Abstract
Purpose
Exome sequencing (ES) can rapidly identify disease-causing variants responsible for rare, single-gene diseases, and potentially reduce the duration of the diagnostic odyssey. Our study examines how parents and families value ES.
Methods
We developed a discrete choice experiment (DCE) survey that was administered to parents of children with rare diseases. The DCE included 14 choice tasks with 6 attributes and 3 alternatives. A valuation-space model was used to estimate willingness to pay, willingness to wait for test results, and minimum acceptable chance of a diagnosis for changes in each attribute.
Results
There were n = 319 respondents of whom 89% reported their child had genetic testing, and 66% reported their child had a diagnosis. Twenty-six percent reported that their child had been offered ES. Parents were willing to pay CAD$6590 (US$4943), wait 5.2 years to obtain diagnostic test results, and accept a reduction of 3.1% in the chance of a diagnosis for ES compared with operative procedures.
Conclusion
Timely access to ES could reduce the diagnostic odyssey and associated costs. Before ES is incorporated routinely into care for patients with rare diseases in Canada and more broadly, there must be a clear understanding of its value to patients and families.
Similar content being viewed by others
INTRODUCTION
There are more than 7000 single-gene disorders, and while individually rare, these diseases collectively contribute significantly to morbidity, mortality, and health-care costs.1 Moreover, more than half of patients with rare diseases are undiagnosed and most rare diseases are currently untreatable, creating a tremendous burden on the individual, their families, and society. Patients with rare diseases traditionally experience a prolonged and expensive diagnostic odyssey culminating in a delayed diagnosis or, frequently, no diagnosis at all. Patients and their families have expressed frustration throughout the diagnostic odyssey.2,3 In addition to the emotional burden, this diagnostic odyssey is a financial burden to the health-care system, costing more than US$10,000 per patient.4,5,6,7,8,9
The advent of next-generation sequencing has, and will continue to, revolutionize our ability to identify human disease genes. Current implementation of next-generation sequencing, including exome sequencing (ES), has advanced our ability to rapidly identify disease-causing variants, primarily for rare diseases. In contrast to genome sequencing, which sequences the entire genome of an individual, ES targets the protein coding regions of the genome (less than 2% of the genome).10,11 Several thousand genetic diseases are due to pathogenic variants in protein coding genes.10,11,12 However, availability of clinical ES in Canada is limited outside of the research setting. Timely access to ES has the potential to reduce the time to achieve a diagnosis and the costs associated with the diagnostic odyssey in patients with rare diseases. In a recent study exploring suspected monogenic disorders, the average cost per patient spent on diagnostic investigations and assessments was AU$4733 (US$3692) and the cost per successful diagnosis was AU$27,050 (US$21,099).6 Further, those who had clinical ES had lower average costs per patient and the cost per successful diagnosis was AU$5047 (US$3937).6 The costs reported by Stark et al.6 are higher than the costs of clinical ES in commercial laboratories, which are approximately US$2654 to US$6000 and in some cases up to US$9000 (costs per trio; costs based on personal communications, November 2018, with nine North American commercial diagnostic laboratories).
Previous qualitative and mixed methods research exploring parent attitudes toward obtaining a diagnosis and related diagnostic information found that a diagnosis can have social, personal, and medical impacts for patients.13 Parents value genetic testing and a diagnosis for informing disease treatment, understanding disease etiology, prognosis, and family planning.3,14,15,16,17,18,19,20 However, it remains unclear how parents value ES and a diagnosis when weighing the benefits and risks of diagnostic testing. Although attitude studies provide some insights, preference studies require respondents to explicitly trade off risks and benefits of an intervention to understand how much risk respondents are willing to accept for receiving particular benefits.21,22 Discrete choice experiments (DCEs) are an example of preference elicitation that is increasingly applied in a health context. In a DCE, respondents evaluate and choose between alternative health-care options that are described by common benefits and risks. These benefits and risks are systematically varied between health-care alternatives, such that respondents have to make trade-offs when making a choice.23
Before technologies such as ES are incorporated routinely into care for patients with rare diseases in Canada and more broadly, there must be a clear understanding of its personal and clinical utility, limitations, and associated economic impact.24,25 Without evidence of patient and family value of ES for informing a diagnosis, it is unclear whether this test should be supported and reimbursed in a publicly funded health-care system. This study aimed to fill this gap by determining the personal utility of ES, and, more broadly, the value of a diagnosis for parents of children with rare diseases.
MATERIALS AND METHODS
Study sample
We administered a bilingual (English and French) survey online to a sample of Canadian parents or guardians (>18 years) of children with, or suspected of having, a rare disease. Parents were recruited using three methods: (1) mail-outs at two sites (Alberta Children’s Hospital Clinical Genetics Service, Calgary, AB, Canada and the Children’s Hospital of Eastern Ontario Department of Genetics, Ottawa, ON, Canada) to recruit existing patients; (2) prospective in-clinic recruitment at the same two sites; and (3) postings on websites and social media accounts of rare disease organizations.
For the mail-out, letters were mailed from both sites to parents of children with childhood-onset genetic diseases seen by a medical geneticist after January 2013. Mail-out recruitment was conducted from May 2016 to March 2017. For prospective in-clinic recruitment, letters were provided to parents of children with childhood-onset genetic diseases who were being seen in-clinic by a medical geneticist at both sites. Prospective in-clinic recruitment was conducted from May 2016 to February 2017 at both sites. Parents of children who were seen before January 2013, seen for prenatal or cancer-specific visits, had adult onset diseases, diseases with an incidence greater than 1 in 2000, or multifactorial diseases (i.e., isolated birth defects, idiopathic autism) were not included in recruitment at either site. Nonresponders for the mail-out and prospective in-clinic recruitment were mailed one follow-up letter. Lastly, survey information and instructions for how to participate were posted by eight rare disease organizations using their websites and social media (Facebook and Twitter). These organizations have memberships that include patients, families, and caregivers across Canada who are impacted by a variety of rare diseases. All participants had the option to request a paper survey.
Study design
A discrete choice experiment (DCE) was included in the survey to estimate the value of an ES-based diagnosis for parents of children with rare diseases or suspected genetic conditions.21,23,26,27 In DCEs, respondents evaluate a series of alternative scenarios described by specific attributes (choice tasks) and must choose the scenario they prefer (example in Supplementary Fig. 1).23 Respondent preferences can be estimated from DCE data and expressed as a mathematical function, which is called utility. Measures of value (e.g., willingness to pay, willingness to wait, and minimum acceptable chance of diagnosis) can be derived from the estimated utility.
Attribute development in our DCE was informed by literature review and focus groups (n = 15 parents of children and n = 8 adults with rare diseases).13,28,29 We worked closely with medical geneticists (K.M.B., F.P.B., M.I.) and genetic counselors (T.H., B.M.) to ensure clinical face validity. A draft survey was pretested using one-on-one cognitive interviews with n = 5 parents of children with rare diseases from the Calgary site.30 During these interviews we assessed parents' understanding of the full survey, including background information and the DCE (attributes and levels), as well as asked for their comments and feedback on this information. After modifications based on the pretesting (minor edits to wording of attributes and levels, wording of questions throughout the survey, grammatical edits, and changes to the structure of some survey questions), n = 22 parents of children with rare diseases were recruited from the Calgary site for quantitative pilot testing. After minor modifications based on pilot testing (levels of the negative impact attribute underwent wording edits, and the cost and time attributes had the upper and lower limits expanded to include more levels to cover a broader range of cost and time), the final DCE included six attributes with 4–5 levels each (Table 1). The final survey included the following components: (1) screening questions, (2) background information on genetics and genetic testing, (3) questions exploring experiences with rare diseases or suspected genetic conditions, (4) questions exploring experiences with ES, (5) DCE, (6) questions exploring opinions on costs of obtaining a diagnosis, and (7) demographics. Within our survey we used the terminology “whole exome sequencing” (WES). However, current preferred terminology is “exome sequencing” (ES) and as such we use the term ES throughout.
We used a balanced overlap method to create 50 DCE versions with 12 choice tasks each consisting of three alternatives.26,31 Two repeated choices were added to test preference stability (Supplementary Table 1). Thus, the DCE consisted of 14 choice tasks. We used a dual-response choice format, which allowed respondents to state they would not choose any scenario described in the choice task. This opt-out option was considered a fourth choice alternative. Based on the number of choice tasks and attributes, a sample size of 250 respondents was the estimated minimum required to achieve an acceptable level of statistical precision (standard error of all attributes less than 0.05).26,32,33
Statistical methods
Descriptive statistics were used to examine the demographic and experience variables, and fixed choice tasks. Our analysis of the DCE data defined the utility that each respondent n obtains from a particular health-care option j described in a particular choice set t as:
The parameter \(\alpha _{{\mathrm{opt - out}}}\) is a constant of the opt-out alternative, \(\beta _{1n}\) to \(\beta _{12n}\) are the effects of changes in each attribute on the utility, and \(\varepsilon _{njt}\) is an extreme value distributed error. All attributes, except for chance of diagnosis, time to result, and cost, were dummy coded. This means the estimated parameters are relative to the reference level denoted in Table 1.
We used a valuation-space model34 to estimate three versions of the utility function above: first, utility was measured as respondents’ willingness to pay (WTP) for changes in each attribute. Second, utility was measured as respondents’ willingness to wait (WTW) for their test results for changes in each attribute. Third, utility was measured as respondents’ minimum acceptable chance of a diagnosis (MAC) for changes in each attribute.
We assumed that preferences were randomly distributed. Specifically, preferences for the cost of diagnostic testing, time to obtain an answer from the diagnostic test, and chance of a diagnosis from the test were assumed to be log-normal distributed.35 The remaining parameters were assumed to follow a normal distribution. As a result of this specification, two parameters were estimated for each attribute: mean valuation and standard deviation of the preference distribution. Finally, we assumed that respondents’ valuation of not undergoing any diagnostic test was constant. We assessed model performance using likelihood ratio, the Akaike information criterion (AIC), and the Bayesian information criterion (BIC). All WTP costs are reported in Canadian and US dollars (using 1 March 2017 Bank of Canada exchange rate of 0.75).
We also predicted uptake of ES, other genetic testing methods, “operative procedures” for diagnostic purposes (e.g., biopsies), and a series of other tests and procedures using the WTP utility function. The attributes describing each of the diagnostic testing approaches are in Supplementary Table 2. As a sensitivity analysis, we varied two attribute levels. First, the chance of a diagnosis varied from 35% to 40% for the other genetic testing methods alternative to understand the relative value of ES over single-gene testing, depending on the likelihood of receiving a diagnosis. This sensitivity analysis was used to explore the competition between single-gene testing and ES. Second, a low-cost (CAD$3000/US$2250) and high-cost (CAD$6000/US$4500) scenario were compared for ES, representing the low and high cost range of clinical testing given that ES has limited availability in Canada and all testing is currently sent to commercial diagnostic laboratories in the United States.
Ethics and consent
Informed consent was obtained from participants prior to beginning the survey. The University of Calgary (REB14-1137) and Children’s Hospital of Eastern Ontario (REB14/233X) Research Ethics Boards approved this study.
RESULTS
Respondent characteristics and experiences
Overall, 668 recruitment letters were administered from both sites and the weblink was clicked on 515 times by people accessing it via rare disease organization websites or social media (n = 1183). A total of 704 individuals started and 319 individuals completed the survey (response rate: 60%; completion rate: 27%).
The mean age was 41 years (range: 22–73 years) and 90% were female (Supplementary Table 3). Most (83%) respondents reported at least some postsecondary education, 48% reported full-time employment, 57% resided in large urban population centers (population ≥100,000), 85% were married or in a common-law relationship, 56% reported a household income of ≥CAD$80,000, and 89% identified as white. Some respondents (17%) reported that, in addition to their children having a rare disease or suspected genetic condition, they also had one (Supplementary Table 3). A summary of reported rare diseases is reported in Supplementary Table 4.
Most (81%) respondents reported having one biological child with a rare disease and 14% reported ≥2. The mean age of respondents’ children was 9.8 years (range: 1–44 years) (Supplementary Table 5). Overall, 66% reported their child had received a diagnosis. The mean age of children at time of symptom onset was 1.1 years (range: 0–18 years) and at diagnosis was 4.4 years (range: 0–43 years). The mean duration between year of symptom onset and year of diagnosis was 3.1 years (range: 0–31 years). Of those who reported their child had not received a diagnosis, the mean duration spent seeking a diagnosis is 5.8 years (range: 1–28 years).
Most respondents (89%) reported their child had undergone some form of genetic testing. Almost half (45%) of respondents had previously heard about ES and 75% had a positive attitude toward ES. About one quarter (26%) of respondents reported that their child had been offered ES and of those respondents, 55% had a diagnosis from ES.
Overall DCE findings
The DCE results are presented in Fig. 1–3 and Supplementary Table 6. Regardless of the measure of utility (i.e., WTP, WTW, or MAC), we observed similar patterns of relative attribute importance. Increased knowledge and changes in management or access to services were the most valued attributes. Positive impacts were valued more than negative impacts, and among the negative impacts the worst was receiving a poor prognosis. Respondents preferred ES and genetic tests compared with other tests and procedures or operative procedures.
Willingness to wait for changes in each attribute, in years. Positive values indicate the number of years a respondent is willing to wait for a change in attribute level from the reference level; negative values indicate reduction in waiting time required for a change in attribute level from the reference level.
Minimum acceptable chance of a diagnosis for changes in each attribute, in %. Positive values indicate the reduction in chance of diagnosis that a respondent is willing to accept for a change in attribute level from the reference level; negative values indicate the increase in chance of diagnosis required for a change in attribute level from the reference level.
On average, respondents’ choice of diagnostic tests was affected by all attributes and levels except the possible consequence of lifestyle restrictions. This means that respondents were willing to trade different aspects of diagnostic tests against each other. The specific trade-offs among the attributes are described quantitatively in the following sections.
Respondents’ valuation of diagnostic tests
Respondents were, on average, willing to pay $6590 (US$4943) to have ES (Fig. 1), wait 5.2 years for the test results (Fig. 2), or accept a reduction of 3.1% in the chance of receiving a diagnosis (Fig. 3), when ES was compared with operative procedures. Respondents were, on average, willing to pay $4360 (US$3270) for receiving other genetic testing, wait 3.5 years for test results, or accept a reduction of 2.2% in the chance of receiving a diagnosis rather than an operative procedure. For having a series of other tests and procedures, respondents were, on average, willing to pay $3940 (US$2955), wait 3.1 years for test results, or accept a reduction of 1.8% in the chance of receiving a diagnosis rather than an operative procedure. This means that if all other attributes were equal, respondents preferred ES over other genetic testing methods, followed by a series of other tests and procedures, and operative procedures.
Respondents’ valuation of positive and negative impacts of receiving a diagnosis
For the positive impact of a diagnosis, respondents most valued obtaining knowledge about cause, progression and future with disease, and risk to family. Specifically, respondents were, on average, willing to pay $8050 (US$6038) for such information (Fig. 1), wait 6.2 years for the test results (Fig. 2), or accept a reduction of 3.8% in the chance of receiving a diagnosis (Fig. 3). Respondents valued improvements in disease management, for which they were, on average, willing to pay $7690 (US$5768), wait 5.6 years for the test results, or accept a reduction of 3.5% in the chance of receiving a diagnosis. Respondents were, on average, willing to pay $7510 (US$5633) for access to disease-specific services, programming, and support groups; wait 5.7 years for the test results; or accept a reduction of 3.5% in the chance of receiving a diagnosis.
For the negative impact of a diagnosis, compensating respondents for a poor prognosis required a price reduction of $3440 (US$2580), a decreased waiting time for results of 2.4 years, or an increase in the chance of diagnosis of 1.5%. To compensate respondents for their child being subject to discrimination and/or labeling required a price reduction of $2150 (US$1613), a decreased waiting time for results of 1.3 years, or an increase in the chance of diagnosis of 1.0%.
Respondents’ valuation of chance of a diagnosis, time to diagnostic test results, and cost
Respondents were willing to pay $2280 (US$1710) for every additional percent increase in the chance of receiving a diagnosis (Fig. 1) or wait an additional 1.7 years (Fig. 2). In contrast, the compensation of every additional year that respondents had to wait for their test results required a price reduction of $1860 (US$1395) or an increase in the chance of diagnosis of 0.9% (Fig. 3). Respondents were willing to accept a $1000 price increase if it reduced their waiting time for test results by 1.5 years or increased the chance of a diagnosis by 1.0%.
Heterogeneity in respondents’ valuation of diagnostic test attributes
Respondents’ preferences for the diagnostic test attribute varied significantly (p < 0.05), with the exception of other genetic testing methods. Most (89.8%) respondents preferred ES over operative procedures, while 10.2% of respondents preferred operative procedures (everything else being equal). Most (84.4%) respondents positively valued a series of other tests and procedures compared with operative procedures, while 15.6% had the opposite preference. For negative impact of a diagnosis, 50.4% of respondents positively valued lifestyle restrictions and 49.6% negatively valued lifestyle restrictions.
Probability of diagnostic test uptake
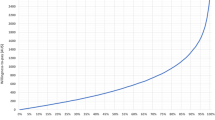
We calculated uptake of different diagnostic tests that a child might undergo for the scenarios outlined in Supplementary Table 2. Given the assumed levels and access to all diagnostic options, we found that nobody would choose operative procedures or a series of other tests and procedures (uptake <0.001%). Figure 4 therefore shows the estimated uptake of ES compared with other genetic testing methods, depending on the chance of diagnosis for other genetic testing methods. In the low-cost scenario, 91% of respondents were predicted to choose ES (with chance of diagnosis constant at 40%) over other genetic testing methods with a 35% chance of diagnosis. The uptake of ES decreased to 46% as the chance of diagnosis using the other genetic testing methods increased to 40%. This means that in the case both genetic diagnostics have similar precision, uptake is predicted to be higher for the other genetic testing methods. Similarly, in the high-cost scenario, 84% of respondents were predicted to choose ES (with chance of diagnosis constant at 40%) over other genetic testing methods with a 35% chance of diagnosis. The uptake of ES decreased to 31% as the chance of diagnosis using other genetic testing methods increased to 40%. These results demonstrate the importance of the chance of diagnosis in the value-based comparison of ES and other genetic testing methods.
Probability of diagnostic test uptake for exome sequencing compared with other genetic testing methods. Chance of a diagnosis is varied from 35% to 40% for the other genetic testing method alternatives to understand the relative value of exome sequencing (ES) over single-gene testing, depending on the likelihood of receiving a diagnosis. A low-cost (CAD$3000) and high-cost (CAD$6000) scenario were compared for ES, representing the low and high cost range of clinical testing. Operative procedures and series of other tests and procedures have an uptake probability significantly lower than 0.0001%.
DISCUSSION
This study estimated personal utility of ES and, more broadly, the value of a diagnosis for parents of children with rare diseases. We found that parents of children with rare diseases place a high value on ES compared with other diagnostic testing if all other attributes were the same. Parents would be willing to pay $6590 (US$4943), wait 5.2 years to obtain a diagnostic test result, and accept a reduction of 3.1% in the chance of receiving a diagnosis for ES testing compared with operative procedures.
Our WTP estimates are higher than in the study by Regier et al., which found that parents of children with developmental disabilities were willing to pay an average of CAD$1656 (US$1523) for genetic testing (array genomic hybridization) that was more advanced than conventional cytogenetic testing.36 Conversely, a DCE study exploring parent preferences for genome sequencing for children with rare diseases (focused on variants of unknown significance and secondary findings) found that cost had the lowest relative importance to parents.37
In our study, the positive impacts of receiving a diagnosis (changes to medical management, facilitates access to diseases specific services, and provides knowledge about the cause, progression, and future impact of the disease) were found, on average, to have a greater impact on respondents’ choices than the negative impacts (lifestyle restrictions, subject to discrimination and/or being labeled, receive a poor prognosis). Additionally, each level of the positive impact attribute was valued similarly. Consistent with previous research, our results suggest that parents place intrinsic value on information and knowledge, suggesting that changes in clinical outcomes alone may not be suitable to measure the full benefit of the diagnostic tests. This is important with respect to the valuation of benefits for next-generation sequencing; valuation should include nonhealth outcomes to capture benefits beyond health outcomes.24,25 The negative impact of a diagnosis attribute demonstrated that chance of discrimination or a poor prognosis significantly reduced respondents' WTP, WTW, and MAC, while lifestyle restrictions was not valued significantly. Previous research identified parental concern with genetic testing; specifically, receiving a diagnosis that leads to discrimination, their child being negatively labeled, and lifestyle restrictions.19,20,38,39 We found that parents value the time to obtain an answer from diagnostic testing, whether the diagnostic test(s) resulted in a diagnosis or not, and are willing to pay for a reduction in time to obtain an answer. This aligns with previous research that suggests parents value an earlier diagnosis because it can lead to timely care and treatment.2,36
Our findings are consistent with previous qualitative research exploring attitudes and experiences of parents who have children with rare diseases or suspected genetic conditions. Levenseller et al.40 found that most parents would want to receive all results from testing such as ES, including secondary findings, and indicated the benefits of knowing all information outweighed the risks of future discrimination. Sapp et al.17 found that while parents had positive attitudes toward information about variants that have treatment or preventive measures, they still valued any health information and expressed discomfort in not learning information. Further, Sapp et al.17 state that parents “were optimistic that receiving their child’s results could empower them to identify novel ways to control aspects of their child’s condition.” Similarly, Harris et al.18 found that most participants would want to receive all findings and indicated that learning any information was better than learning none, even secondary findings with unknown clinical implications or treatments. A DCE study exploring parent preferences for genome sequencing for children with rare diseases, focusing on variants of unknown significance and secondary findings, found that most parents would prefer to receive all findings from genome sequencing regardless of uncertain or secondary findings.37 However Peyron et al.37 also found preference heterogeneity in that there is a proportion of parents who do not want all findings and would prefer a “targeted diagnostic approach” specific to the condition they are seeking a diagnosis for. Lastly, parents value genetic testing and receiving a diagnosis for disease treatment, understanding disease etiology, and family planning.3,14,15,16,17,18,19,20
There are several strengths to our study. To our knowledge, this is the first utility-based survey to obtain quantitative benefit–risk data on the value of a diagnosis for parents of children with rare diseases. Given our comprehensive recruitment strategy, we were able to capture respondents from every province and territory across Canada, whose children have a wide range of rare diseases. As such, our results may be generalizable to the broader pediatric rare disease population in Canada.
Our results should be considered in the context of certain limitations. First, the DCE choice tasks were hypothetical and represent stated preferences rather than revealed preferences inferred from decisions actually made in practice. Additionally, the probability of diagnostic test uptake scenario comparisons were hypothetical and assumed the chance of a diagnosis was the same for all children, which may not be true in practice. All DCEs can be impacted by the use of simplifying heuristics by respondents. However, to what extent these affect external validity is unclear since similar choice behaviors are observed in practice. Second, we were unable to explore differences in preferences between individual diseases due to sample size. However, our model accounted for uncertainty and heterogeneity in preferences. Third, by recruiting through rare disease organizations, we may have captured parents who were more engaged in their child’s care than parents who are not part of these organizations. Our sample was also highly educated (>80% of respondents reporting postsecondary education) and may not be representative of the education level of all parents of children with rare diseases, so that our estimates of willingness to pay and the value of information may be overestimated. Lastly, by using social media we may have missed parents who do not have access to social media or a computer with Internet. However, we used a comprehensive recruitment approach at two sites and offered participants the option to complete the survey by paper. Previous research exploring the use of social media to conduct surveys in rare disease populations found that this method can be useful and faster for accessing large cohorts of rare disease patients or their families and caregivers, in addition to traditional recruitment methods.41
Timely access to ES has the potential to reduce the time to achieve a diagnosis and the costs associated with the diagnostic odyssey in patients with rare diseases. Before technologies like ES are incorporated routinely into care delivery for patients with rare diseases across Canada and more broadly, there must be a clear understanding of their value to patients and families.
Change history
18 July 2019
In the original version of this Article, financial support was not fully acknowledged. The PDF and HTML versions of the Article have now been corrected to include the following: SH acknowledges that the University of Aberdeen’s Health Economics Research Unit is funded by the Chief Scientist Office of the Scottish Government Health and Care Directorates.
18 July 2019
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Boycott KM, Vanstone MR, Bulman DE, MacKenzie AE. Rare-disease genetics in the era of next-generation sequencing: discovery to translation. Nat Rev Genet. 2013;14:681–691.
de Ru MH, Bouwman MG, Wijburg FA, van Zwieten MCB. Experiences of parents and patients with the timing of Mucopolysaccharidosis type I (MPS I) diagnoses and its relevance to the ethical debate on newborn screening. Mol Genet Metab. 2012;107:501–507.
Lewis C, Skirton H, Jones R. Living without a diagnosis: the parental experience. Genet Test Mol Biomarkers. 2010;14:807–815.
Kingsmore SF, Saunders CJ. Deep sequencing of patient genomes for disease diagnosis: when will it become routine? Sci Transl Med. 2011;3:87ps23–87ps23.
Monroe GR, Frederix GW, Savelberg SMC, et al. Effectiveness of whole-exome sequencing and costs of the traditional diagnostic trajectory in children with intellectual disability. Genet Med. 2016;18:949–956.
Stark Z, Schofield D, Alam K, et al. Prospective comparison of the cost-effectiveness of clinical whole-exome sequencing with that of usual care overwhelmingly supports early use and reimbursement. Genet Med. 2017;19:867–874.
Soden SE, Saunders CJ, Willig LK, et al. Effectiveness of exome and genome sequencing guided by acuity of illness for diagnosis of neurodevelopmental disorders. Sci Transl Med. 2014;6:265ra168–265ra168.
van Nimwegen KJM, Schieving JH, MAAP Willemsen, et al. The diagnostic pathway in complex paediatric neurology: a cost analysis. Eur J Paediatr Neurol. 2015;19:233–239.
Anderson M, Elliott EJ, Zurynski YA. Australian families living with rare disease: experiences of diagnosis, health services use and needs for psychosocial support. Orphanet J Rare Dis. 2013;8:22.
Bamshad MJ, Ng SB, Bigham AW, et al. Exome sequencing as a tool for Mendelian disease gene discovery. Nat Rev Genet. 2011;12:745–755.
Biesecker LG, Green RC. Diagnostic clinical genome and exome sequencing. N Engl J Med. 2014;370:2418–2425.
McKusick-Nathans Institute of Genetic Medicine Johns Hopkins University School of Medicine. Online Mendelian Inheritance in Man (OMIM) gene map statistics. 2019. https://www.omim.org/statistics/geneMap. Accessed 7 January 2019.
Esquivel-Sada D, Nguyen MT. Diagnosis of rare diseases under focus: impacts for Canadian patients. J Community Genet. 2018;9:37–50.
Chen L-S, Xu L, Huang T-Y, Dhar SU. Autism genetic testing: a qualitative study of awareness, attitudes, and experiences among parents of children with autism spectrum disorders. Genet Med. 2013;15:274–281.
Palmer CGS, Martinez A, Fox M, Sininger Y, Grody WW, Schimmenti LA. Ethnic differences in parental perceptions of genetic testing for deaf infants. J Genet Couns. 2008;17:129–138.
Withrow KA, Burton S, Arnos KS, Kalfoglou A, Pandya A. Consumer motivations for pursuing genetic testing and their preferences for the provision of genetic services for hearing loss. J Genet Couns. 2008;17:252–260.
Sapp JC, Dong D, Stark C, et al. Parental attitudes, values, and beliefs toward the return of results from exome sequencing in children. Clin Genet. 2014;85:120–126.
Harris ED, Ziniel SI, Amatruda JG, et al. The beliefs, motivations, and expectations of parents who have enrolled their children in a genetic biorepository. Genet Med. 2012;14:330–337.
Narcisa V, Discenza M, Vaccari E, Rosen-Sheidley B, Hardan AY, Couchon E. Parental interest in a genetic risk assessment test for autism spectrum disorders. Clin Pediatr (Phila). 2013;52:139–146.
Anderson JA, Meyn MS, Shuman C, et al. Parents perspectives on whole genome sequencing for their children: qualified enthusiasm? J Med Ethics. 2017;43:535–539.
Phillips KA, Johnson FR, Maddala T. Measuring what people value: a comparison of “attitude” and “preference” surveys. Health Serv Res. 2002;37:1659–1679.
Feero W, Wicklund C, Veenstra DL. The economics of genomic medicine: insights from the IOM roundtable on translating genomic-based research for health. JAMA. 2013;309:1235–1236.
Bridges JF, Hauber AB, Marshall D, et al. Conjoint analysis applications in health—a checklist: a report of the ISPOR Good Research Practices for Conjoint Analysis Task Force. Value Health. 2011;14:403–413.
Regier DA, Weymann D, Buchanan J, Marshall DA, Wordsworth S. Valuation of health and nonhealth outcomes from next-generation sequencing: approaches, challenges, and solutions. Value Health. 2018;21:1043–1047.
Phillips KA, Deverka PA, Marshall DA, et al. Methodological issues in assessing the economic value of next-generation sequencing tests: many challenges and not enough solutions. Value Health. 2018;21:1033–1042.
Johnson FR, Lancsar E, Marshall D, et al. Constructing experimental designs for discrete-choice experiments: report of the ISPOR Conjoint Analysis Experimental Design Good Research Practices Task Force. Value Health. 2013;16:3–13.
Hauber AB, González JM, Groothuis-Oudshoorn CGM, et al. Statistical methods for the analysis of discrete choice experiments: a report of the ISPOR Conjoint Analysis Good Research Practices Task Force. Value Health. 2016;19:300–315.
Coast J, Horrocks S. Developing attributes and levels for discrete choice experiments using qualitative methods. J Health Serv Res Policy. 2007;12:25–30.
Vass C, Rigby D, Payne K. The role of qualitative research methods in discrete choice experiments: a systematic review and survey of authors. Med Decis Making. 2017;37:298–313.
Collins D. Pretesting survey instruments: an overview of cognitive methods. Qual Life Res. 2003;12:229–238.
Lighthouse Studio 9.0.1 [computer program]. Orem, Sawtooth Software: Utah; 2017.
Hensher DA, Rose JM, Greene WH. Applied choice analysis: a primer. Cambridge, UK: Cambridge University Press; 2005.
Yang J-C, Johnson FR, Kilambi V, Mohamed AF. Sample size and utility-difference precision in discrete-choice experiments: a meta-simulation approach. J Choice Model. 2015;16:50–57.
Train K, Weeks M. Discrete choice models in preference space and willingness-to-pay space. In: Alberini A, Scarpa R, eds. Applications of simulation methods in environmental and resource economics. Dordrecht: Springer; 2005. p. 1–16.
Daly A, Hess S, Train K. Assuring finite moments for willingness to pay in random coefficient models. Transportation. 2012;39:19–31.
Regier DA, Friedman JM, Makela N, Ryan M, Marra CA. Valuing the benefit of diagnostic testing for genetic causes of idiopathic developmental disability: willingness to pay from families of affected children. Clin Genet. 2009;75:514–521.
Peyron C, Pélissier A, Béjean S. Preference heterogeneity with respect to whole genome sequencing. A discrete choice experiment among parents of children with rare genetic diseases. Soc Sci Med. 2018;214:125–132.
Lipstein EA, Nabi E, Perrin JM, Luff D, Browning MF, Kuhlthau KA. Parents’ decision-making in newborn screening: opinions, choices, and information needs. Pediatrics. 2010;126:696–704.
Geelen E, Van Hoyweghen I, Doevendans PA, Marcelis CLM, Horstman K. Constructing “best interests”: genetic testing of children in families with hypertrophic cardiomyopathy. Am J Med Genet A. 2011;155:1930–1938.
Levenseller BL, Soucier DJ, Miller VA, Harris D, Conway L, Bernhardt BA. Stakeholders’ opinions on the implementation of pediatric whole exome sequencing: implications for informed consent. J Genet Couns. 2014;23:552–565.
Davies W. Insights into rare diseases from social media surveys. Orphanet J Rare Dis. 2016;11:151.
Acknowledgements
This work was performed under the Care4Rare Canada Consortium funded by Genome Canada and the Ontario Genomics Institute (OGI-147), the Canadian Institutes of Health Research (CIHR), Ontario Research Fund, Genome Alberta, Genome British Columbia, Genome Quebec, and Children’s Hospital of Eastern Ontario Foundation. We thank Isabel Jordan and the Rare Disease Foundation, Durhane Wong-Rieger and the Canadian Organization for Rare Disorders, Gail Ouellette and Regroupement Québécois des Maladies Orphelines, Maureen Latocki and MitoCanada, Marla Spiegel and Muscular Dystrophy Canada, Mary Sunderland and the Foundation for Fighting Blindness, Chrisanne Campos and the Genetic Aortic Disorder Association of Canada, and Orphanet Canada for their assistance with participant recruitment. We also thank all of the respondents for their time and participation in this study, and Ken Deal for his guidance on the development of the discrete choice experiment. D.A.M. is supported by a Canada Research Chair in Health Systems and Services Research, and Arthur J.E. Child Chair in Rheumatology. T.H. is supported by a Frederick Banting & Charles Best Canada Graduate Scholarship–Doctoral Award from CIHR. K.M.B. is supported by a CIHR Foundation Grant (FDN-154279). SH acknowledges that the University of Aberdeen’s Health Economics Research Unit is funded by the Chief Scientist Office of the Scottish Government Health and Care Directorates.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
The authors declare no conflicts of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Marshall, D.A., MacDonald, K.V., Heidenreich, S. et al. The value of diagnostic testing for parents of children with rare genetic diseases. Genet Med 21, 2798–2806 (2019). https://doi.org/10.1038/s41436-019-0583-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-019-0583-1
Keywords
This article is cited by
-
The value of knowing: preferences for genetic testing to diagnose rare muscle diseases
Orphanet Journal of Rare Diseases (2024)
-
The value of genomic testing in severe childhood speech disorders
European Journal of Human Genetics (2024)
-
QALYs and rare diseases: exploring the responsiveness of SF-6D, EQ-5D-5L and AQoL-8D following genomic testing for childhood and adult-onset rare genetic conditions in Australia
Health and Quality of Life Outcomes (2023)
-
Key drivers of family-level utility of pediatric genomic sequencing: a qualitative analysis to support preference research
European Journal of Human Genetics (2023)
-
The Value of Genomic Testing: A Contingent Valuation Across Six Child- and Adult-Onset Genetic Conditions
PharmacoEconomics (2022)