Abstract
Germline variants that affect the proofreading activity of polymerases epsilon (POLE) and delta (POLD1) predispose to colorectal adenomas and carcinomas, among other cancers. All cancer-associated pathogenic variants reported to date consist of non-disruptive genetic changes affecting the sequence that codifies the exonuclease domain (ED). Generally, disruptive (frameshift, stop-gain) POLE and POLD1 variants and missense variants outside the ED do not predispose to cancer. However, this statement may not be true for some, very specific variants that would indirectly affect the proofreading activity of the corresponding polymerase. We evaluated, by using multiple approaches, the possibility that POLD1 c.883G>A; p.(Val295Met), -a variant located 9 amino acids upstream the ED and present in ~0.25% of hereditary cancer patients-, affects POLD1 proofreading activity. Our findings show cumulative evidence that support no alteration of the proofreading activity and lack of association with cancer. The variant is classified as likely benign according to the ACMG/AMP guidelines.
Similar content being viewed by others
Introduction
Germline pathogenic non-disruptive variants in the region coding for the exonuclease domain (ED) of polymerases epsilon and delta, cause increased risk to colorectal cancer (CRC), adenomatous polyposis and other tumor types, including endometrial, breast, ovarian and brain cancers; which defines the polymerase proofreading-associated polyposis (PPAP) [1, 2]. The alteration of POLE or POLD1 proofreading activity, either in the germline or in a tumor (somatic), causes defective DNA repair during replication, which translates into an accumulation of specific genetic changes in associated tumors (>100 variants per Mb (var/Mb) and COSMIC mutational signature SBS10, or SBS14 and SBS20 when combined with mismatch repair (MMR) deficiency) [1, 3]. Available data indicate that there is no association of loss-of-function variants and of variants located outside the ED with cancer, as they do not alter the polymerase proofreading [2, 4, 5]. Nevertheless, although never empirically proven, it has been speculated that some variants located outside the ED might indirectly affect proofreading, thus having a similar effect than ED pathogenic variants. Such is the case of POLE c.1420G>A; p.(Val474Ile), which affects a residue three amino acids downstream the ED, and causes an effect on proofreading when tested in a yeast model [6]. However, exome sequencing of a carrier’s CRC revealed neither hyper/ultra-mutation nor accumulation of the transversions observed in proofreading-defective tumors. Here we present the results of multiple studies performed to elucidate the potential pathogenicity of POLD1 c.883G>A; p.(Val295Met), a recurrently identified variant affecting a residue in close proximity to POLD1 ED.
Methods
POLD1 p.Val295Met carriers
Variant carriers were identified among: (i) 2,309 unrelated familial/early-onset cancer patients subjected to a multi-gene hereditary cancer panel [5]; (ii) 504 unrelated cancer patients that include high risk breast and/or ovarian families, patients with personal or familial history of different tumor types previously associated with PPAP (CRC and polyposis excluded), patients with other multiple tumors, and patients fulfilling the criteria for TP53 genetic testing [5]; and (iii) 529 families with familial/early onset CRC and/or polyposis and no germline pathogenic variants in other known high-penetrance CRC genes [7]. The characteristics of the cohorts are detailed elsewhere [5, 7]. Informed consent was obtained from all subjects and the study received the approval of IDIBELL Ethics Committee.
In silico pathogenicity prediction
In silico predictions were extracted from Varsome [8], which uses BayesDel_addAF, DEOGEN2, EIGEN, FATHMM-MKL, M-CAP, MVP, MutationAssessor, MutationTaster, PrimateAI, REVEL and SIFT for pathogenicity predictions and GERP++ for conservation.
Tumor mutational signatures
Mutational signature analysis from tumor exome sequencing data was performed with DeconstructSigs [5].
3D structure modeling and predictions
The cryo-EM structure of human POLD1 determined at 3.08 angstroms resolution (PDB ID: 6tny, chain A) and a 3D model based on the crystallographic structure of the homologous yeast protein Pol3 (PDB ID: 3iay, chain A) [7], were used in this study. 3D stability predictions were performed with I-Mutant 3.0 (http://gpcr.biocomp.unibo.it/cgi/predictors/I-Mutant3.0/I-Mutant3.0.cgi), CUPSAT (http://cupsat.tu-bs.de), PoPMuSiC (http://dezyme.com), MAESTRO (https://biwww.che.sbg.ac.at/maestro/web/), INPS-3D (http://inpsmd.biocomp.unibo.it/inpsSuite/default/index3D), DeepDDG (http://protein.org.cn/ddg.html) and DynaMut (http://biosig.unimelb.edu.au/dynamut/).
Case-control studies
POLD1 p.(Val295Met) allele frequencies in breast cancer patients, CRC patients and controls were obtained from a population-based multi case-control series (MCC-Spain, www.mccspain.org).
Variant repository
Variant and phenotype information of the families carrying POLD1 p.Val295Met has been submitted to LOVD (https://www.lovd.nl/3.0/home).
Results and discussion
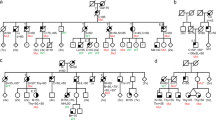
By searching for pathogenic variants in POLE and POLD1 affecting the proofreading activity of polymerases epsilon and delta, our group identified POLD1 (LRG_785, t1) c.883G>A; p.(Val295Met), a variant located 9 amino acids upstream the ED, in a total of 16 families (19 carriers). Of those, two families belonged to the series of 529 familial/early onset colorectal cancer (CRC) and/or polyposis families [7] (Families 1 & 2; Table 1); 11 to the 2,309 familial/early-onset cancer patients [5] (Families 3–13); and three to the 504 unrelated cancer patients with selected phenotypes [5] (Materials and Methods for details) (Families 14–16).
The tumor spectrum of POLD1 p.(Val295Met) carriers mainly included breast and/or ovarian cancer (11/19 carriers) and CRC (8/19). In two families, POLD1 p.(Val295Met) co-occurred with a BRCA2 pathogenic variant, and in one family, with the likely pathogenic variant POLD1 p.(Asp316Gly) in trans (Table 1).
POLD1 c.883G>A; p.(Val295Met) was not predicted pathogenic (benign computational verdict based on 10 benign predictions vs. 1 pathogenic prediction by FATHMM-MKL) and affected a non-conserved amino acid (GERP + + = 3.64). Nevertheless, further analyses were performed to elucidate its actual involvement in cancer predisposition, due to its recurrence (16 families) and its proximity to the ED.
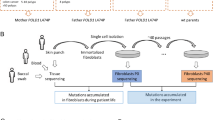
Exome sequencing data from an MMR-proficient CRC developed at age 48 by a POLD1 p.(Val295Met) carrier (Family 2) revealed no hypermutation (~5 var/Mb). Mutational signature analysis revealed a subtle presence of proofreading defective-associated signatures SBS10 (2% signature contribution), and SBS20 (3.5% contribution), associated with combined MMR deficiency and POLD1 pathogenic variant, despite the absence of detectable MMR deficiency (Supplementary Fig. 1). No somatic POLE or POLD1 ED variants were identified. These findings led us to study the mutational burden and signatures in other MMR-proficient tumors harboring the POLD1 variant. We identified one tumor, among 42 non-treated, stage II, MMR-proficient CRCs with exome sequencing data [9], with a somatic POLD1 p.(Val295Met) and no additional suspicious POLE or POLD1 variants. The tumor harbored ~50 var/Mb, and no trace of POLE/D1-associated signatures (Supplementary Fig. 1).
Being the valine 295 not conserved in yeast, we were not able to perform a yeast-based proofreading assay to assess the effect of the variant [5]. The suspicion of a potential (weak) effect of the variant on the proofreading activity of polymerase δ, -based on the results of the analysis of tumor mutational signatures in one of the tumors and the proximity of the variant to the ED-, led us to perform an in-depth analysis of the effect of the variant on the structural 3D conformation changes of the ED and/or the alteration of the DNA binding cleft, which has been observed to be the ED structural (3D) region most directly associated with the proofreading activity of the polymerases [5, 10, 11].
We used the cryo-EM structure of human POLD1 and a 3D model based on the crystallographic structure of the homologous yeast protein [7], to study the effect of p.Val295Met. While the cryo-EM structure and the 3D model superpose perfectly with a root mean square deviation of less than 1.9 angstroms (PDBeFold method) (Fig. 1A), the DNA binding site is placed at different positions (Fig. 1B and C). Single-stranded DNA from the 3D model is in the same position as in bacteriophage T4 polymerase complex (PDB ID: 1NOY), which fits with the position of the DNA when the exonuclease is working. Despite the proximity of Val295 to the ED in the linear sequence (9 amino acids upstream), both the cryo-EM structure and the 3D model show that residue Val295 is distant from the DNA binding pocket of the exonuclease, suggesting lack of effect on the proofreading activity.
Compared to the wildtype residue (Val295), Met295 was predicted, by 4 out of 7 different methods, to locally alter the 3D stability of the protein (Supplementary Table 1). As predicted by DynaMut, this local structure destabilization is caused by direct steric clashes with neighboring residues (Supplementary Fig. 2). However, when comparing the 3D conformation changes of the whole protein by Normal Mode Analysis (NMA; implemented in DynaMut) between POLD1 wildtype and POLD1 p.(Val295Met), the two variants showed similar structural profiles; i.e. low fluctuation for the whole protein except for the C-terminal region in the cryo-EM structure (Supplementary Fig. 3); indicating that Met295 has no effect on the global protein dynamics.
Based on the carriers’ phenotypes, we evaluated the allele frequency of POLD1 c.883G>A in CRC and breast cancer patients and matched controls from the same geographic area (source: MCC_Spain [12]). The case-control study showed lack of association with either cancer type (Table 2). Moreover, the observed allele frequencies (0.19% for CRC patients, 0.18% for breast cancer patients and 0.16%-0.20% for controls) were highly similar to those observed in the prospective hereditary cancer cohort (0.24%) and in Spanish individuals (0.22%; source: http://csvs.babelomics.org/) (Table 2). To investigate the potentially associated phenotypes, we separately analyzed the allele frequencies in hereditary/early-onset breast and/or ovarian cancer patients, and in hereditary/early-onset CRC and/or polyposis patients included in the prospective hereditary cancer cohort. While the p.(Val295Met) allele frequency in breast and/or ovarian phenotypes (0.17%) was similar to the one observed in the above-mentioned cases and controls, the frequency in patients with colorectal phenotypes was higher (0.50%), suggesting a potential association with the disease (Table 2). To validate this observation in an independent series of patients, we accessed the exome sequencing data obtained from 1,006 familial/early-onset CRC probands via Canvar (https://canvar.icr.ac.uk; accessed October 2020). The allele frequency for POLD1 p.(Val295Met) was 0.05% (1/1,778), same that those observed in European non-cancer population (MAF: 0.05%; 119/239,056; source: gnomAD 2.1.1), arguing against its involvement in colorectal cancer predisposition. Interestingly, the frequency of the variant was higher in Spanish population (0.22%), compared to that observed in Europeans (0.05%), suggesting an enrichment in that geographical area.
Our findings suggest that POLD1 p.(Val295Met) does not affect the proofreading activity of POLD1 and is not associated with cancer predisposition. Considering the gathered evidence, application of the ACMG/AMP guidelines [5, 13] resulted in the classification of POLD1 p.Val295Met as likely benign (BS1, BP4; Supplementary Table 2).
References
Rayner E, van Gool IC, Palles C, Kearsey SE, Bosse T, Tomlinson I, et al. A panoply of errors: polymerase proofreading domain mutations in cancer. Nat Rev Cancer. 2016;16:71–81.
Palles C, Cazier JB, Howarth KM, Domingo E, Jones AM, Broderick P, et al. Germline mutations affecting the proofreading domains of POLE and POLD1 predispose to colorectal adenomas and carcinomas. Nat Genet. 2013;45:136–44.
Alexandrov LB, Kim J, Haradhvala NJ, Huang MN, Tian AQ, Wu Y. et al. The repertoire of mutational signatures in human cancer. Nature. 2020;578:94–101.
Mur P, Palles C, Tomlinson I, Valle L. Reply to: “Development of an MSI-positive colon tumor with aberrant DNA methylation in a PPAP patient”. J Hum Genet. 2020;65:513–4.
Mur P, García-Mulero S, Del Valle J, Magraner-Pardo L, Vidal A, Pineda M, et al. Role of POLE and POLD1 in familial cancer. Genet Med. 2020;22:2089–100.
Esteban-Jurado C, Gimenez-Zaragoza D, Munoz J, Franch-Exposito S, Alvarez-Barona M, Ocana T, et al. POLE and POLD1 screening in 155 patients with multiple polyps and early-onset colorectal cancer. Oncotarget. 2017;8:26732–43.
Bellido F, Pineda M, Aiza G, Valdés-Mas R, Navarro M, Puente DA, et al. POLE and POLD1 mutations in 529 kindred with familial colorectal cancer and/or polyposis: review of reported cases and recommendations for genetic testing and surveillance. Genet Med. 2016;18:325–32.
Kopanos C, Tsiolkas V, Kouris A, Chapple CE, Albarca Aguilera M, Meyer R, et al. VarSome: the human genomic variant search engine. Bioinformatics. 2019;35:1978–80.
Sanz-Pamplona R, Lopez-Doriga A, Paré-Brunet L, Lázaro K, Bellido F, Alonso MH, et al. Exome sequencing reveals AMER1 as a frequently mutated gene in colorectal cancer. Clin Cancer Res. 2015;21:4709–18.
Barbari SR, Kane DP, Moore EA, Shcherbakova PV. Functional analysis of cancer-associated DNA polymerase ε variants in saccharomyces cerevisiae. G3 (Bethesda) 2018;8:1019–29.
Hamzaoui N, Alarcon F, Leulliot N, Guimbaud R, Buecher B, Colas C, et al. Genetic, structural, and functional characterization of POLE polymerase proofreading variants allows cancer risk prediction. Genet Med. 2020;2:1533–41.
Castaño-Vinyals G, Aragonés N, Pérez-Gómez B, Martín V, Llorca J, Moreno V, et al. Population-based multicase-control study in common tumors in Spain (MCC-Spain): rationale and study design. Gac Sanit. 2015;29:308–15.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24.
Funding
This work was funded by the Spanish Ministry of Science and Innovation, co-funded by FEDER funds [SAF2016-80888-R, PID2019-111254RB-I00, Severo Ochoa SVP-2014-068895 contract (LM-P)]; Instituto de Salud Carlos III, co-funded by FEDER funds [CIBERONC CB16/12/00234, CIBERESP, Sara Borrell contract (PM) and PI19/00553, CIBERONC ayuda iniciación a la investigación (EE)]; EMBO Short-Term Fellowship [8374 (LM-P)]; Government of Catalonia [AGAUR 2017SGR1282, CERCA Program]; and Fundación Olga Torres. This study was facilitated by COST Action CA17118, supported by COST (European Cooperation in Science and Technology).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Mur, P., Magraner-Pardo, L., García-Mulero, S. et al. Solving the enigma of POLD1 p.V295M as a potential cause of increased cancer risk. Eur J Hum Genet 30, 485–489 (2022). https://doi.org/10.1038/s41431-021-00926-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-021-00926-6
This article is cited by
-
No April fools in clinical genomics
European Journal of Human Genetics (2022)