Abstract
Purpose
Elucidating the genetic architecture underlying autism spectrum disorder (ASD) will aid in the understanding of its genetic etiology and clinical diagnosis.
Methods
A comprehensive set of coding de novo variants (DNVs) from 4504 trios with ASD and 3012 control/sibling trios from several large-scale sequencing studies were collected and combined. Multiple in-depth analyses including DNVs burden, clinical phenotypes, and functional networks underlying the combined data set were used to evaluate the nonrandom occurrence of multiple extreme DNVs (loss-of-function and damaging missense variants) in the same patients.
Results
We observed a significant excess of multiple extreme DNVs among patients with ASD compared with controls. Meanwhile, patients with ASD carrying 2+ extreme DNVs had significantly lower IQs than patients carrying 0 or 1 DNV. Moreover, much closer functional connectivity than expected was observed among 2 or more genes with extreme DNVs from the same individuals. In particular, we identified 56 key genes as more confident ASD genes compared with other known ASD genes. In addition, we detected 23 new ASD candidate genes with recurrent DNVs, including VIP, ZWILCH, MSL2, LRRC4, and CAPRIN1.
Conclusions
Our findings present compelling statistical evidence supporting an oligogenic model and provide new insights into the genetic architecture of ASD.
Similar content being viewed by others
INTRODUCTION
Autism spectrum disorders (ASDs) are a group of neurodevelopmental disorders characterized by significant impairments in social communication, interactions, and repetitive behaviors.1,2 ASD exhibits high heritability with substantial clinical and genetic heterogeneity.3,4,5 Recently, de novo variants (DNVs) have been considered significant contributors to the etiology of sporadic ASD in several trio-based studies and have become an important resource for the discovery of pathogenic genes.6,7,8,9,10
The ability to decipher the genetic etiology of ASD remains challenging due to its complex genetic architecture.3,11 A single proband often carries two or more gene-disruptive DNVs in different genes.7,10,12 For example, two genes (CHD8 and CUBN) with truncating DNVss have been found in an ASD proband with a severe clinical phenotype in a cohort of 189 patients, including 19 patients harboring multiple genes with loss-of-function (LoF) or damaging missense (D-Mis) DNVs (defined as extreme DNVs; Supplementary Methods).10 In addition, a trio-based exome sequencing study has found that one proband carrying a SCN2A splicing DNM was also a carrier of a second extreme DNV in PTPN14.7 Moreover, genome sequencing in 32 families with ASD has revealed one patient in whom three candidate genes were affected by extreme DNVs, including a de novo frameshift insertion in USP54 and damaging missense DNVs in KIAA1217 and FAT3.13
However, the rarity of coding DNVs and limited cohort sizes suggest that it is very difficult to provide sufficient statistical support to clarify whether the phenomenon of multiple DNVs occurring in a proband has a larger effect on ASD or leads to a more severe phenotype in patients than only one or no DNV.3,14 An alternative solution is a combined analysis of DNVs from collective large-scale sequencing studies, which may be a productive approach to provide further insights into the genetic landscape and pathophysiology of ASD.3,15 In this study, by utilizing the combined DNV data sets, we first analyze the prevalence of individuals with one, two, or more DNVs in ASD and controls to find the enrichment of individuals with more variants in ASD. Then we investigate the relationship between number of DNVs and the IQ of individuals, and find more mutated genes correspond to a lower IQ in a patient. By evaluating functional connectivity of multiple mutated genes within the same patient, including protein interaction and coexpression relationship, we observed a clear enrichment signal in pairs of multiple mutated genes compared with the background or other random pairs of mutated genes (Fig. S1). This evidence supports the existence of an oligogenic model in ASD, which can improve our understanding of the genetic etiology of ASD.
MATERIALS AND METHODS
Data collection and annotation
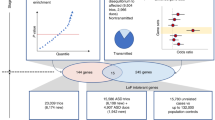
As the workflow indicates (Fig. S1), a total of 36,262 DNVs from 4504 ASD trios and 41,577 DNVs from 3012 control trios were collected from multiple massively parallel sequencing studies in the NPdenovo database (Table S1).16 The institutional review board of Wenzhou Medical University approved our study as human subjects exempt because only de-identified data was accessed. All variants were annotated with the ANNOVAR software.17 The details are described in the Supplementary Materials and Methods.
Prioritization of candidate genes and identification of ASD-associated genes
A Bayesian model tool called transmission and de novo association (TADA)18 was used to prioritize the candidate genes by combining LoF and D-Mis DNVs and four background DNV rates (DNVRs), including sequence context (DNVR-SC), GC content (DNVR-GC), multiple factors (DNVR-MF), and local DNA methylation level (DNVR-DM) retrieved from the mirDNMR database.19 TADA P values were corrected with the false discovery rate (FDR) approach to obtain the TADA q values.
Gene conservation and damage assessment
The enrichment levels of ASD-associated genes in the targets of CHD820 and the gene-level constraints, including residual variation intolerance scores (RVIS), probability of LoF intolerance scores (pLI) in ExAC, and haploinsufficiency, were analyzed as described in the Supplementary Materials and Methods.
Functional network analysis and spatiotemporal expression patterns
The expression data of candidate and key genes in human brain were acquired from the Human Brain Transcriptome database (HBT, http://hbatlas.org/)21 and BrainSpan database (http://www.brainspan.org/).22 The protein–protein interaction (PPI) networks, coexpression networks, and clustering analysis were performed as described in the Supplementary Materials and Methods.
Gene Ontology enrichment analysis
The Gene Ontology (GO) enrichment analysis was performed using WebGestalt (http://www.webgestalt.org/) and provided results on biological processes, cellular components, and molecular functions. Moreover, a plug-in of Cytoscape, ClueGO v2.3.3, was used to perform a more detailed enrichment analysis.
Identification of an oligogenic model and potential genetic rules in ASD
All patients and controls were divided into three groups according to the number of genes (0, 1, and 2+) carrying protein-coding DNVs, extreme DNVs, and nonextreme DNVs. The analyses of an oligogenic model, phenotype information, and functional connectivity among the multiple mutated genes were described in the Supplementary Materials and Methods.
RESULTS
Identification of new ASD candidate genes using combined data set
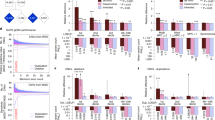
We have collected 4504 ASD trios together with 3012 unaffected control/sibling trios from currently trio-based exome/genome sequencing studies (Table S1). After eliminating noncoding variants, 5123 coding DNVs were retrieved, including 4705 single-nucleotide variants (SNVs) and 418 insertions/deletions (indels) (Table S2). The rate of coding DNVs was approximately 1–1.19 events per trio across five separate studies that conducted sequencing of more than 50 trios, and 1.14 DNVs per trio in the combined data (Tables S1 and S3). The DNV rate in our combined data set is comparable with each study (Padjust >0.05, two-tailed Poisson rate test corrected by FDR; Table S3). In addition, no significant difference in the rates of synonymous SNVs and nonframeshift indels was observed between all cases and controls (Padjust = 0.37, two-tailed Poisson rate test corrected by FDR; Table S3). To prioritize ASD-associated genes, we utilized the TADA18 to identify associated genes based on four different background de novo variant rates (DNVRs) including DNVR-GC, DNVR-SC, DNVR-MF, and DNVR-DM (Supplementary Methods). As a result, we identified 98 strong associated genes with q values <0.1 and 246 associated genes with q values <0.3 in ASD cohorts, but only one gene (SH3D19) was found with q value <0.3 in the control using the same detection strategy (Fig. 1a, Fig. S2a, b). Among the 98 genes with q values <0.1, 80 genes (80/98, 81.6%) were simultaneously identified when using at least three of the different background DNVRs, which will be used for subsequent analysis as ASD-associated genes (Fig. S2c). In addition, 174 were also obtained from the 246 genes with q values <0.3 (Fig. S2d).
Identification of known and strong candidate genes in autism spectrum disorder (ASD). (a) Genes with TADA q values <0.1 were shared by four background de novo variant rate (DNVR) methods in 4504 trios: sequence context (DNVR-SC), GC content (DNVR-GC), multiple factors (DNVR-MF), and local DNA methylation level (DNVR-DM). (b) Intersections between gene sets of SFARI-1&2, TADA q values <0.3 and <0.1 (Sanders et al.23), and TADA q values <0.1 (this study). (c) A scatter diagram for candidate-23 genes. Extreme de novo variants (DNVs), loss-of-function and damage missense DNVs; expected DNVR, average of four background DNVRs; q value, average of TADA q value based on four background DNVRs.
For comparison, we compiled a known ASD gene set containing 192 genes retrieved from the SFARI Gene Database (categories 1 and 2 only) and a high-risk ASD gene set identified from a recent gene prioritization study.23 We found 23 new candidate genes from 80 strong associated genes (q < 0.1) and 88 potential candidate genes from the 174 associated genes (q < 0.3) by excluding the known ASD genes (Fig. 1b and Fig. S2e). Among 23 genes, 6 (P4HA2, SETBP1, KIRREL3, LRRC4,SARM1, and MYO1E) were affected by 3 extreme DNVs, and the other 17 genes contained 2 extreme DNVs. In particular, VIP was the gene with the highest q value and contained 2 LoF DNVs (TADA q value = 7.7 × 10−3) (Fig. 1c, Fig. S2f, Tables S4 and S5).
Evolutionary constraint and functional characteristics of ASD candidate-23 genes
We performed a series of functional analyses to evaluate the possible contribution of the candidate-23 genes to ASD (Table S6). While CHD8 target genes were frequently reported as associated with ASD,7,20 8 of the 23 candidate genes are among theCHD8 targets (P = 5.47 × 10-3, one-tailed Fisher’s exact test). The RVIS for candidate genes were significantly enriched in the top 25th percentile (P = 2.17 × 10−5), which indicates them as functional important genes. Moreover, there were 12 candidates with pLI ≥ 0.9 (P = 1.87 × 10-4) and ten genes with the 25th percentile of missense Z-scores (Z ≥ 1.74) in ExAC (Supplementary Methods). Notably, 22 candidate genes have been identified as haploinsufficient genes, and 11 genes were in the top quartile of haploinsufficiency percentile (P = 0.01).
To investigate the expression profiles of the candidate-23 genes during human brain development, we analyzed spatial and temporal gene expression data from HBT21 and found that 8 genes with mean expression levels across all 16 brain regions were in the top 20% of highly expressed genes in HBT, which is unexpected by chance (P = 0.02, top 20% of expression level = 8.302; Fig. S3a). Interestingly, the 22 genes with expression data in HBT showed a significant enrichment of expression in the top 20% of highly expressed genes before birth and displayed the obvious hierarchical clustering in the neocortex and other brain regions (Fig. S3b). Similar analyses were performed based on RNA-seq data from BrainSpan22 and showed significant enrichment for in total 23 candidate genes in the top 20% of all genes before birth consistent with the analyses in HBT (Fig. S4). Furthermore, fold change analysis in BrainSpan showed that 16 of 23 candidate genes with the fold change (prenatal/postnatal) of mean expression levels were >1 (Table S7).
We then constructed a genetic interaction network using GeneMANIA (Supplementary Methods), which showed more credible genetic interactions between ASD candidates identified here and known ASD-associated genes than expected by chance (P = 0.026 for nodes, P = 2.8 × 10-4 for edges; 100,000 permutation tests; Fig. 2a). Notably, an analysis of relationships among genes using the coexpression network from HBT showed that the expression levels of ASD candidates were highly correlated with known ASD-associated genes (absolute r ≥ 0.7; P = 8.0 × 10−3 for nodes, P = 5.8 × 10−4 for edges; Fig. 2b). Similarly, new ASD candidates also revealed the high correlation with known ASD-associated genes based on the coexpression network from the BrainSpan database (absolute r ≥ 0.7;P = 1.0 × 10−5 for nodes, P = 1.0 × 10−5 for edges; Fig. S5). Furthermore, GO enrichment analysis of the 80 high-confidence ASD-associated genes (TADAq value <0.1) revealed multiple biological processes and cellular components containing the candidates associated with the nervous system (Fig. 2c and Table S8), such as nervous system development (GO:0007399) (included 5 candidate genes in 26 enriched ASD genes), chromosome organization (GO:0051276) in biological processes, and neuron projection (GO:0043005) in cellular components.
Network and Gene Ontology (GO) enrichment analysis of candidate-23 genes. (a) Genetic interaction network of 23 candidates in GeneMANIA. Nodes denote genes and gray represents the strong known autism spectrum disorder (ASD) genes from the gene sets of SFARI-1&2 and TADA q values <0.1 (Sanders et al.23). Edges denote the interactions, and the thickness denotes the degree of connectivity. (b) Coexpression network analysis of the 23 candidates with the strong known ASD genes in the Human Brain Transcriptome database (HBT). The red edges indicate positive coefficients, and the blue indicates negative coefficients. (c) GO enrichment analysis of 80 high-confidence ASD genes (q < 0.1) using WebGestalt. The top 5 q values for GO terms were selected. The number above each bar represents the number of 23 candidate genes / the number of enriched ASD candidate genes for this GO term. Permutation tests of network genes and connections were performed using 100,000 iterations.
Occurrences of the oligogenic model in ASD
In previous studies, the overall DNVR is elevated in patients with ASD compared with normal individuals.8,10,24 Based on an in-depth analysis of coding DNVs from this combined data set, we found significantly more individuals who carried multiple genes (≥2 or 2+) with DNVs in ASD cohorts than controls (odds ratio [OR] = 1.3, P = 2.8 × 10−6, one-tailed Fisher’s exact test), but no differences for individuals carrying 0 or 1 genes with DNVs (Fig. 3a). The analysis was restricted to extreme DNVs and revealed a much more significant enrichment trend in individuals with ASD who carried 1 or 2+ mutated genes (OR = 1.4, P = 1.4 × 10−9 for 1 gene; OR = 2.0, P = 7.5 × 10−10 for 2+genes). Consistently, by comparing the occurrence of 0, 1, or 2+ genes with extreme DNVs (extreme genes) in ASD patients using control as baseline, we also found that the patients carrying 2+ extreme genes were significantly more than in those carrying 0 or 1 extreme genes (2+ vs. 0: OR = 2.2, P = 3.3 × 10−12 and 2+ vs. 1: OR = 1.5,P = 5.1 × 10−4; Fig. 3b). When considering nonextreme DNVs (coding DNVs that are not classified as extreme), the enrichment in individuals carrying 2+ genes in ASD cohorts was still significant, although to a lesser extent than in patients carrying 2+ extreme DNVs, but it was not significant for patients carrying 0 or 1 gene with DNVs (Fig. S6a, b). We also investigated the occurrence of 0, 1, or 2+ genes with extreme DNVs in 2508 patients with ASD in the largest subset data sets from one study6 (Fig. S7a, b), the results were consistent with the observation from the combined data set, which indicates the DNV occurrence differences were not caused by data from different studies. Therefore, the observation of the strong enrichment of 2+ genes with extreme DNVs in ASD patients suggests the existence of oligogenic model in a fraction of patients with ASD.
The oligogenic model in 4504 autism spectrum disorder (ASD) trios. (a, b) The oligogenic model influences the burden of 4504 patients compared with 3012 controls in each and pairs of groups. P values and odds ratios (ORs) (95% lower confidence interval [CI]) were calculated by one-tailed Fisher’s exact test. (c) Violin plot shows the distribution of three types of IQ in patients. P values were calculated by two-tailed pairwiset test, and false discovery rate (FDR) correction was used to obtain Padjust.FSIQ full-scale IQ,NVIQ nonverbal IQ,VIQ verbal IQ. (d) The oligogenic model does not influence the burden of female compared with male individuals.P values and ORs (95% CI) were calculated by two-tailed Fisher’s exact test. Coding: genes with coding variants; extreme: genes with loss-of-function (LoF) and damaging missense (D-Mis) variants; “0,” “1,” and “2+” mean that the individuals contain 0, 1, or 2+ genes with coding or extreme de novo variants (DNVs).
To evaluate the relationships between one important ASD phenotypic feature (IQ) and the number of DNVs within each individual, we compared IQ among patients with ASD who carried 0, 1, and 2+ extreme genes. As a result, a clear and statistically significant relationship between the number of mutated genes and full-scale IQ (FSIQ) was observed, with a lower FSIQ corresponding to the presence of a higher number of mutated genes (Fig. 3c). Although patients who carried DNVs showed a significant overall decrease in FSIQ compared with patients without DNVs,25,26 the FSIQ of patients carrying 2+ mutated genes was significantly less than that of patients carrying 0 or 1 mutated genes (Padjust for 2+ vs. 0: 4.7 × 10−3, Padjust for 2+ vs. 1: 0.01, FDR-corrected two-tailed pairwise t test; Fig. 3c). Moreover, similar relationships between the number of mutated genes and nonverbal IQ (NVIQ) or verbal IQ (VIQ) were also found. Notably, the NVIQ of patients carrying 2+ mutated genes was significantly less than that of patients harboring no mutated genes (Padjust = 2.2 × 10−3) and was slightly less than that of patients with only 1 mutated gene (Padjust = 0.02); similar results were observed for VIQ (2+ vs. 0: Padjust = 0.02; 2+ vs. 1: Padjust = 0.02). In addition, similar trends for FSIQ, NVIQ, and VIQ were observed in the subset data sets from one study6 (Fig. S7c).
Given the sex differences in ASD and the hypothetically greater genetic burden in female patients,25,27 we further tested whether sex differences existed. The female:male ratio among patients carrying 2+ genes with coding or extreme DNVs was not significantly different from the ratio for patients carrying 0 or 1 mutated genes (P > 0.05/4 for all comparisons, two-tailed Fisher’s exact test), indicating that there was no significant difference between the sexes (Fig. 3d).
Potential genetic rules underlying the oligogenic model of ASD
We further investigated the relationship between the number of mutated known ASD genes in each patient to explore the relevance of occurrence of multiple DNVs in one patient to the ASD diagnosis (Tables S9 and S10). Because extreme DNVs tend to be disease-causal in ASD,4,5,25 we performed a comparative analysis of extreme genes and known ASD-associated genes. We found that 190 (V+VII in Fig. 4a) of 1062 mutated genes from patients carrying 1 extreme gene, and 114 (VI+VII in Fig. 4a) of 579 mutated genes from patients carrying 2+ extreme genes were reported as known ASD-associated genes. Notably, 56 known ASD-associated genes (gene-56, VII in Fig. 4a) and 36 non–ASD-associated genes (gene-36, IV in Fig. 4a) were found at the intersection between 1 and 2+ extreme gene sets (Table S9). However, almost no intersections were observed between ASD-associated genes and the 1 and 2+ extreme gene sets in controls, and the overlap between the 1 and 2+ extreme gene sets contained very few genes (Fig. S8). Interestingly, the proportion of known ASD-associated genes among 2+ extreme genes was higher than among 1 extreme gene (19.7% vs. 17.9%), although this difference was not statistically significant (Fig. 4b). Furthermore, a slight increase in the number of extreme DNVs in ASD-associated genes was observed in carriers of 2+ extreme genes compared with carriers of 1 extreme gene (2+ vs. 1: OR = 1.2, P = 0.07, two-tailed Fisher’s exact test). Consistent with this result, we observed a slightly increased burden in patients harboring 2+ ASD-associated genes with extreme DNVs (2+ vs. 1: OR = 1.2, P = 0.07, Fig. 4b).
Potential genetic rules underlying the oligogenic model of autism spectrum disorder (ASD). (a) Intersections between 1 and 2+ extreme gene sets in patients and ASD-associated genes (SFARI-1&2, TADA q value <0.3 [Sanders et al.23] and TADA q value <0.3 [this study]). (b) The percentage of ASD-associated genes, extreme de novo variants (DNVs) within ASD-associated genes, and individuals carrying mutated ASD genes among 1 extreme (I+V+IV+VII) and 2+ extreme (II+VI+IV+VII) gene sets. P values and odds ratios (ORs) (95% confidence interval [CI]) were calculated by two-tailed Fisher’s exact test. (c) The percentage of patients carrying 1 extreme (I+V) and 2+ extreme (II+VI) genes classified by different ranges of full-scale IQ (FSIQ). The total distribution of FSIQs was tested by Pearson’s chi-squared test. P values and ORs (95% CI) for each range of FSIQ were calculated by two-tailed Fisher’s exact test. (d, e) Boxplot of protein–protein interaction (PPI) scores in STRING and Pearson’s coexpression correlation coefficients in BrainSpan, each divided into four groups: background, 1 extreme (I+V), random 2+ extreme (II+VI, based on total genes), and pairs of 2+ extreme (II+VI, based on pairs of genes in each individual). P values were calculated by two-tailed Wilcox rank sum test, andPadjust was obtained based on false discovery rate (FDR) correction.
Because IQ values among ASD patients vary within a large range from below 60 to above 1009,28 and IQ values among the patients carrying 2+ mutated genes were generally lower than those carrying 1 mutated gene (Fig. 3c), we sought to address the effect of extreme DNVs on IQ (Table S10). A difference in the distribution of FSIQ was observed between patients carrying 2+ extreme genes and those carrying 1 extreme gene (Ptotal = 0.03; Fig. 4c). Specifically, FSIQ values for patients carrying 2+ extreme genes were significantly absent from the upper range of >100 (2+ vs. 1: OR = 0.5, Padjust = 0.029) but were enriched in the lower range of ≤60 (2+ vs 1: OR = 1.7, Padjust = 0.035). In addition, no obvious difference in the FSIQ distribution in the normal FSIQ range of 60-100 was observed among patients carrying 1 or 2+ extreme genes.
Given that the occurrence of 2+ genes with extreme DNVs in the same individual was greater in the ASD cohort and may decrease the patient’s IQ, we sought to investigate the possible functional connectivity among the multiple mutated genes detected within the same patients. By evaluating PPI relationship scores, we observed significant enrichments in pairs of 2+ extreme genes compared with the background (Padjust = 5.0 × 10−8, FDR-corrected two-tailed Wilcoxon rank sum test) and random pairs of 1 (Padjust = 8.8 × 10−8) or 2+ (Padjust = 5.1 × 10−7) extreme genes (Fig. 4d). For coexpression correlation coefficients, significant enrichment was observed in pairs of 2+ extreme genes compared with the background (Padjust = 3.7 × 10−3; Fig. 4e).
Identification of key ASD genes
While known ASD-associated genes were collected from different sources with different evidence with different strength, here we are trying to filter for more confident known ASD genes or key ASD genes compared with other known ASD genes using DNV frequency data. Gene-56 contained extreme DNVs in an average of 3.2 individuals per gene, which is a significantly higher value than other gene groups, such as gene-134 (genes in only 1 extreme gene set, V in Fig. 4a) with 1.5 individuals per gene (gene-56 vs. gene-134: P = 6.1 × 10−13; two-tailed Poisson rate test) and gene-58 (genes in only 2 extreme gene sets, VI in Fig. 4a) with 1.1 individuals per gene (gene-56 vs. gene-58: P = 3.1 × 10−14; Fig. S9a). Moreover, almost all genes (52/56) were strongly correlated with ASD evaluated using TADA (q < 0.3) (Fig. 5a). We further assessed the gene-level constraints of gene-56 using RVIS and pLI values, and found that more than 30 constrained genes in gene-56 group fell in the “hot zone” (RVIS ≤25th percentile and pLI scores of ≥0.9) compared with the background (Fig. S9b and Fig. S10) and other known ASD gene sets, including gene-134 (P = 5.2 × 10−3), gene-58 (P = 2.5 × 10−5), and gene-32 (genes harboring no extreme DNVs, III in Fig. 4a; P = 0.014; Fig. 5b). The above observations indicated that the genes of gene-56 are more vulnerable to stringent evolutionary selection against functional genetic variation than background or the other known ASD genes.
Key genes implicated in autism spectrum disorder (ASD) as identified by the oligogenic model. (a) Scatter diagram showing the 56 ASD risk genes. (b) The comparison between the enrichment levels of gene-56 and background, gene-836, gene-429, gene-134, gene-58, and gene-32 in residual variation intolerance (RVIS) and probability of loss-of-function intolerance (pLI) score. P values and odds ratios (ORs) (95% confidence interval [CI]) were calculated by two-tailed Fisher’s exact test. (c) Biological process terms of gene-56 identified using the ClueGO enrichment analysis. (d) Protein–protein interaction (PPI) network between gene-56 and ASD-associated genes in STRING. The main biological process terms were identified by Gene Ontology (GO) enrichment analysis (C1–C4). A permutation test of network genes and connections was performed using 100,000 iterations.DNV de novo mutation,DNVR de novo variant rate.
To characterize the functional classification of gene groups, we performed ClueGO and found 39 significantly enriched gene sets converging on six major modules in 66.07% (37/56) of the gene-56 group (enrichment q value <1.0 × 10−3; FDR-corrected hypergeometric test) (Fig. 5c). However, no significant GO enrichment was observed in other known ASD genes like the gene-58 and gene-134 sets. As expected, genes in gene-56 are mainly enriched in synaptic formation, transcriptional regulation, and chromatin remodeling pathways that are known in ASD (Fig. 5c). For instance, the top two significantly enriched gene sets were involved in regulation of postsynaptic membrane potential (GO:0060078) (9 genes, q = 3.9 × 10−8) and covalent chromatin modification (GO:0016569) (12 genes, q = 5.9 × 10−6) (Table S11). Furthermore, we also discovered new gene sets enriched in membrane depolarization during action potential, representing potential functional groups that may be involved in ASD (Fig. 5c and Table S11).
We further assessed the functional relevance of gene-56 genes to other known ASD-associated genes using PPI. As a result, a single significant network with high-confidence interconnections was derived from 102 of the 226 ASD-associated genes (P = 0.05 for nodes, P = 0.0016 for edges; Fig. 5d), forming four main functional subclusters, consistent with previous findings.6,7 Strikingly, many of the known recurrent hit genes in gene-56 exhibited high connectivity to other ASD-associated genes in these clusters, such as CHD8 and ASH1L, which have roles in chromatin remodeling;SCN2A, which is involved in regulating action potentials; and SYNGAP1 and PTEN, which are implicated in synaptic function.
DISCUSSION
Previously, a single ASD proband often carried multiple deleterious DNVs; whether this phenomenon occurs by chance is difficult to assess due to the limited individual sizes.7,9,13 Our analysis revealed nonrandom occurrence of this phenomenon using large comprehensive data sets, and provided multiple lines of evidence supporting the existence of an oligogenic model. Based on burden analysis, we observed a significant enrichment of multiple genes with extreme DNVs compared with patients with no or only one mutated gene and the controls. Remarkably, the investigation of clinical phenotypes suggested that an increased number of extreme DNVs in a proband decreases the IQ. For example, the IQ values (IQ = 39, VIQ = 21 and NVIQ = 60) of proband 13012 harboring a de novo DIP2A insertion and a damaging de novo missense SNV in RELN are lower than proband 13106 (IQ = 100, VIQ = 140 and NVIQ = 79) carrying only one de novo nonsense variant in DIP2A (Table S2 and Table S10). By examining the proportion of ASD-associated genes present in patients, we observed an increased burden of individuals with ASD who harbored multiple genes with extreme DNVs. Furthermore, the analysis of PPI and coexpression networks among extreme genes showed higher functional connectivity in the pairs of 2+ extreme genes group, thus further establishing a potential link between an oligogenic model and ASD.
An oligogenic model has been proposed in a fraction of ASD cases with different forms of genetic variations.9,10,12,25,29,30 Briefly, the observation of inherited heterozygous variants of multiple known autism susceptibility genes in 23 of 339 ASD probands has implied the possibility of existence of the oligogenic signal.30 In addition, the occurrence of multiple rare copy-number variants (CNVs) in several ASD cases displaying increasing phenotypic severity may indicate the additive or epistatic effects of this model.29,31 Specifically, trio-based exome/genome sequencing studies have shown that the association of de novo events with inherited deleterious variants may contribute to ASD in a few individuals.9,10,25 However, all these observations have relied on a limited number of ASD cases and lack sufficient statistical power to confirm the model. We showed clear evidence for the involvement of the oligogenic model in patients with ASD using large-scale exome/genome sequencing data; compared with the previous methods, we only focused on DNVs in coding regions to better document an oligogenic model of ASD, because these DNVs are more accessible in a large number of patients and controls and are more likely to be deleterious. In fact, the rate of individuals affected by multiple genes with extreme DNVs is estimated to be approximately 6.3% among sporadic ASD cases in our combined data set (Table S12). In addition, by investigating the previous coding DNV data sets,6,7 we did not detect a substantial difference in the proportion of individuals carrying multiple genes with extreme DNVs (Ptotal = 0.95 and Padjust = 1; Table S12). The remaining probands may carry a combination of coding variants and other forms of genetic variation, such as noncoding DNVs, CNVs, structural variants, or rare inherited variants in coding regions.
Twenty-three novel strong candidates harboring recurrent extreme DNVs were discovered. A targeted sequencing study has identified 91 neurodevelopmental disorder risk genes between ASD and DD,4 including our candidates HIVEP3, NFIA,SETBP1, and ZC3H4. Another study performing genome sequencing on 2626 ASD individuals has also identified 18 candidate genes for ASD,5 including our candidatesDIP2C and PHF3. Furthermore, the other genes also show strong associations with ASD. For example, CAPRIN1 with two LoF DNVs in this study (pLI = 0.9998, RVIS percentile = 21.20%) regulates the transport and translation of the messenger RNAs (mRNAs) involved in synaptic plasticity in neurons and cell proliferation. Additionally, heterozygous knockout of CAPRIN1 decreases social interactions, thus resulting in an ASD-like behavior.32DVL3 encoding a cytoplasmic phosphoprotein that regulates cell proliferation carried two LoF DNVs in two ASD patients (pLI = 0.9954, RVIS percentile = 9.17%). A mouse model with DVL1 and DVL3 knockout has linked fetal brain development with adult ASD-like behaviors,33 and the DVL3 PDZ domain is associated with typical autism by influencing the SHANK3 PDZ domain.34
The genetic landscape of ASD has been estimated to involve 400–1000 genes conferring risk.3,35 However, which ASD-associated genes are the core genes that contribute to ASD remains unclear. In this study, 56 ASD-associated genes were critically relevant to ASD and represented key ASD causal genes based on their contributions to the aberrant phenotype using evidence from function-related analyses of biological processes, evolutionary constraint, gene coexpression, and protein interaction networks. Moreover, several genes in gene-56, such as CHD8, CHD2,SCN1A, SYNGAP1, SHANK3, and PTEN, have well-documented roles in ASD pathogenesis, on the basis of highly recurrent extreme DNVs4,5,6,7,26 and a series of model systems, such as human stem cells,36 mice, Drosophila,4 and zebrafish.37 The other genes of gene-56 might be involved in a variety of neuropsychiatric disorders or phenotypes related to neurodevelopment. In addition, several genes of gene-36 with recurrent extreme damaging variants, such as RAI1,38NLGN2,39 and RGMA,40 have been found to probably contribute to ASD according to recent findings in animal models. Altogether, our findings highlight the importance of gene-56 and provide considerable clues for studies aiming to obtain a deeper understanding of ASD pathogenesis and more effective diagnostics and therapies.
Change history
01 August 2019
In the original version of the published Article the word "mutation" was used throughout the text. The ACMG changed its terminology from "mutation" to "variant" prior to the publication of the Article and the requested text changes were not incorporated by the typesetter. The PDF and HTML versions of the Article have now been modified accordingly.
References
Constantino JN, Charman T. Diagnosis of autism spectrum disorder: reconciling the syndrome, its diverse origins, and variation in expression. Lancet Neurol. 2016;15:279–291.
Lord C, Bishop SL. Recent advances in autism research as reflected in DSM-5 criteria for autism spectrum disorder. Annu Rev Clin Psychol. 2015;11:53–70.
Vorstman JAS, Parr JR, Moreno-De-Luca D, et al. Autism genetics: opportunities and challenges for clinical translation. Nat Rev Genet. 2017;18:362–376.
Stessman HA, Xiong B, Coe BP, et al. Targeted sequencing identifies 91 neurodevelopmental-disorder risk genes with autism and developmental-disability biases. Nat Genet. 2017;49:515–526.
RK CY, Merico D, Bookman M, et al. Whole genome sequencing resource identifies 18 new candidate genes for autism spectrum disorder. Nat Neurosci. 2017;20:602–611.
Iossifov I, O’Roak BJ, Sanders SJ, et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature. 2014;515:216–221.
De Rubeis S, He X, Goldberg AP, et al. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature. 2014;515:209–215.
Sanders SJ, Murtha MT, Gupta AR, et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature. 2012;485:237–241.
O’Roak BJ, Vives L, Girirajan S, et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature. 2012;485:246–250.
O’Roak BJ, Deriziotis P, Lee C, et al. Exome sequencing in sporadic autism spectrum disorders identifies severe de novo mutations. Nat Genet. 2011;43:585–589.
de la Torre-Ubieta L, Won H, Stein JL, Geschwind DH. Advancing the understanding of autism disease mechanisms through genetics. Nat Med. 2016;22:345–361.
Turner TN, Coe BP, Dickel DE, et al. Genomic patterns of de novo mutation in simplex autism. Cell. 2017;171:710–22 e712.
Jiang YH, Yuen RK, Jin X, et al. Detection of clinically relevant genetic variants in autism spectrum disorder by whole-genome sequencing. Am J Hum Genet. 2013;93:249–263.
Ronemus M, Iossifov I, Levy D, Wigler M. The role of de novo mutations in the genetics of autism spectrum disorders. Nat Rev Genet. 2014;15:133–141.
Acuna-Hidalgo R, Veltman JA, Hoischen A. New insights into the generation and role of de novo mutations in health and disease. Genome Biol. 2016;17:241.
Li J, Cai T, Jiang Y, et al. Genes with de novo mutations are shared by four neuropsychiatric disorders discovered from NPdenovo database. Mol Psychiatry. 2016;21:290–297.
Wang K, Li M, Hakonarson H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010;38:e164.
He X, Sanders SJ, Liu L, et al. Integrated model of de novo and inherited genetic variants yields greater power to identify risk genes. PLoS Genet. 2013;9:e1003671.
Jiang Y, Li Z, Liu Z, et al. mirDNMR: a gene-centered database of background de novo mutation rates in human. Nucleic Acids Res. 2017;45:D796–D803.
Cotney J, Muhle RA, Sanders SJ, et al. The autism-associated chromatin modifier CHD8 regulates other autism risk genes during human neurodevelopment. Nat Commun. 2015;6:6404.
Kang HJ, Kawasawa YI, Cheng F, et al. Spatio-temporal transcriptome of the human brain. Nature. 2011;478:483–489.
Hawrylycz MJ, Lein ES, Guillozet-Bongaarts AL, et al. An anatomically comprehensive atlas of the adult human brain transcriptome. Nature. 2012;489:391–399.
Sanders SJ, He X, Willsey AJ, et al. Insights into autism spectrum disorder genomic architecture and biology from 71 risk loci. Neuron. 2015;87:1215–1233.
Awadalla P, Gauthier J, Myers RA, et al. Direct measure of the de novo mutation rate in autism and schizophrenia cohorts. Am J Hum Genet. 2010;87:316–324.
Krumm N, Turner TN, Baker C, et al. Excess of rare, inherited truncating mutations in autism. Nat Genet. 2015;47:582–588.
O’Roak BJ, Stessman HA, Boyle EA, et al. Recurrent de novo mutations implicate novel genes underlying simplex autism risk. Nat Commun. 2014;5:5595.
Jacquemont S, Coe BP, Hersch M, et al. A higher mutational burden in females supports a “female protective model” in neurodevelopmental disorders. Am J Hum Genet. 2014;94:415–425.
Samocha KE, Robinson EB, Sanders SJ, et al. A framework for the interpretation of de novo mutation in human disease. Nat Genet. 2014;46:944–950.
Nava C, Keren B, Mignot C, et al. Prospective diagnostic analysis of copy number variants using SNP microarrays in individuals with autism spectrum disorders. Eur J Hum Genet. 2014;22:71–78.
Schaaf CP, Sabo A, Sakai Y, et al. Oligogenic heterozygosity in individuals with high-functioning autism spectrum disorders. Hum Mol Genet. 2011;20:3366–3375.
Gau SS, Liao HM, Hong CC, et al. Identification of two inherited copy number variants in a male with autism supports two-hit and compound heterozygosity models of autism. Am J Med Genet B Neuropsychiatr Genet. 2012;159B:710–717.
Ohashi R, Takao K, Miyakawa T, Shiina N. Comprehensive behavioral analysis of RNG105 (Caprin1) heterozygous mice: reduced social interaction and attenuated response to novelty. Sci Rep. 2016;6:20775.
Belinson H, Nakatani J, Babineau BA, et al. Prenatal beta-catenin/Brn2/Tbr2 transcriptional cascade regulates adult social and stereotypic behaviors. Mol Psychiatry. 2016;21:1417–1433.
Saupe J, Roske Y, Schillinger C, et al. Discovery, structure-activity relationship studies, and crystal structure of nonpeptide inhibitors bound to the Shank3 PDZ domain. ChemMedChem. 2011;6:1411–1422.
Mullins C, Fishell G, Tsien RW. Unifying views of autism spectrum disorders: a consideration of autoregulatory feedback loops. Neuron. 2016;89:1131–1156.
SS MS, Magdalon J, Griesi-Oliveira K, et al. Rare RELN variants affect Reelin-DAB1 signal transduction in autism spectrum disorder. Hum Mutat. 2018;39:1372–1383.
Bernier R, Golzio C, Xiong B, et al. Disruptive CHD8 mutations define a subtype of autism early in development. Cell. 2014;158:263–276.
Rao NR, Abad C, Perez IC, et al. Rai1 haploinsufficiency is associated with social abnormalities in mice. Biology (Basel). 2017;6:E25.
Wohr M, Silverman JL, Scattoni ML, et al. Developmental delays and reduced pup ultrasonic vocalizations but normal sociability in mice lacking the postsynaptic cell adhesion protein neuroligin2. Behav Brain Res. 2013;251:50–64.
O’Leary C, Cole SJ, Langford M, et al. RGMa regulates cortical interneuron migration and differentiation. PLoS ONE. 2013;8:e81711.
Acknowledgements
The project was funded by the National Natural Science Foundation of China (31771404 and 31571357) and China Scholarship Council (201608330306).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
The authors declare no conflicts of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Du, Y., Li, Z., Liu, Z. et al. Nonrandom occurrence of multiple de novo coding variants in a proband indicates the existence of an oligogenic model in autism. Genet Med 22, 170–180 (2020). https://doi.org/10.1038/s41436-019-0610-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-019-0610-2
Keywords
This article is cited by
-
Statistical methods for assessing the effects of de novo variants on birth defects
Human Genomics (2024)
-
Truncating PICK1 Variant Identified in Azoospermia Affected Mitochondrial Dysfunction in Knockout Mice
Current Medical Science (2023)
-
Three generation families: Analysis of de novo variants in autism
European Journal of Human Genetics (2023)
-
A cryptic microdeletion del(12)(p11.21p11.23) within an unbalanced translocation t(7;12)(q21.13;q23.1) implicates new candidate loci for intellectual disability and Kallmann syndrome
Scientific Reports (2023)
-
Dystrophin genetic variants and autism
Discover Mental Health (2022)