Abstract
We examined tau haplotype frequencies in two different ethnical groups from the Basque Country (BC): Roma people and residents of European ancestry (general population). In addition, we analyzed the spatial distribution of tau haplotypes in Eurasian populations to explore the genetic affinities of the Romani groups living in Europe in a broader scope. The 17q21.31 genomic region was characterized through the genotyping of two diagnostic single nucleotide polymorphisms, SNPs (rs10514879 and rs199451), which allow the identification of H1 and H2 haplotypes. A significant heterozygous deficit was detected in the Romani for rs10514879. The H2 haplotype frequency proved to be more than twice in the BC general population (0.283) than in the Roma people (0.127). In contrast, H2 frequency proved to be very similar between Basque and Hungarian Romani, and similar to the H2 frequencies found in northwestern India and Pakistan as well. Several statistical analyses unveiled genetic structuring for the MAPT diversity, mirrored in a significant association between geography and genetic distances, with an upward trend of H2 haplotype frequencies from Asia to Europe. Yet, Roma samples did not fit into this general spatial patterning because of their discrepancy between geographical position and H2 frequency. Despite the long spatial coexistence in the Basque region between the residents of European ancestry and the Roma, the latter have preserved their Asian genetic ancestry. Bearing in mind the lack of geographical barriers between both ethnical groups, these findings support the notion that sociocultural mores might promote assortative matings in human populations.
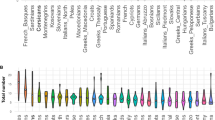
Similar content being viewed by others
Introduction
Over the recent years, much thought and investigation have been done on the structural diversity of the 17q21.31 genomic region of human chromosome 17, due essentially to its association with several neurodegenerative disorders such as Alzheimer's and Parkinson's diseases, sporadic frontotemporal dementia, progressive supranuclear palsy, amyotrophic lateral sclerosis and corticobasal degeneration, among others (Skipper et al. 2004; Ballatore et al. 2007; Zody et al. 2008; Arendt et al. 2016; Woerman et al. 2016). These pathologies are all characterized by the presence of abundant filamentous deposits of hyperphosphorylated Tau protein in nerve cells and glial cells, known as neurofibrillary tangles (NFTs). Such proteins are coded by the microtubule-associated protein tau (MAPT) gene and are involved in the regulation of the nucleation, elongation, and stabilization of neuronal microtubules (Lee et al. 2001; Ballatore et al. 2012).
The MAPT gene is located within a 1.5-Mb linkage disequilibrium block of human chromosome 17 (Myers et al. 2007). A substantial part of this genomic block (970 kb) may be inverted in some chromosomes, which allows the identification of two distinct haplotype clades, termed H1 (direct orientation) and H2 (inverted orientation) (Baker et al. 1999; Stefansson et al. 2005). In addition to the complex genomic architecture of the 17q21.31 inversion, the tau region has also attracted the attention of the scientific community owing to the enigmatic origin of H1 and H2 haplotype clades in our species and the intriguing spatial spreading of this structural polymorphism (Zody et al. 2008; Rao et al. 2010). Earlier investigations found notable differences in the geographical distribution of the MAPT-related haplotypes (Evans et al. 2004). Haplotype H1 has been widely predominant in all populations analyzed so far. Conversely, tau H2 haplotype is nonexistent or very scarce, with frequencies always <0.1 across the continents, excepting for populations of European ancestry, where H2 can reach frequencies of up to 0.38 (Evans et al. 2004, Stefansson et al. 2005).
In the present study we examined tau haplotype frequencies in two ethnically distinct human groups from the same region of northern Spain: the Roma people settled for generations in the Basque Country, and the general population from this geographical area. Our main anthropological interest was obviously focused on the Romani population. The Roma people or Gypsies constitute a broadly disseminated ethnic group, mostly characterized by the lack of a specific nation-state, the practice of different religions, the existence of different languages, and the presence of a wide spectrum of divergent groups separated by very strict sociocultural mores, including those concerning marriage patterns (Kalaydjieva et al. 2005). In the Roma society, the primary unit is the group, and groups are members of metagroups. They live in a closed-society structure, with unusual admixture with other populations and a relatively high rate of consanguineous matings in several Roma communities (Assal et al. 1991; Martinez-Frias and Bermejo 1992).
Both the origins and the early demographic history of the Roma have been controversial topics among the scientists owing to the lack of well-documented records (Hancock 1993). Yet, some linguistic and genetic evidences seem to indicate that the Roma people originated in India. Nowadays, most philologists allege that Romani language probably evolved in the northwestern region of India as a result of the confluence of widely spoken languages close to Sanskrit (Beníšek 2010; Bakker and Monrad 2011). Concerning genetic evidences, unambiguous signals of the Indian ancestry of the Roma people are obtained from Y chromosome haplogroup H1a1a-M82 (Rai et al. 2012), mtDNA haplogroup M (Mendizabal et al. 2011) and the pathogenic 1267delG mutation in the gene encoding for the epsilon subunit of the acetylcholine receptor (CHRNE) causing autosomal recessive congenital myasthenia, found on the same ancestral chromosomal background in Roma, Indian and Pakistani subjects (Morar et al. 2004). The combined evidence suggests that the Roma migrated from the Punjab region of northwest India 1000–1500 years ago and traveled through Asia (along Persia, today’s Armenia and Turkey). The mainstream moved into the Balkans and Greece and some of them into Eastern Europe ahead of the Turks. Early diaspora appeared in Western Europe around the period from the fourteenth to the fifteenth century, and another wave of migrations to Western Europe started after the abolition of serfdom in the Habsburg Empire in 1841, and recently from 1989 after the disappearance of the Iron Curtain (Kalaydjieva et al. 2005).
The current Roma population in Europe is estimated at around 10 million people (European Communities 2004), and the largest communities are located in Central and Eastern Europe. According to recent estimates, Bulgaria, Hungary and Romania are the Eastern European states with the highest number of Roma people. In Western Europe, Spain features the highest number of Romani (Corsi et al. 2008), with a population ranging between 700,000–970,000 inhabitants mostly concentrated in Andalusia (La Parra et al. 2013). It has been argued that the arrival to Spain was through the Pyrenees in the early fifteenth century, with the first documented presence of Roma individuals in Barcelona in 1425 (Callén et al. 2005).
Bearing in mind that the 17q21.31 inversion is a structural polymorphism, the probability of MAPT genomic region being affected by back-mutations or recurrent mutations is practically zero. For that reason, analysis of tau haplotype frequencies might be suitable to detect ancient genetic affinities and the potential effects of gene flow and genetic drift in shaping the gene pool of the human populations. As previously mentioned, in this work we have examined the frequencies of the tau haplotypes in two human groups from the Basque Country, with particular emphasis on the Roma people, through the analysis of two diagnostic single-nucleotide polymorphism (SNP) markers (Donnelly et al. 2010; Steinberg et al. 2012). To widen the scope of our results, we also explored the geographical distribution of tau H2 haplotype in the Eurasian context using data compiled in the relevant literature, in an attempt to provide further evidence on the Asian ancestry of the Roma people settled in Europe for centuries.
Materials and methods
Population samples
In this study, a total of 67 Roma individuals living in Biscay province (Basque Country, North Spain) were genotyped. Only those individuals that self-identified as Roma people were considered in the sampling process. Likewise, the Roma collection included only persons with Iberian origins to minimize the within-group genetic heterogeneity. Samples were collected in health centers and public schools with high Romani attendance and in Roma family houses. A methodical description of the sample can be consulted in Poveda et al. (2012). Written informed consent was obtained from all study participants and from their parents or guardians in the case of minors. Permission to carry out the study in the public centers was requested from the head of each center.
In addition, a sample of 23 individuals from the resident population of Bilbao was included in the analysis as a reference group. For this latter group, no autochthony criterion (e.g., Basque origins of the subject and its ancestors) was considered in the sampling process; thus, this collection was considered to be representative of the Spanish population. To avoid the existence of close genetic kinship among the members of these samples, ancestry of individuals was assessed by biographical information traced back at least two generations. In this way, all voluntary donors were healthy individuals without kinship relationships of first or second degree.
Ethical guidelines for research with human beings were adhered to as stipulated by the Institutional Review Board from the University of the Basque Country (UPV/EHU). The study protocol was approved by the Ethics Committee of the cited institution.
SNP typing
Genomic DNA was isolated from saliva samples using the Oragene DNA sample collection kit OG-250 (DNA Genotek, Ottawa, Ontario, Canada) according to manufacturer’s instructions. Genetic characterization of the 17q21.31 genomic region was carried out by analysis of the SNPs rs10514879 and rs199451 using high resolution melting (HRM) assay. We typed the SNP markers using the following primer pairs: 5’ TGA GAT CCG GCA GAT AAA TG TG 3’ (forward) and 5’ AAT GGA CTC TGA AAT CTC ACT GTC 3’ (reverse) for rs10514879 (Donnelly et al. 2010), and 5’ TCA GAG ACT CAA GCT AAT AG 3’ (forward) and 5’ TGA TTC ACC ATA CTC CTT TCC C 3’ (reverse) for rs199451 (Steinberg et al. 2012). Purified DNA templates were amplified in a Bio-Rad C1000 real-time thermal cycler (Bio-Rad Laboratories, Hercules, CA) using a Bio-Rad CFX96 optical reaction module. Real-time PCR reactions were performed in a final volume of 5 µl using the following reagents: 0.15 µM of each primer, 2.5 µL of SsoFast™ EvaGreen® supermix (Bio-Rad) and 15 ng of genomic DNA. The PCR conditions for the SNP rs10514879 were as follows: initial denaturation at 98° C for 3 min; 39 cycles at 98° C for 10 s and 61.2° C for 30 s (58.2° C for the SNP rs199451). The plate read was taken after an initial step of 10 s at 95° C and 65° C for 2 min. The melt curve was from 65° C to 95° C with an increment of 0.5° C each after 5 s. Melting profiles were analyzed with the Bio-Rad Precision Melt Analysis Software v1.0 (Bio-Rad).
Statistical analysis
Allele frequencies of the SNP loci examined were estimated by direct counting.
Hardy–Weinberg equilibrium (HWE) was assessed by a Fisher’s exact probability test to estimate P-values, using the Arlequin v3.5 program (Excoffier and Lischer 2010). Allelic combinations of the SNP loci rs10514879 and rs199451 were then used for the assignment of the MAPT haplotypes H1, H2’ and H2D. According to previously published works, alleles A and G of rs10514879 allow accurate identification of H1 and H2 tau haplotypes, respectively (Donnelly et al. 2010). Likewise, allele G of rs199451 discriminates the haplotypes without duplications (H1 and H2’), whereas allele A is suitable to identify haplotype H2D, which is the variant with interspersed duplications of H2 (Steinberg et al. 2012).
Tau haplotype frequencies from previous studies were compiled to assess the genetic relationships of both BC Roma and BC general population in a broader geographical scope. A total of 33 Eurasian populations previously typed for the targeted SNPs were utilized for the assays. Populations considered in such analyses, sample sizes, and the corresponding references can be consulted in Supplementary Table S1. Bearing in mind the paucity of data for tau H2’ and H2D haplotypes in the Indian populations, comparative analyses were specifically focused on H1 and H2. These haplotype frequencies were employed in determining the fraction of genetic variability attributable to differences within and among populations through the analysis of molecular variance, AMOVA (Excoffier et al. 1992; Weir and Cockerham 1984). AMOVA tests were carried out considering two (Asia and Europe) and three (Europe, Middle East and South Asia) population clusters classed according to geography. In the first of them, Caucasian populations from the Middle East (Palestinians, Bedouins, Samaritans and Druze) were included into the European cluster. Two alternative AMOVA tests were planned, in an attempt to ascertain maximum genetic variance between groups (FCT) and minimum genetic variance among populations within groups (FSC): one of them including Romani samples (Basque Country and Hungarian Olah Roma) within the European group (case a), and an alternative one where both Roma collections were classified as Asian populations (case b).
We further investigated the spatial trend of the haplotype frequency distributions in a system of moving coordinates to detect potential frequency gradients (genetic clines). This analysis was performed by calculating the linear regression of the tau haplotype frequencies (H1and H2) in a series of populations using the GenoCline program (Peña et al. 2016). This software rotates a virtual coordinate axis in consecutive iterations of one degree each until completing 360 degrees. The position of all the populations is projected on this rotated axis in every iteration. Then the program carries out both the linearity test and the Pearson product-moment correlation coefficient between haplotype frequencies and the coordinates of the populations with respect to the virtual axis. A statistically significant association (P < 0.05 in the linearity test) between both variables will be indicative of a spatially patterned distribution of the haplotype frequencies, that is, of a haplotype frequency cline. In those cases where, for the same tau haplotype, more than one statistically significant association is obtained between frequency and spatial distribution, the program will select that direction of the axis for which the coefficient of determination (r2) rendered its maximum value. Finally, a Mantel test of matrix correspondence (Mantel 1967) was applied to evaluate the concordance between genetic and geographical distances of the populations involved. To that end, haplotype frequencies were employed to compute FST genetic distances among populations (Reynolds et al. 1983) with the Arlequin v3.5 program. Geographical distances were calculated from geographical coordinates and considering the curvature of the terrestrial surface using the GenoCline software (Peña et al. 2016).
Results
Allele frequencies for diagnosis SNPs (rs10514879 and rs199451) as well as haplotype frequencies in the two Basque Country samples (BC Roma and BC general population) are summarized in Table 1. In accordance with the relative population isolation of the Romani owing to sociocultural constraints, observed heterozygosity values were lower in the Roma people than in the BC general population for both SNP markers. Accordingly, average heterozygosity was also lower in the Roma (0.163) than in the general population (0.341). No significant departure from HWE expectations was detected in the reference population of the Basque Country (Table 2). In contrast, rs10514879 showed a significant deviation from HWE expectations in the Romani sample. As can be inferred from the observed (Ho: 0.134) and expected (He: 0.222) heterozygosity values, heterozygous deficit seems to underlie the disequilibrium. Regarding tau haplotypes, the frequency of H2 in the BC general population (H2’ + H2D: 0.283) proved to be more than twice of that obtained for the Romani population (0.127).
AMOVA results based on tau haplotypes (Table 3) revealed statistically significant values for the fixation indices FCT and FSC (P < 0.0001), thereby indicating spatial patterning (or geographical structuring) of the interregional and intraregional genetic diversity, respectively. In all AMOVA analyses, the combination yielding the highest value of genetic heterogeneity between continental clusters (FCT) and the lowest value of genetic variation among populations within groups (FSC) was the one considering Roma populations from the Basque Country and Hungary as members of the Asian group (cases b in Table 3).
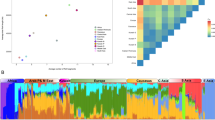
We further investigated the spatial distribution of the haplotype frequencies in a system of moving coordinates to detect potential genetic clines. Regression line of the H2 haplotype frequencies on the rotated geographical coordinates of 35 European and Asian populations are illustrated in Fig. 1. In line with the AMOVA findings, we detected a clear differentiation between European and South Asian populations in terms of tau H2 haplotype frequencies, with a conspicuous upward trend from Asia to Europe. Bearing in mind that H1 and H2 haplotype frequencies are complementary, H1 frequency cline showed exactly the opposite orientation. Linear regression analysis pointed toward a statistically significant influence of the geographic position on the H2 frequency (P < 0.001 in the linearity test), a result endorsed by a relatively high value for the coefficient of determination (r 2 = 0.515). Interestingly, an additional regression analysis performed by excluding the two Romani populations revealed a substantial increase of the linearity between geographical coordinates and H2 haplotype frequencies (r 2 = 0.623), suggesting that the heterogeneity in the association between these variables seems to be introduced by the Roma samples.
Regression line and 95% confidence intervals (dashed lines) in a regression analysis of tau H2 haplotype frequencies on the rotated geographical coordinates (H2 freq = 0.4256 − coord × 0.000083) of 35 European and Asian populations (coefficient of determination, r 2 = 0.515). Populations examined in this study are highlighted with a frame. Solid circles are European populations, solid squares are Middle Eastern populations, and solid triangles represent South Asian populations. Romani populations are designated by stars. Population labels: BC-Roma (Basque Country Roma), BC-resid (Basque Country general population), BC-Spain (Iberian Basques), BC-French (French Basques), UK (British), IT-Sardn (Sardinia, Italy), IT-Bergm (Bergamo, Italy), IT-Bresc (Brescia, Italy), IT-Tuscn (Tuscany, Italy), HU-Roma (Hungarian Roma), Palestn (Palestinians), and Samartn (Samaritans)
No overlapping of the haplotype frequencies was detected between European and Southern Asian populations. In South Asia, H2 haplotype frequency peaks at 0.122 in Mohanna, whereas the lowest frequency in Europe (0.157) has been reported for Denmark (Donnelly et al. 2010). Results generated by a Mantel test of matrix correspondence also indicated a statistically significant association between pairwise FST genetic distances (computed from haplotype frequencies) and the geographical distances of the targeted populations (r = 0.432; P < 0.0002), which further supports the increasing trend of the H2 frequencies from Asia to Europe. The matrix of pairwise FST genetic distances, statistical significance (P-values), geographical coordinates of populations and pairwise geographical distances can be consulted in supplemental material (Supplementary Tables 2, 3, 4 and 5, respectively).
As for the Romani samples, both the study population as well as the Hungarian Roma plotted relatively distant from the statistical trend represented by the regression line. Tau H2 frequency proved to be very similar between Romani from the Basque Country (0.093) and from Hungary (0.102), and therefore, visibly lower than in Europeans. For that reason the two samples of the Basque region (BC Roma and residents of European ancestry) remained clearly separated on the regression graph. As expected according to ancestry, H2 frequency in the Basque general population (0.273) fitted to the variation range in Europe, which oscillates between 0.157 in Denmark and 0.375 in Sardinia (Donnelly et al. 2010). A Fisher’s exact test unveiled statistically significant differences (P < 0.00001) for H1 and H2 haplotype frequencies between BC Roma and BC general population.
Discussion
In this study we have examined the frequencies of the tau haplotypes in two different human groups from the Basque Country: resident population (with European ancestry) and the Roma people settled in this northern region of the Iberian Peninsula. We also analyzed the geographical distribution of tau H2 haplotype in Eurasian populations to explore the genetic affinities of the Romani groups living in Europe in a wider context.
Among the most notable results was a different proportion of the inversion at the 17q21.31 genomic region in the Roma from the Basque Country relative to the resident population of this territory. Thus, BC Roma carried the tau H1 haplotype at a significantly higher proportion than the European-ancestry residents, in congruence with previous results obtained in a similar analysis between Roma (Hungarian Olah Roma) and non-Roma samples from Hungary (Almos et al. 2008). Such a finding supports the notion that 17q21.31 structural variation might be used in combination with other autosomal genetic markers as a potential ancestry informative marker (AIM) in evolutionary, forensic, and population admixture studies.
Another important finding of this work was the spatial patterning of the MAPT diversity, inferred from the strong association between geography and tau haplotype frequencies. Yet, it should be highlighted that the two targeted Roma samples remained separated from the main trend represented by the regression line because, in spite of being geographically located in Europe, both collections feature H2 haplotype frequencies within the variation range of the South Asian populations (basically from the Indian subcontinent), which oscillates between zero in Thoti, Pathan and Kachari, and 0.122 in Mohanna (Donnelly et al. 2010). Likewise, all northern Indian and Pakistani groups showed H2 frequencies comparable to those observed in the Romani populations settled in European territories (Hungary and the Spanish Basque Country), with the only exception of the Pathan sample. Such similarity regarding the MAPT inversion polymorphism might be a genetic trace of the geographical origin of the current Romani diaspora in Europe or, at least, it suggests a close genetic affinity between both human groups.
In recent years, several investigations based on a variety of genetic markers, including genome-wide-SNP loci (Mendizabal et al. 2012; Moorjani et al. 2013), and uniparentally inherited markers, such as Y-chromosome and mitochondrial DNA (mtDNA) lineages or haplogroups (Martínez-Cruz et al. 2016), have consistently situated the origin of the proto-Roma population in the northwestern region from the Indian subcontinent. Particularly, present-day Indian populations from Kashmir and Punjab have been postulated as strong candidates for being the source of the Indian ancestry in Roma. The close genetic relationships between European Romani and northern Indian populations derived from our analysis of the polymorphic inversion on the 17q21.31 genomic region strongly support this hypothesis, which is also compatible with findings from earlier linguistic and sociocultural studies (see Fraser 1992).
Our findings on the MAPT inversion polymorphism failed to find clear evidence of genetic heterogeneity between Romani groups by genetic drift effects or differential admixture with European host populations, as has been stated in precedent studies (Mendizabal et al. 2012; Martínez-Cruz et al. 2016). Two potential reasons might be argued to account for this discrepancy. First, the limited number of Romani samples included in our study due to the extreme paucity of MAPT population data in this ethnic group did not allow an exhaustive analysis of the genetic diversity among the European Roma. The second, and perhaps the more important, reason are the intrinsic features of the structural polymorphism examined. The polymorphic inversion at 17q21.31 is an extremely large block (∼ 970 kb) of high linkage disequilibrium. Accordingly, this genetic marker is more conservative of ancient population genetic relationships or, in other words, evolutionarily stable, because the probability of the MAPT genomic region being affected by recurrent or back-mutations is practically zero. The evolutionary stability of the MAPT inversion would also account for the close genetic relationships between the European Romani populations and the hypothetical proto-Roma populations from northwestern India and neighboring zones of Pakistan. Nevertheless, in explaining the genetic affinity between the European Romani and the northern Indian populations the effects of the genetic drift cannot be ruled out, bearing in mind both the small demographic size of the migrant Roma groups and the relative population isolation of the Romani people across time owing to a strong ethnicity.
As we have noted before, heterozygosity was visibly lower in the BC Roma population than in the BC general population. The significant heterozygous deficit (Hardy-Weinberg disequilibrium) detected in the Roma group for the SNP rs10514879 could be explained by the genetic effects of their closed-society structure and their reluctance to interethnic marriages, a phenomenon reflected in the high rates of endogamy and consanguinity among the Spanish Roma groups (Martinez-Frias and Bermejo 1992). In a survey of the prevalence of congenital anomaly syndromes in a Spanish Romani population, Martinez-Frias and Bermejo (1992) estimated that the proportion of consanguineous matings in the Roma group was 16 to 19.5 times that in non-Roma. Likewise, these authors associated the high inbreeding level of the Romani population with a higher proportion of homozygotes for recessive conditions in the offspring and, accordingly, with a higher rate of recessive syndromes, namely seven times higher in the Roma community. Deeply rooted sociocultural mores could be a major factor limiting gene flow and population admixture by preventing the integration of immigrants into the recipient population and by increasing ethnic endogamy (Alfonso-Sánchez et al. 2001; 2005).
Summarizing, MAPT inversion has shown to be a robust, conservative and stable marker for evolutionary studies dealing with the genetic ancestry of a given human group. The analysis of the MAPT inversion polymorphism indicated that the European Romani groups displayed much more genetic affinity with populations from the northwestern region of the Indian subcontinent than with European populations, which in principle would be in agreement with the hypothesis of the origin of the proto-Roma group in this Asian geographical area. In addition, even though MAPT inversions are conservative markers from the evolutionary viewpoint, they indirectly permitted evaluating the effect of stochastic evolutionary forces (genetic drift and gene flow) in the Eurasian context, taking into account that the survey of the spatial distribution of tau H2 frequencies unveiled a significant association between geography and genetics with a genetic cline from Asia to Europe.
An important added value of this work is the contribution of useful data from a highly endogamous human society. Roma population genetics represents a case study for understanding how a complex demographic history can impact the genetic make-up of a human population. In light of our findings, it can be deduced that, in spite of the prolonged coexistence between the Roma of the Basque Country and the rest of the European-ancestry community of this region, the Romani people have preserved a genetic background very similar to the hypothetical ancestral populations from the Indian subcontinent and with scarce influence of the European gene pool, which is in agreement with the findings reported for Hungarian Olah Romani (Almos et al. 2008). The deeply rooted ethnicity and sociocultural restrictions of the BC Romani population could have acted as a barrier to random mating through continuing strong preference for in-marriage, so that a native gene pool with a negligible admixture rate would have been preserved. The results of this study constitute a good example of how ethnopsychology and sociocultural factors may have notably impacted the genetic make-up of human populations.
References
Alfonso-Sánchez MA, Aresti U, Peña JA, Calderón R (2005) Inbreeding levels and consanguinity structure in the Basque province of Guipúzcoa (1862-1980). Am J Phys Anthropol 127:240–252
Alfonso-Sánchez MA, Peña JA, Aresti U, Calderón R (2001) An insight into recent consanguinity within the Basque area in Spain. Effects of autochthony, industrialization and demographic changes. Ann Hum Biol 28:505–521
Almos PZ, Horváth S, Czibula A, Raskó I, Sipos B, Bihari P, Béres J, Juhász A, Janka Z, Kálmán J (2008) H1 tau haplotype-related genomic variation at 17q21.3 as an Asian heritage of the European Gypsy population. Heredity 101:416–419
Arendt T, Stieler JT, Holzer M (2016) Tau and tauopathies. Brain Res Bull 126:238–292
Assal S, Susanszky E, Czeizel A (1991) High consanguinity rate in Hungarian gypsy communities. Acta Paediatr Hung 31:299–304
Baker M, Litvan I, Houlden H, Adamson J, Dickson D, Perez-Tur J, Hardy J, Lynch T, Bigio E, Hutton M (1999) Association of an extended haplotype in the Tau gene with progressive supranuclear palsy. Hum Mol Genet 8:711–715
Bakker P, Monrad A (2011) Roma: linguistic archaeology of nomads. AmS-Varia 53:35–44
Ballatore C, Brunden KR, Huryn DM, Trojanowski JQ, Lee VM, Smith III AB (2012) Microtubule stabilizing agents as potential treatment for Alzheimer’s disease and related neurodegenerative tauopathies. J Med Chem 55:8979–8996
Ballatore C, Lee VMY, Trojanowski JQ (2007) Tau-mediated neurodegeneration in Alzheimer’s disease and related disorders. Nat Rev Neurosci 8:663–672
Beníšek M (2010) The quest for a Proto-Romani infinitive. Romani Stud 20:47–86
Callén E, Casado JA, Tischkowitz MD, Bueren JA, Creus A, Marcos R, Dasí A, Estella JM, Muñoz A, Ortega JJ, de Winter J, Joenje H, Schindler D, Hanenberg H, Hodgson SV, Mathew CG, Surrallés J (2005) A common founder mutation in FANCA underlies the world’s highest prevalence of Fanconi anemia in Gypsy families from Spain. Blood 105:1946–1949
Corsi M, Crepaldi C, Lodovici MS, Boccagni P, Vasilescu C (2008) Ethnic minority and Roma women in Europe: A case for gender equality? Final Report. Expert Group on Gender equality, social inclusion, health and long term care.
Donnelly MP, Paschou P, Grigorenko E, Gurwitz D, Mehdi SQ, Kajuna SL, Barta C, Kungulilo S, Karoma NJ, Lu RB, Zhukova OV, Kim JJ, Comas D, Siniscalco M, New M, Li P, Li H, Manolopoulos VG, Speed WC, Rajeevan H, Pakstis AJ, Kidd JR, Kidd KK (2010) The distribution and most recent common ancestor of the 17q21 inversion in humans. Am J Hum Genet 86:161–171
European Communities (2004) The situation of Roma in an enlarged European Union. Office for Official Publications of the European Communities, Luxembourg
Evans W, Fung HC, Steele J, Eerola J, Tienari P, Pittman A, Silva Rd, Myers A, Vrieze FW, Singleton A, Hardy J (2004) The tau H2 haplotype is almost exclusively Caucasian in origin. Neurosci Lett 369:183–185
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Fraser A (ed.) (1992) The Gypsies. Blackwell Publishers, Oxford
Fung HC, Evans J, Evans W, Duckworth J, Pittman A, de Silva R, Myers A, Hardy J (2005) The architecture of the tau haplotype block in different ethnicities. Neurosci Lett 377:81–84
García-Obregón S, Alfonso-Sánchez MA, Pérez-Miranda AM, de Pancorbo MM, Peña JA (2007) Polymorphic Alu insertions and the genetic structure of Iberian Basques. J Hum Genet 52:317–327
Ghidoni R, Signorini S, Barbiero L, Sina E, Cominelli P, Villa A, Benussi L, Binetti G (2006) The H2 MAPT haplotype is associated with familial frontotemporal dementia. Neurobiol Dis 22:357–362
Hancock I (1993) The emergence of a Union Dialect of North American Vlax Romani, and its implications for an international standard. International Journal of the Society of Language 99:91–104
Kalaydjieva L, Morar B, Chaix R, Tang H (2005) A newly discovered founder population: the Roma/Gypsies. BioEssays 27:1084–1094
La Parra D, Gil-González D, Jiménez A (2013) Los procesos de exclusión social y la salud del pueblo gitano en España. Gac Sanit 27:385–386
Lee VM, Goedert M, Trojanowski JQ (2001) Neurodegenerative tauopathies. Annu Rev Neurosci 24:1121–1159
Mantel N (1967) The detection of disease clustering and a generalized regression approach
Martínez-Cruz B, Mendizabal I, Harmant C, de Pablo R, Ioana M, Angelicheva D, Kouvatsi A, Makukh H, Netea MG, Pamjav H, Zalán A, Tournev I, Marushiakova E, Popov V, Bertranpetit J, Kalaydjieva L, Quintana-Murci L, Comas D (2016) Origins, admixture and founder lineages in European Roma. Eur J Hum Genet 24:937–943
Martinez-Frias ML, Bermejo E (1992) Prevalence of congenital anomaly syndromes in a Spanish gypsy population. J Med Genet 29:483–486
Mendizabal I, Lao O, Marigorta UM, Wollstein A, Gusmao L, Ferak V, Ioana M, Jordanova A, Kaneva R, Kouvatsi A, Kucinskas V, Makukh H, Metspalu A, Netea MG, de Pablo R, Pamjav H, Radojkovic D, Rolleston SJ, Sertic J, Macek Jr. M, Comas D, Kayser M (2012) Reconstructing the population history of European Romani from genome-wide data. Curr Biol 22:2342–2349
Mendizabal I, Valente C, Gusmão A, Alves C, Gomes V, Goios A, Parson W, Calafell F, Alvarez L, Amorim A, Gusmão L, Comas D, Prata MJ (2011) Reconstructing the Indian origin and dispersal of the European Roma: a maternal genetic perspective. PLoS One 6:e15988
Moorjani P, Patterson N, Loh PR, Lipson M, Kisfali P, Melegh BI, Bonin M, Kádaši L, Rieß O, Berger B, Reich D, Melegh B (2013) Reconstructing Roma history from genome-wide data. PLoS One 8:e58633
Morar B, Gresham D, Angelicheva D, Tournev I, Gooding R, Guergueltcheva V, Schmidt C, Abicht A, Lochmuller H, Tordai A, Kalmar L, Nagy M, Karcagi V, Jeanpierre M, Herczegfalvi A, Beeson D, Venkataraman V, Warwick Carter K, Reeve J, de Pablo R, Kucinskas V, Kalaydjieva L (2004) Mutation history of the Roma/Gypsies. Am J Hum Genet 75:596–609
Myers AJ, Gibbs JR, Webster JA, Rohrer K, Zhao A, Marlowe L, Kaleem M, Leung D, Bryden L, Nath P, Zismann VL, Joshipura K, Huentelman MJ, Hu-Lince D, Coon KD, Craig DW, Pearson JV, Holmans P, Heward CB, Reiman EM, Stephan D, Hardy J (2007) A survey of genetic human cortical gene expression. Nat Genet 39:1494–1499
Peña JA, Alfonso-Sánchez MA, Gómez-Pérez L (2016) GenoCline version 1.0 User Manual. http://genocline.sourceforge.net
Poveda A, Ibáñez ME, Rebato E (2012) Heritability and genetic correlations of obesity-related phenotypes among Roma people. Ann Hum Biol 39:183–189
Rai N, Chaubey G, Tamang R, Pathak AK, Singh VK, Karmin M, Singh M, Rani DS, Anugula S, Yadav BK, Singh A, Srinivasagan R, Yadav A, Kashyap M, Narvariya S, Reddy AG, van Driem G, Underhill PA, Villems R, Kivisild T, Singh L, Thangaraj K (2012) The phylogeography of Y-chromosome haplogroup H1a1a-M82 reveals the likely Indian origin of the European Romani populations. PLoS One 7:e48477
Rao PN, Li W, Vissers LE, Veltman JA, Ophoff RA (2010) Recurrent inversion events at 17q21. 31 microdeletion locus are linked to the MAPT H2 haplotype. Cytogenet Genome Res 129:275–279
Reynolds J, Weir BS, Cockerman CC (1983) Estimation of the coancestry coefficient: bases for a short term genetic distance. Genetics 105:767–779
Skipper L, Wilkes K, Toft M, Baker M, Lincoln S, Hulihan M, Ross OA, Hutton M, Aasly J, Farrer M (2004) Linkage disequilibrium and association of MAPT H1 in Parkinson disease. Am J Hum Genet 75:669–677
Stefansson H, Helgason A, Thorleifsson G, Steinthorsdottir V, Masson G, Barnard J, Baker A, Jonasdottir A, Ingason A, Gudnadottir VG, Desnica N, Hicks A, Gylfason A, Gudbjartsson DF, Jonsdottir GM, Sainz J, Agnarsson K, Birgisdottir B, Ghosh S, Olafsdottir A, Cazier JB, Kristjansson K, Frigge ML, Thorgeirsson TE, Gulcher JR, Kong A, Stefansson K (2005) A common inversion under selection in Europeans. Nat Genet 37:129–137
Steinberg KM, Antonacci F, Sudmant PH, Kidd JM, Campbell CD, Vives L, Malig M, Scheinfeldt L, Beggs W, Ibrahim M, Lema G, Nyambo TB, Omar SA, Bodo J-M, Froment A, Donnelly MP, Kidd KK, Tishkoff SA, Eichler EE (2012) Structural diversity and African origin of the 17q21. 31 inversion polymorphism. Nat Genet 44(8):872–880
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Winkler S, König IR, Lohmann-Hedrich K, Vieregge P, Kostic V, Klein C (2007) Role of ethnicity on the association of MAPT H1 haplotypes and subhaplotypes in Parkinson’s disease. Eur J Hum Genet 15:1163–1168
Woerman AL, Aoyagi A, Patel S, Kazmi SA, Lobach I, Grinberg LT, McKee AC, Seeley WW, Olson SH, Prusiner SB (2016) Tau prions from Alzheimer’s disease and chronic traumatic encephalopathy patients propagate in cultured cells. Proc Natl Acad Sci USA 113:E8187–E8196
Zody MC, Jiang Z, Fung HC, Antonacci F, Hillier LW, Cardone MF, Graves TA, Kidd JM, Cheng Z, Abouelleil A, Chen L, Wallis J, Glasscock J, Wilson RK, Reily AD, Duckworth J, Ventura M, Hardy J, Warren WC, Eichler EE (2008) Evolutionary toggling of the MAPT 17q21.31 inversion region. Nat Genet 40:1076–1083
Acknowledgements
The authors are particularly indebted to all voluntary donors from the Roma community and from the resident population of the Spanish Basque Country, who generously cooperated to the development of this research. We also would like to express our gratitude to Kale Dor Kayiko (KDK) for their collaboration. This study was supported by grants from: Bilbao Bizkaia Kutxa (BBK; 87014/97012/07007), Spanish Ministry of Science and Innovation (MICINN; GCL2010-15511), Industry Department of the Basque Government (SAIOTEK; SA2010/00035), and by three pre-doctoral grants: from the Basque Government (I.E.), and from the Ministry of Education of Spain (A.P.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Miguel A. Alfonso-Sánchez and Ibone Espinosa contributed equally to this work.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Alfonso-Sánchez, M.A., Espinosa, I., Gómez-Pérez, L. et al. Tau haplotypes support the Asian ancestry of the Roma population settled in the Basque Country. Heredity 120, 91–99 (2018). https://doi.org/10.1038/s41437-017-0001-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-017-0001-x
This article is cited by
-
Neolithic expansion and the 17q21.31 inversion in Iberia: an evolutionary approach to H2 haplotype distribution in the Near East and Europe
Molecular Genetics and Genomics (2023)
-
On the Trail of Spatial Patterns of Genetic Variation
Evolutionary Biology (2022)