhibiting E3-Pelo interaction (Fig. 7). This may explain why viral infection only slightly decreases Pelo-Hbs1 complex accumulation in vector male reproductive organs.
Pns11-induced tubule assembly is dependent on Hsp70 activity; however, Hbs1 effectively inhibits Hsp70 activity. Virus-activated ubiquitin ligase E3 directly targets and mediates the ubiquitinated degradation of Pelo via the 26 S proteasome pathway, which also causes the degradation of Hbs1. Importantly, Pns11 effectively competes with Pelo to bind to E3, thus antagonising E3-mediated degradation of the Pelo-Hbs1 complex, ensuring a slightly reduced accumulation of Pelo-Hbs1 complex and promoting Pns11 tubule assembly.
Discussion
Currently, the exact mechanism of sperm-mediated paternal virus transmission by male insects to offspring in nature has been largely ignored. The role of insect sperm-specific components in mediating paternal virus transmission needs to be explored. In Drosophila, the Pelo-Hbs1 complex, a well-known ribosome rescuer in translational quality control, is essential for spermatogenesis and germline stem cell maintenance16,26. In this study, we determine that the Pelo-Hbs1 complex is highly expressed in the male reproductive organ of leafhopper R. dorsalis, and is abundantly distributed on the sperms. The knockdown of Pelo or Hbs1 expression in male leafhoppers significantly impairs sperm morphology and function, potentially disrupting meiosis and spermatid individualisation during spermatogenesis, as also described in Drosophila16. We thus identify a leafhopper sperm protein complex Pelo-Hbs1. We further provide insight into how leafhopper sperm protein complex Pelo-Hbs1 is modulated to promote sperm-mediated paternal transmission of viruses without significantly impairing sperm function.
Electron microscopic observations show that RGDV exploits Pns11 tubules to traverse the apical plasmalemma from the infected testis epithelium to the sperm lumen for targeting the sperms. Virus-induced tubules not only facilitate rapid viral dissemination but also aid in the evasion of insect immune defences45,46,47. Viral tubular protein Pns11 interacts directly with Pelo, allowing the Pelo-Hbs1 complex to mediate the dissemination of virus-induced tubules for targeting the sperms. Pre-treatment with Pelo antibody interferes with this interaction, significantly hindering the attachment of virus-induced tubules to live sperm and preventing subsequent tubule-mediated paternal transmission. Importantly, such tubule-sperm binding does not affect the sperm morphological and functional characteristics. Thus, the Pelo-Hbs1 complex serves as the receptor for virus-induced tubules to facilitate sperm-mediated paternal virus transmission (Fig. 3j).
Pelo is a known interacting partner of Hbs1, forming a complex to perform ribosomal rescue during translation termination48. We note a slight reduction in Pelo-Hbs1 protein levels in leafhopper testes during viral infection despite normal Pelo or Hbs1 transcript levels. The knockdown of Hbs1 expression by dsHbs1 treatment results in the reduced Pelo protein level but not its mRNA level. Similarly, the knockdown of Pelo expression by dsPelo treatment leads to the reduced Hbs1 protein level without altering its mRNA level, suggesting that the degradation of Pelo or Hbs1 proteins occurs in the absence of either interacting partner. This coordinated regulation at the post-translational level of the Pelo-Hbs1 complex has been extensively observed in mammals29. One explanation for this finding is that Pelo stability is disrupted in the absence of Hbs1, and the destabilisation of its tertiary structure leads to the premature degradation of the protein in vivo. In accordance with this hypothesis, treating infected leafhopper testes with a specific dose of proteasome inhibitor results in an increase in Pelo and Hbs1 levels, indicating that the proteolytic degradation pathway is responsible for the reduced regulation of Pelo level under viral infection. Thus, Pelo and its binding partner Hbs1 are targeted for degradation when they are in an unbound state during the direct binding of virus-induced tubules to Pelo-Hbs1 complex.
Typically, the RING domain of E3 ubiquitin ligase facilitates the attachment of K48-linked polyubiquitin chains to target proteins for subsequent proteasomal degradation49,50. Here, we show that RGDV infection activates the expression of a sperm-specific E3 ubiquitin ligase to promote viral propagation in leafhopper testes. The interaction between the RING domain of E3 ubiquitin ligase and Pelo on the sperms promotes E3-mediated ubiquitinated degradation of Pelo through the K48-linked polyubiquitination via the 26 S proteasome pathway, which also synergistically causes the degradation of its binding partner Hbs1 (Fig. 7). During viral propagation in the leafhopper testes, both Pns11 and the E3 RING domain interact with the eRF1-1 region of Pelo, suggesting that Pns11 competitively inhibits E3-Pelo interaction. We further show that RGDV activates the Pns11-Pelo complex to compete with E3 for binding to Pelo, thereby suppressing the E3-mediated ubiquitinated degradation of the Pelo-Hbs1 complex. This process ultimately ensures the slight reduction of the Pelo-Hbs1 complex during viral infection. We have shown that RGDV infection activates the expression of leafhopper sperm-specific HongrES1, which is essential for sperm maturation in the male reproductive organ and contributes to paternal virus transmission13. Collectively, the adverse impact on sperm function resulting from a virus-mediated slight reduction in Pelo-Hbs1 complex accumulation could be alleviated by virus-activated HongrES1, thereby ensuring effective paternal virus transmission.
Hbs1 was originally identified for its ability to rescue the stalled ribosomes by suppressing Hsp70 activity23, while the proper folding and assembly of plant reovirus-induced tubules is dependent on the ATPase activity of Hsp7037. We confirm that the leafhopper Pelo-Hbs1 complex functions as a suppressor of virus-induced tubule assembly by inhibiting Hsp70 activity (Fig. 7). However, virus-induced slight reduction in Pelo-Hbs1 complex ultimately leads to an increase in Hsp70 expression in virus-infected leafhopper testes (Fig. 7). These processes promote tubule-mediated paternal virus transmission without significantly impairing sperm function.
Insect ribosome-rescuer Pelo-Hbs1 complex was previously identified as the factor involved in spermatogenesis in Drosophila51,52. We further identify it as a leafhopper sperm protein complex to facilitate paternal virus transmission. Furthermore, a lack of Pelo protein in Drosophila reduces the availability of free ribosomes and restricts the high-level synthesis of viral proteins, thus impeding viral replication27. Similarly, the Pelo-Hbs1 complex also negatively regulates the infection of plant potyviruses in Arabidopsis and Nicotiana benthamiana leaves, potentially through the recognition of Pelo with a functional G1-2A6-7 motif in the P3 cistron of potyviruses53. However, the exact role of the Pelo-Hbs1 complex still remains poorly understood. In mammals, it has recently been found that Pelo catalysers the oligomeric assembly of NOD-like receptor family proteins via activating their ATPase enzymatic activity54. It would be a topic in the future to understand the more evolutionary conserved functions of the Pelo-Hbs1 complex during persistent transmission of arboviruses by insect vectors in nature.
Methods
Insects, viruses, cells and antibodies
Nonviruliferous R. dorsalis individuals were reared in a greenhouse at 25 ± 1 °C with 75 ± 5% relative humidity and a 16- h light/8-h dark. RGDV-infected rice plants were collected from Luoding City, Guangdong Province, China and propagated via transmission by R. dorsalis. Briefly, nonviruliferous R. dorsalis individuals were allowed to feed on RGDV-infected rice plants for two days and then transferred to rice seedlings. The offspring produced by RGDV-positive parents were reared to establish a RGDV-positive population. The Sf9 cell line was cultured in Sf900 III medium (Gbico, 12658019) with 5% foetal bovine serum at 28 °C. Rabbit polyclonal antibodies against Pns11 and P8 of RGDV, as well as Pelo and Hbs1 of R. dorsalis, were prepared as described previously35,55. E3 antibody was produced by Genscript Biotech Corporation, Nanjing, China. Pelo and Hbs1-specific IgGs were conjugated to fluorescein isothiocyanate (FITC) (Invitrogen, F1907) to generate Pelo-FITC and Hbs1-FITC. Hbs1, Pns11, and E3-specific IgGs were conjugated to rhodamine (Invitrogen, 46112) to generate Hbs1-rhodamine, Pns11-rhodamine, and E3-rhodamine. Pelo-specific IgG was conjugated to Alexa Fluor 647 (Invitrogen, A37573) to generate Pelo-Alexa Fluor 647. Ubiquitin (ab134953) and His-FITC (ab1206) were purchased from Abcam, while Flag-FITC (MA1-91878-D488), Flag-Alexa Fluor 647 (MA1-142-a647), Actin dye phalloidin-Alexa Fluor 488 (A12379), and Myc-rhodamine (MA1-980-A555) were purchased from Invitrogen. Rabbit polyclonal antibody against histone H3 was purchased from Proteintech (17168-1-AP). A mouse monoclonal antibody against Hsp70 was purchased from Abmart (M20041S). Mouse monoclonal antibodies against GST (HT601), β-actin (HC201), β-tubulin (HC101-01), His (HT501) and Flag (HT201) were purchased from Transgene Biotech. Mouse monoclonal antibodies against Myc (D191042) and rabbit polyclonal antibodies against GAPDH (D110016) were purchased from Sangon Biotech.
RT-qPCR assay
To investigate the gene expression levels of Pelo, Hbs1, E3, and Hsp70 in insect vectors, the total RNAs of 50 intact insects infected with RGDV or treated with dsRNAs were extracted using TRIzol reagent (Invitrogen) following the manufacturer’s instructions. Reverse transcription was then performed using RevertAid reverse transcriptase (Promega, EP0441) and the appropriate primers following the manufacturer’s instructions. All RT-qPCRs were performed in triplicate using the SYBR Green PCR Master Mix Kit (GenStar, A304-10) according to the manufacturer’s instructions. The relative quantification of the genes was performed using the 2−ΔΔCt method56. The housekeeping gene elongation factor 1 of R. dorsalis served as the internal reference. The primers used in RT-qPCR assays are shown in Supplementary Table 1.
Western blot assay
To detect the accumulation levels of proteins from RGDV or R. dorsalis, the total proteins from 100 nonviruliferous, viruliferous, or dsRNAs treated intact insects were extracted and detected by western blot assay with Pns11-, P8-, Pelo-, Hbs1-, E3-, Hsp70-, or H3 IgG as the primary antibody and goat anti-rabbit IgG-peroxidase (Invitrogen, 32460) or goat anti-mouse IgG-peroxidase (Invitrogen, 32430) as the secondary antibody. The bands were detected using the Luminata Classico Western HRP Substrate (Millipore, WBKLS0500), and images were obtained using the Molecular Imager ChemiDoc XR + System (Bio-Rad). ImageJ (https://imagej.nih.gov/ij/) was used for signal quantitation.
Y2H assay
Y2H assay was used to examine the interaction relationship for Pelo, Hbs1, E3, Hsp70, Pns11, and P8. The full-length ORFs of Pns11, P8, Pelo, and E3 were inserted into the bait vector pGBKT7 to generate pGBKT7-Pns11, pGBKT7-P8, pGBKT7-Pelo, and pGBKT7-E3, respectively. The N-terminal segment (amino acids 1–200, E3-N) and C-terminal segment (amino acids 201-658, E3-C) of E3 were inserted into the bait vector pGBKT7 to generate pGBKT7-E3-N, pGBKT7-E3-C, respectively. The full-length ORFs of Pelo, Hbs1, Hsp70, and Pns11 were inserted into the prey vector pGADT7 to generate pGADT7-Pelo, pGADT7-Hbs1, pGADT7-Hsp70, and pGADT7-Pns11, respectively. The N-terminal segment (amino acids 1–130, Pelo-N), middle segment (amino acids 136–268, Pelo-M), and C-terminal segment (amino acids 271–370, Pelo-C) of Pelo was inserted into the prey vector pGADT7 to generate pGADT7-Pelo-N, pGADT7-Pelo-M, and pGADT7-Pelo-C, respectively. The bait and prey plasmids were co-transformed into the yeast strain AH109. The pGBKT7-53/pGADT7-T interaction served as a positive control, while the pGBKT7-Lam/pGADT7-T served as a negative control. Transformants were then screened on the QDO (SD/-Trp-Leu-His-Ade) culture medium. The primers used in Y2H are shown in Supplementary Table 1.
GST Pull-down assay
GST pull-down assay was used to confirm the interaction between Pelo and Pns11, Pelo and Hbs1, Hsp70 and Pns11, or Pelo and E3. In brief, ORFs of Pns11 and Pelo were cloned into pGEX-4T-3 to express GST-fusion protein. Then, ORFs of Pelo, Hbs1, Hsp70, and E3 were cloned into pET28a to express His-tag-fusion protein. The primers used in the GST pull-down are shown in Supplementary Table 1. All recombinant proteins were expressed in the Rosetta E. coli strain (Biomed, BC204-01). GST-Pns11 or GST-Pelo was incubated with glutathione-Sepharose beads (Amersham, 17-0756-01) for 5 h at 4 °C, centrifuged for 5 min at 500 × g, and then the supernatant was discarded. Either His-Pelo or His-Hsp70 was added to the beads binding with Pns11, or His-Hbs1 or His-E3 was added to the beads binding with Pelo. Then, the mixture was incubated for 5 h at 4 °C. The beads were collected and washed with PBS and the immunoprecipitated proteins were detected by western blot assay with His (Sangon, HT501) and GST (Transgene, HT601) antibodies.
Immunofluorescence microscopy
The male and female reproductive organs from nonviruliferous and viruliferous R. dorsalis adults were dissected and fixed in 4% paraformaldehyde in PBS (pH 7.2) for 2 h at room temperature. Samples were washed with PBS and permeabilised in 2% Triton X-100 (Sigma-Aldrich, T8787) for 40 min. The samples were washed with PBS and immunolabelled with Pns11-, Pelo-, Hbs1-, or E3-specific IgG conjugated to FITC, rhodamine, or Alexa Fluor 647, and visualised with a Leica TCS SP5 confocal microscope.
Effects of synthesised dsRNAs or purified proteins on RGDV infection in the male reproductive organs
The dsRNAs targeting Pelo (dsPelo), Hbs1 (dsHbs1), E3 (dsE3), and GFP (dsGFP) were synthesised in vitro using the T7 RiboMAX Express RNAi System (Promega Biotech, P1700). The primers used for dsRNA synthesis are shown in Supplementary Table 1. Approximately 600 third-instar nymphs of R. dorsalis were allowed to feed on RGDV-infected rice plants for two days and transferred to healthy rice seedlings for two weeks. Afterwards, 0.5 μg/μL dsRNAs were microinjected into the nonviruliferous and viruliferous male adults at the conjunction site between prothorax and mesothorax using a Nanoject II Auto-Nanoliter Injector (Spring). The dsGFP treatment served as a control. Five days later, the transcript levels of Pelo, Hbs1, E3, β-actin, and β-tubulin were tested with RT-qPCR assay, while the accumulation levels of Pelo, Hbs1, E3, P8, Pns11, β-actin, and β-tubulin were determined by western blot assay as described above. To detect the effect of Hbs1 on RGDV infection in the male R. dorsalis reproductive system, 0.01 μg/μL purified Hbs1 or GFP proteins expressed in E. coli Rosetta strain were microinjected into 100 male adults infected with RGDV for two weeks. Three days later, the accumulation levels of Hbs1, Pelo, Hsp70, P8, and Pns11 in the male reproductive organ were determined by western blot assay as described above.
Effect of synthesised dsRNAs on sperm morphology and the fitness of male leafhoppers and their offspring
To evaluate the effect of the dsRNA’s on the mortality rates of male adults, 50 nonviruliferous fifth instar nymphs were microinjected with dsPelo, dsHbs1, and dsGFP as control. After emergence, the males were kept separately in small cages, and the mortality rates were daily calculated. Meanwhile, dsRNAs-treated testes were dissected, fixed, permeabilised, immunolabelled with Pelo-FITC and Hbs1-rhodamine, and examined by immunofluorescence microscopy, as described above. Alternatively, after emergence, 10 pairs consisting of one nonviruliferous virgin female with one nonviruliferous male leafhopper microinjected with dsPelo or dsHbs1 were selected for oviposition on single rice seedlings in glass tubes. Female fecundity rates were calculated according to the average number of eggs laid by copulated females. Egg development duration was timed and hatch rates were calculated by the total number of neonates divided by the total number of neonates plus the number of non-hatched eggs. The combinations of nonviruliferous virgin females mating with nonviruliferous males microinjected with dsGFP were used as control. These experiments were conducted in triplicate.
Neutralising Pns11-sperm binding
To detect the direct interaction between Pns11 and Pelo on sperms, the mature live sperms excised from nonviruliferous males were pre-incubated with pre-immune or Pelo antibody (0.5 μg/μL) for 30 min and then incubated with purified Pns11 proteins (1.0 μg/μL) for 1 h. Subsequently, the sperms smeared on poly-lysine-treated glass slides were fixed, permeabilised, immunolabelled with Pns11-rhodamine (0.5 μg/μL), and then stained with DAPI (2.0 μg/mL). The sperms were visualised with a Leica TCS SP5 confocal microscope as described above.
To detect the effects of Pelo antibody on paternal transmission of RGDV in viruliferous male adults, viruliferous males were microinjected with pre-immune or Pelo antibody (0.5 μg/μL). Then, one nonviruliferous virgin female was mated with one viruliferous male adult microinjected with Pelo or pre-immune antibody for 3 days. Fifty pairs were performed for each mating combination. After mating, the males were tested with RT-PCR assays to determine the presence of RGDV. The offspring of each mating combination were also tested for RGDV by RT-PCR assays. The primers used in the RT-PCR assays are shown in Supplementary Table 1.
Baculovirus expression assay
To study the relationship for Pns11, Pelo, Hbs1, or Hsp70 in Sf9 cells, Flag-tagged Pelo (Pelo-Flag), His-tagged Hbs1 (Hbs1-His), Flag-tagged Hbs1 (Hbs1-Flag), His-tagged Hsp70 (Hsp70-His), and Myc-tagged Pns11 (Pns11-Myc) were cloned into pFast-bac1 vector and then introduced into E. coli DH10Bac strain (Biomed, BC112-01) to construct the recombinant bacmids. The recombinant bacmids infected Sf9 cells using Cellfectin II Reagent (ThermoFisher Scientific, 10362100) to obtain recombinant baculoviruses. To observe the effect of Pelo or Hbs1 on the expression and localization of Pns11, the recombinant baculoviruses expressing either Pelo-Flag or Hbs1-His, were co-infected or triply infected with Pns11-Myc in Sf9 cells. At 48 hpi, the cells were fixed, permeabilised, and immunolabelled with the appropriate antibodies, and then processed for immunofluorescence microscopy. To investigate the effect of the Hsp70 activity inhibitor on Pns11 tubule formation, Sf9 cells expressing Pns11-Myc were treated with DMSO or VER155008 (5, 10, and 20 μM). At 36 hpi, Sf9 cells were fixed, permeabilised, immunolabelled with Myc-rhodamine, and examined by immunofluorescence microscopy.
To observe the relationship between Hsp70 and Pns11, Sf9 cells were co-infected with the recombinant bacmids expressing Hsp70-His and Pns11-Myc. At 18 and 24 hpi, the cells were fixed, permeabilised, immunolabelled with His-FITC and Myc-rhodamine, and processed for immunofluorescence microscopy. To investigate the relationship for Hsp70, Hbs1, and Pns11, Sf9 cells were triply infected with the recombinant bacmids expressing Hsp70-His, Hbs1-Flag, and Pns11-Myc. At 24 and 36 hpi, the cells were fixed, permeabilised, immunolabelled with His-FITC, Flag-Alexa Fluor 647, and Myc-rhodamine, and processed for immunofluorescence microscopy.
Protein degradation assay
To investigate whether Pelo was degraded by the 26 S proteasome, the protease inhibitor MG132 (10 μM) was microinjected into viruliferous male adults. After microinjection for 24 h, approximately 100 testes of viruliferous males were dissected and tested with western blot assay using Pelo, Hbs1 or Pns11 antibody. Males treated with DMSO served as the control. To study the degradation mode of Pelo, the recombinant bacmids expressing Myc-tagged E3 (E3-Myc) were constructed. Thereafter, Sf9 cells singly expressing Pelo or co-expressing Pelo and E3-Myc were treated with MG132 (1 μM) for 6 h. The cells were then harvested using NP-40 lysis buffer for protein extraction. The accumulation levels of Pelo were detected by western blot assay as described above.
Co-IP for ubiquitination assay in Sf9 cells
HA-tagged ubiquitin (ubiquitin-WT-HA) and a mutant that only retained the 48th lysine site (ubiquitin-K48-HA) were inserted into the Fast-bac1 vector and then introduced into E. coli DH10Bac to construct the recombinant bacmids. Then, Sf9 cells were co-expressing Pelo-Flag with ubiquitin-WT-HA or ubiquitin-K48-HA, or triply expressing Pelo-Flag, ubiquitin-WT-HA/ ubiquitin-K48-HA and E3-Myc for two days. Correspondingly, Pns11-His was co-expressed with Pelo-Flag, ubiquitin-WT-HA, ubiquitin-K48-HA, or E3-Myc in Sf9 cells. The cells were harvested using NP-40 lysis buffer and incubated with Flag agarose beads (AlpalifeBio, KTSM1338) for 3 h at 4 °C. All beads were collected via a magnetic grate. The pellets were washed with PBS buffer (pH 7.2) 4-5 times and then mixed with 5 × loading buffer. Western blot assay was performed to analyse the ubiquitination of Pelo in the immunoprecipitant.
In vitro ubiquitination assay
To investigate the E3-mediated Pelo ubiquitination in vitro, ORFs of E1, E2, E3, and Pelo of R. dorsalis were cloned into vector pET-28a for fusion with His tag. The ORF of Pelo fused with the Flag tag at the C-terminal was cloned into the pET-28a vector. The recombinant proteins were expressed in the E. coli Rosetta strain and purified. The primers are listed in Supplementary Table 1. For the in vitro ubiquitination assay, 0.01 μg E1, 0.03 μg E2, 0.03 μg E3, 0.1 μg Pelo, and 0.1 μg ubiquitin (ThermoFisher Scientific) were incubated in ubiquitination buffer (1 M Tris, pH 7.2, 100 mM ATP, 200 mM MgCl2, 40 mM DTT, 20 ×) at 30 °C. After 90 mins, proteins were collected and analysed by western blot assay using ubiquitin and Flag antibodies as described above.
Competitive binding assay
To investigate the competitive relationship among E3, Pelo, and Pns11, a competitive binding assay was performed as previously described38. Briefly, E3 and Pns11 were cloned into the pET-28a vector to express E3-His and Pns11-His. Pelo was cloned into the pGEX-4T-3 vector to express GST-Pelo. The recombinant proteins E3-His, Pns11-His and GST-Pelo were expressed in E. coli Rosetta and purified. Thereafter, GST-Pelo was incubated with glutathione-Sepharose beads for 5 h at 4 °C, the mixture was centrifuged for 5 min at 500 × g, and the supernatant was discarded. Subsequently, 100, 200, or 400 μL E3-His (0.2 mg/mL) mixed with 200 μL Pns11-His (0.5 mg/mL), or 100, 200, or 400 μL Pns11-His (0.5 mg/mL) mixed with 200 μL E3-His (0.2 mg/mL) were added to the beads and incubated for 2 h at 4 °C. The beads were centrifuged and washed with PBS buffer (pH 7.2) 4-5 times. The bead-bound proteins were detected using E3, Pns11, or GST antibody by western blot assays as described above.
Electron microscopy
The male and female reproductive organs dissected from nonviruliferous, viruliferous, or dsGFP- and dsPelo-treated R. dorsalis individuals were fixed, dehydrated and embedded. Samples were sectioned on an ultramicrotome (Leica EM UC6) with a diamond knife, as previously described13. Sections were then incubated with Pns11- Pelo-, Hbs1-, or E3-specific IgG and immunolabelled with goat antibody against rabbit IgG conjugated to 15 nm-diameter gold particles (Sigma-Aldrich, G7402). Samples were observed under a transmission electron microscope (H-7650; Hitachi).
Sequence homology analysis
The amino acid sequences encoded by Pelo, Hbs1, or E3 were predicted using the online software Smart (http://smart.embl-heidelberg.de/). The Pelo, Hbs1, or E3 amino acid sequences of R. dorsalis, Drosophila melanogaster, Aedes aegypti, Homo sapiens, and Mus musculus were aligned using the ESPript 3.0 software (https://espript.ibcp.fr/ESPript/cgi-bin/ESPript.cgi).
Statistical analyses and reproducibility
To analyse the probability of discrepancies within a specific range, all quantitative data presented in the text and figures were analysed with an unpaired two-tailed t test or Tukey’s HSD multiple test in GraphPad Prism 8 software (GraphPad Software, San Diego, CA, USA).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data needed to evaluate the conclusions in the paper are present in source data files. The nucleotide sequences of Pelo, Hbs1 and E3 from R. dorsalis generated in this study were deposited in the Genbank under accession code PP091976.1, PP092032.1, PP092036.1, respectively. Source data are provided with this paper.
References
Weaver, S. C. & Reisen, W. K. Present and future arboviral threats. Antivir. Res. 85, 328–345 (2010).
Wang, H., Chen, Q. & Wei, T. Complex interactions among insect viruses-insect vector-arboviruses. Insect Sci. https://doi.org/10.1111/1744-7917.13285 (2023).
Schopen, S., Labuda, M. & Beaty, B. Vertical and venereal transmission of California group viruses by Aedes triseriatus and Culiseta inornata mosquitoes. Acta Virol. 35, 373–382 (1991).
Silverstein, P. S., Aljouda, N. A. & Kumar, A. Recent developments in vertical transmission of ZIKA virus. Oncotarget 7, 62797–62798 (2016).
Mao, Q. et al. Viral pathogens hitchhike with insect sperm for paternal transmission. Nat. Commun. 10, 955 (2019).
Heath, C. J. et al. Evidence of transovarial transmission of Chikungunya and Dengue viruses in field-caught mosquitoes in Kenya. PLoS Negl. Trop. Dis. 14, e0008362 (2020).
Huo, Y. et al. Transovarial transmission of a plant virus is mediated by vitellogenin of its insect vector. PLoS Pathog. 10, e1003949 (2014).
Jia, D. et al. Insect symbiotic bacteria harbour viral pathogens for transovarial transmission. Nat. Microbiol. 2, 17025 (2017).
Liao, Z. et al. Virus-Induced Tubules: A vehicle for spread of virions into ovary oocyte cells of an insect vector. Front. Microbiol. 8, 475 (2017).
Mao, Q. et al. Insect bacterial symbiont-mediated vitellogenin uptake into oocytes to support egg development. mBio 11, e01142–20 (2020).
Wei, J. et al. Vector development and vitellogenin determine the transovarial transmission of begomoviruses. Proc. Natl. Acad. Sci. USA 114, 6746–6751 (2017).
Wu, W. et al. Interaction of viral pathogen with porin channels on the outer membrane of insect bacterial symbionts mediates their joint transovarial transmission. Philos. Trans. R. Soc. B Biol. Sci. 374, 20180320 (2019).
Wan, J. et al. Arboviruses and symbiotic viruses cooperatively hijack insect sperm-specific proteins for paternal transmission. Nat. Commun. 14, 1289 (2023).
Hu, S. G., Du, H., Yao, G. X. & Zhang, Y. L. Molecular cloning and identification of mouse epididymis-specific gene mHong1, the homologue of rat HongrES1. Asian J. Androl. 14, 626–634 (2012).
Ma, L. et al. Spink13, an epididymis-specific gene of the Kazal-type serine protease inhibitor (SPINK) family, is essential for the acrosomal integrity and male fertility. J. Biol. Chem. 288, 10154–10165 (2013).
Li, Z., Yang, F., Xuan, Y., Xi, R. & Zhao, R. Pelota-interacting G protein Hbs1 is required for spermatogenesis in Drosophila. Sci. Rep. 9, 3226 (2019).
Yang, F. et al. The RNA surveillance complex Pelo-Hbs1 is required for transposon silencing in the Drosophila germline. EMBO Rep. 16, 965–974 (2015).
Burnicka-Turek, O. et al. Pelota interacts with HAX1, EIF3G and SRPX and the resulting protein complexes are associated with the actin cytoskeleton. BMC Cell Biol. 11, 28 (2010).
Liakath-Ali, K. et al. An evolutionarily conserved ribosome-rescue pathway maintains epidermal homeostasis. Nature 556, 376–380 (2018).
Graille, M., Chaillet, M. & van Tilbeurgh, H. Structure of yeast Dom34: a protein related to translation termination factor Erf1 and involved in No-Go decay. J. Biol. Chem. 283, 7145–7154 (2008).
Kobayashi, K., Ishitani, R. & Nureki, O. Recent structural studies on Dom34/aPelota and Hbs1/aEF1α: important factors for solving general problems of ribosomal stall in translation. Biophysics 9, 131–140 (2013).
Lee, H. H. et al. Structural and functional insights into Dom34, a key component of no-go mRNA decay. Mol. Cell 27, 938–950 (2007).
Nelson, R. J., Ziegelhoffer, T., Nicolet, C., Werner-Washburne, M. & Craig, E. A. The translation machinery and 70 kd heat shock protein cooperate in protein synthesis. Cell 71, 97–105 (1992).
O’Connell, A. E. et al. Mammalian Hbs1L deficiency causes congenital anomalies and developmental delay associated with Pelota depletion and 80S monosome accumulation. PLoS Genet. 15, e1007917 (2019).
Shoemaker, C. J., Eyler, D. E. & Green, R. Dom34:Hbs1 promotes subunit dissociation and peptidyl-tRNA drop-off to initiate no-go decay. Science 330, 369–372 (2010).
Xi, R., Doan, C., Liu, D. & Xie, T. Pelota controls self-renewal of germline stem cells by repressing a Bam-independent differentiation pathway. Development 132, 5365–5374 (2005).
Wu, X. et al. pelo is required for high efficiency viral replication. PLoS Pathog. 10, e1004034 (2014).
Asad, S. et al. Suppression of the pelo protein by Wolbachia and its effect on dengue virus in Aedes aegypti. PLoS Negl. Trop. Dis. 12, e0006405 (2018).
Shi, H. et al. Bta-miR-2411 attenuates bovine viral diarrhea virus replication via directly suppressing Pelota protein in Madin-Darby bovine kidney cells. Vet. Microbiol. 215, 43–48 (2018).
Chen, Y. et al. Adverse effects of rice gall dwarf virus upon its insect vector Recilia dorsalis (Hemiptera: Cicadellidae). Plant Dis. 100, 784–790 (2016).
Yi, G., Wu, W. & Wei, T. Delivery of rice gall dwarf virus into plant phloem by its leafhopper vectors activates callose deposition to enhance viral transmission. Front. Microbiol. 12, 662577 (2021).
Akita, F. et al. Viroplasm matrix protein Pns9 from rice gall dwarf virus forms an octameric cylindrical structure. J. Gen. Virol. 92, 2214–2221 (2011).
Shimizu, T. et al. Hairpin RNA derived from the gene for Pns9, a viroplasm matrix protein of Rice gall dwarf virus, confers strong resistance to virus infection in transgenic rice plants. J. Biotechnol. 157, 421–427 (2012).
Wei, T. et al. Association of Rice gall dwarf virus with microtubules is necessary for viral release from cultured insect vector cells. J. Virol. 83, 10830–10835 (2009).
Chen, H. et al. Rice gall dwarf virus exploits tubules to facilitate viral spread among cultured insect vector cells derived from leafhopper Recilia dorsalis. Front. Microbiol. 4, 206 (2013).
Jacob, P., Hirt, H. & Bendahmane, A. The heat-shock protein/chaperone network and multiple stress resistance. Plant Biotechnol. J. 15, 405–414 (2017).
Liang, Q. et al. A plant reovirus hijacks the DNAJB12-Hsc70 chaperone complex to promote viral spread in its planthopper vector. Mol. Plant Pathol. 23, 805–818 (2022).
Yu, X. et al. A plant reovirus hijacks endoplasmic reticulum-associated degradation machinery to promote efficient viral transmission by its planthopper vector under high temperature conditions. PLoS Pathog. 17, e1009347 (2021).
Xu, F. et al. HSP70 inhibitor VER155008 suppresses pheochromocytoma cell and xenograft growth by inhibition of PI3K/AKT/mTOR and MEK/ERK pathways. Int. J. Clin. Exp. Pathol. 12, 2585–2594 (2019).
Li, Y. et al. An integrated bioinformatics platform for investigating the human E3 ubiquitin ligase-substrate interaction network. Nat. Commun. 8, 347 (2017).
Ruan, H. B., Nie, Y. & Yang, X. Regulation of protein degradation by O-GlcNAcylation: crosstalk with ubiquitination. Mol. Cell Proteom. 12, 3489–3497 (2013).
Yang, Q., Zhao, J., Chen, D. & Wang, Y. E3 ubiquitin ligases: styles, structures and functions. Mol. Biomed. 2, 23 (2021).
Li, X. et al. CUL3 (cullin 3)-mediated ubiquitination and degradation of BECN1 (beclin 1) inhibit autophagy and promote tumor progression. Autophagy 17, 4323–4340 (2021).
Yu, X. et al. E3 ubiquitin ligase RNF187 promotes growth of spermatogonia via lysine 48-linked polyubiquitination-mediated degradation of KRT36/KRT84. Faseb J. 37, e23217 (2023).
Chen, Q. et al. VDAC1 balances mitophagy and apoptosis in leafhopper upon arbovirus infection. Autophagy 19, 1678–1692 (2023).
Jia, D. et al. A nonstructural protein encoded by a rice reovirus induces an incomplete autophagy to promote viral spread in insect vectors. PLoS Pathog. 18, e1010506 (2022).
Liang, Q. et al. A plant nonenveloped double-stranded RNA virus activates and co-opts BNIP3-mediated mitophagy to promote persistent infection in its insect vector. Autophagy 19, 616–631 (2023).
Young, D. J. & Guydosh, N. R. Rebirth of the translational machinery: The importance of recycling ribosomes. Bioessays 44, e2100269 (2022).
Du, J. et al. A cryptic K48 ubiquitin chain binding site on UCH37 is required for its role in proteasomal degradation. Elife 11, e76100 (2022).
Villamil, M. et al. The ubiquitin interacting motif-like domain of Met4 selectively binds K48 polyubiquitin chains. Mol. Cell Proteom. 21, 100175 (2022).
Castrillon, D. H. et al. Toward a molecular genetic analysis of spermatogenesis in Drosophila melanogaster: characterization of male-sterile mutants generated by single P element mutagenesis. Genetics 135, 489–505 (1993).
Eberhart, C. G. & Wasserman, S. A. The pelota locus encodes a protein required for meiotic cell division: an analysis of G2/M arrest in Drosophila spermatogenesis. Development 121, 3477–3486, (1995).
Ge, L. et al. SUMOylation-modified Pelota-Hbs1 RNA surveillance complex restricts the infection of potyvirids in plants. Mol. Plant. 16, 632–642 (2023).
Wu, X., Yang, Z. H., Wu, J. & Han, J. Ribosome-rescuer PELO catalyzes the oligomeric assembly of NOD-like receptor family proteins via activating their ATPase enzymatic activity. Immunity 56, 926–943 (2023).
Jia, D. et al. Development of an insect vector cell culture and RNA interference system to investigate the functional role of fijivirus replication protein. J. Virol. 86, 5800–5807 (2012).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408 (2001).
Acknowledgements
This project was supported by funds from the National Key Research and Development Programme of China to TW under grant number (2023YFD1400300), the National Natural Science Foundation of China to TW under grant number U23A20197 and 31920103014, the Natural Science Foundation of Fujian Province to HC under grant number 2021J01065.
Author information
Authors and Affiliations
Contributions
X.S., D.J., T.W. and Y.D. designed all experiments. X.S., D.J. and Y.C. performed the experiments for immunofluorescence staining and electron microscopy. X.S., Y.D., W.G. and Y.L. performed the protein interaction and ubiquitination experiments. X.S., Y.C. and H.C. performed the protein expression experiments. T.W., D.J., X.S. and Y.L. analysed the data. T.W., D.J. and X.S. organised the project and wrote the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Paolo Gabrieli and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sun, X., Du, Y., Cheng, Y. et al. Insect ribosome-rescuer Pelo-Hbs1 complex on sperm surface mediates paternal arbovirus transmission. Nat Commun 15, 6817 (2024). https://doi.org/10.1038/s41467-024-51020-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-51020-6
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.