Abstract
Background
PIT1-positive pituitary adenoma (PIT1-PA) is one of the most important lineages of pituitary adenoma (PA), which causes systematic endocrine disorders and a worse prognosis. Tumour-associated fibroblast (TAF) is a crucial stroma cell type in the tumour microenvironment (TME). However, cellular and functional heterogeneity of TAF and immune cells in PIT1-PA have not been fully investigated.
Methods
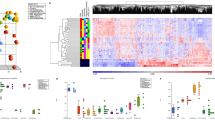
By single-cell RNA sequencing of four PIT1-PAs and further analyses, we characterised the molecular and functional profiles of 28 different cell subtypes.
Results
PA stem cells in PIT1/SF1-positve PA were in a hybrid epithelial/mesenchymal state, and differentiated along the PIT1- and SF- dependent branches. C1Q was overwhelmingly expressed in tumour-associated macrophages, indicating its pro-tumoral functionality. PIT1-PA progression was characterised by lower cell–cell communication strength and higher cell adhesion-associated signals, indicating the immunosuppressive but pro-invasive microenvironment. IFN-γ signal repressed functional remodelling of myofibroblastic TAF (mTAF) towards inflammatory TAF/antigen-presenting TAF. IFN-γ inhibited mTAF phenotypes and N-cadherin expression through STAT3 signal axis. CDH2 knockdown in TAFs abrogated their pro-tumour function in PAs.
Conclusions
Our study builds up a cellular landscape of PIT1-PA TME and highlights anti-tumour function of IFN-γ mediated TAF remodelling, which benefits clinical treatments and drug development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The single-cell RNA sequencing data generated in this paper are available in National Genomics Data Center by accession no. HRA003110 (https://ngdc.cncb.ac.cn/gsa-human/).
Code availability
R scripts used in this study to analyse data and generate figures can be found in Github (https://github.com/lvliang418/single-cell-RNAseq-for-PIT1-PA).
References
Fernandez A, Karavitaki N, Wass JA. Prevalence of pituitary adenomas: a community-based, cross-sectional study in Banbury (Oxfordshire, UK). Clin Endocrinol. 2010;72:377–82.
Asa SL, Mete O, Perry A, Osamura RY. Overview of the 2022 WHO Classification of Pituitary Tumors. Endocr Pathol. 2022;33:6–26.
Asa SL, Casar-Borota O, Chanson P, Delgrange E, Earls P, Ezzat S, et al. From pituitary adenoma to pituitary neuroendocrine tumor (PitNET): an International Pituitary Pathology Club proposal. Endocr Relat Cancer. 2017;24:C5–C8.
Lopes MBS. The 2017 World Health Organization classification of tumors of the pituitary gland: a summary. Acta Neuropathol. 2017;134:521–35.
Bollerslev J, Heck A, Olarescu NC. MANAGEMENT OF ENDOCRINE DISEASE: Individualized Management of Acromegaly. Eur J Endocrinol. 2019;181:R57-r71.
Tampourlou M, Trifanescu R, Paluzzi A, Ahmed SK, Karavitaki N. THERAPY OF ENDOCRINE DISEASE: Surgery in microprolactinomas: effectiveness and risks based on contemporary literature. Eur J Endocrinol. 2016;175:R89–96.
Trouillas J, Burman P, McCormack A, Petersenn S, Popovic V, Dekkers O, et al. Aggressive pituitary tumours and carcinomas: two sides of the same coin? Eur J Endocrinol. 2018;178:C7–C9.
Di Ieva A, Rotondo F, Syro LV, Cusimano MD, Kovacs K. Aggressive pituitary adenomas-diagnosis and emerging treatments. Nat Rev Endocrinol. 2014;10:423–35.
Hanahan D, Weinberg RA. The hallmarks of cancer. Cell. 2000;100:57–70.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
Orimo A, Gupta PB, Sgroi DC, Arenzana-Seisdedos F, Delaunay T, Naeem R, et al. Stromal fibroblasts present in invasive human breast carcinomas promote tumor growth and angiogenesis through elevated SDF-1/CXCL12 secretion. Cell. 2005;121:335–48.
Lv L, Zhang S, Hu Y, Zhou P, Gao L, Wang M, et al. Invasive pituitary adenoma-derived tumor-associated fibroblasts promote tumor progression both in vitro and in vivo. Exp Clin Endocrinol Diabetes. 2017;126:213–21.
Özdemir BC, Pentcheva-Hoang T, Carstens JL, Zheng X, Wu CC, Simpson TR, et al. Depletion of carcinoma-associated fibroblasts and fibrosis induces immunosuppression and accelerates pancreas cancer with reduced survival. Cancer Cell. 2014;25:719–34.
Chen Z, Zhou L, Liu L, Hou Y, Xiong M, Yang Y, et al. Single-cell RNA sequencing highlights the role of inflammatory cancer-associated fibroblasts in bladder urothelial carcinoma. Nat Commun. 2020;11:5077.
Biffi G, Oni TE, Spielman B, Hao Y, Elyada E, Park Y, et al. IL1-induced JAK/STAT signaling is antagonized by TGFβ to shape CAF heterogeneity in pancreatic ductal adenocarcinoma. Cancer Discov. 2019;9:282–301.
Elyada E, Bolisetty M, Laise P, Flynn WF, Courtois ET, Burkhart RA, et al. Cross-species single-cell analysis of pancreatic ductal adenocarcinoma reveals antigen-presenting cancer-associated fibroblasts. Cancer Discov. 2019;9:1102–23.
Guo W, Wang D, Wang S, Shan Y, Liu C, Gu J. scCancer: a package for automated processing of single-cell RNA-seq data in cancer. Brief Bioinforma. 2021;22:1–9.
Hänzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinforma. 2013;14:7.
Zhang S, Cui Y, Ma X, Yong J, Yan L, Yang M, et al. Single-cell transcriptomics identifies divergent developmental lineage trajectories during human pituitary development. Nat Commun. 2020;11:5275.
Murray PJ. Macrophage polarization. Annu Rev Physiol. 2017;79:541–66.
Qiu X, Mao Q, Tang Y, Wang L, Chawla R, Pliner HA, et al. Reversed graph embedding resolves complex single-cell trajectories. Nat Methods. 2017;14:979–82.
La Manno G, Soldatov R, Zeisel A, Braun E, Hochgerner H, Petukhov V, et al. RNA velocity of single cells. Nature. 2018;560:494–8.
Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinforma. 2008;9:559.
Aibar S, González-Blas CB, Moerman T, Huynh-Thu VA, Imrichova H, Hulselmans G, et al. SCENIC: single-cell regulatory network inference and clustering. Nat Methods. 2017;14:1083–6.
Jin S, Guerrero-Juarez CF, Zhang L, Chang I, Ramos R, Kuan C-H, et al. Inference and analysis of cell-cell communication using CellChat. Nat Commun. 2021;12:1088. https://doi.org/10.1038/s41467-021-21246-9.
Leek JT, Johnson WE, Parker HS, Jaffe AE, Storey JD. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics. 2012;28:882–3.
Marques P, Barry S, Carlsen E, Collier D, Ronaldson A, Awad S, et al. Pituitary tumour fibroblast-derived cytokines influence tumour aggressiveness. Endocr Relat Cancer. 2019;26:853–65.
Bromberg JF, Wrzeszczynska MH, Devgan G, Zhao Y, Pestell RG, Albanese C, et al. Stat3 as an oncogene. Cell. 1999;98:295–303.
Conway JR, Lex A, Gehlenborg N. UpSetR: an R package for the visualization of intersecting sets and their properties. Bioinformatics. 2017;33:2938–40.
Jongsma MLM, Neefjes J, Spaapen RM. Playing hide and seek: tumor cells in control of MHC class I antigen presentation. Mol Immunol. 2021;136:36–44.
Würth R, Thellung S, Corsaro A, Barbieri F, Florio T. Experimental evidence and clinical implications of pituitary adenoma stem cells. Front Endocrinol (Lausanne). 2020;11:54.
Lu JQ, Adam B, Jack AS, Lam A, Broad RW, Chik CL. Immune cell infiltrates in pituitary adenomas: more macrophages in larger adenomas and more T cells in growth hormone adenomas. Endocr Pathol. 2015;26:263–72.
Revel M, Sautes-Fridman C, Fridman WH, Roumenina LT. C1q+ macrophages: passengers or drivers of cancer progression. Trends Cancer. 2022;8:517–26. https://doi.org/10.1016/j.trecan.2022.02.006.
Jardine L, Barge D, Ames-Draycott A, Pagan S, Cookson S, Spickett G, et al. Rapid detection of dendritic cell and monocyte disorders using CD4 as a lineage marker of the human peripheral blood antigen-presenting cell compartment. Front Immunol. 2013;4:495.
Jhunjhunwala S, Hammer C, Delamarre L. Antigen presentation in cancer: insights into tumour immunogenicity and immune evasion. Nat Rev Cancer. 2021;21:298–312.
Imanishi T, Saito T. T cell co-stimulation and functional modulation by innate signals. Trends Immunol. 2020;41:200–12.
Mota JM, Leite CA, Souza LE, Melo PH, Nascimento DC, de-Deus-Wagatsuma VM, et al. Post-sepsis state induces tumor-associated macrophage accumulation through CXCR4/CXCL12 and favors tumor progression in mice. Cancer Immunol Res. 2016;4:312–22.
Du Y, Sui Y, Cao J, Jiang X, Wang Y, Yu J, et al. Dynamic changes in myofibroblasts affect the carcinogenesis and prognosis of bladder cancer associated with tumor microenvironment remodeling. Front Cell Dev Biol. 2022;10:833578.
Gocher AM, Workman CJ, Vignali DAA. Interferon-gamma: teammate or opponent in the tumour microenvironment? Nat Rev Immunol. 2021;22:158–172. https://doi.org/10.1038/s41577-021-00566-3.
Kreso A, Dick JE. Evolution of the cancer stem cell model. Cell Stem Cell. 2014;14:275–91.
Atashzar MR, Baharlou R, Karami J, Abdollahi H, Rezaei R, Pourramezan F, et al. Cancer stem cells: a review from origin to therapeutic implications. J Cell Physiol. 2020;235:790–803.
Mantovani G, Giardino E, Treppiedi D, Catalano R, Mangili F, Spada A, et al. Stem cells in pituitary tumors: experimental evidence supporting their existence and their role in tumor clinical behavior. Front Endocrinol (Lausanne). 2019;10:745.
Mertens F, Gremeaux L, Chen J, Fu Q, Willems C, Roose H, et al. Pituitary tumors contain a side population with tumor stem cell-associated characteristics. Endocr Relat Cancer. 2015;22:481–504.
Prager BC, Xie Q, Bao S, Rich JN. Cancer stem cells: the architects of the tumor ecosystem. Cell Stem Cell. 2019;24:41–53.
Puram SV, Tirosh I, Parikh AS, Patel AP, Yizhak K, Gillespie S, et al. Single-cell transcriptomic analysis of primary and metastatic tumor ecosystems in head and neck cancer. Cell. 2017;171:1611–24.e24.
Cui Y, Li C, Jiang Z, Zhang S, Li Q, Liu X, et al. Single-cell transcriptome and genome analyses of pituitary neuroendocrine tumors. Neuro Oncol. 2021;23:1859–71. https://doi.org/10.1093/neuonc/noab102.
Ruan X, Yi J, Hu L, Zhi J, Zeng Y, Hou X, et al. Reduced MHC class II expression in medullary thyroid cancer identifies patients with poor prognosis. Endocr Relat Cancer. 2022;29:87–98.
Chowell D, Morris LGT, Grigg CM, Weber JK, Samstein RM, Makarov V, et al. Patient HLA class I genotype influences cancer response to checkpoint blockade immunotherapy. Science. 2018;359:582–7.
Cho IJ, Lui PP, Obajdin J, Riccio F, Stroukov W, Willis TL, et al. Mechanisms, hallmarks, and implications of stem cell quiescence. Stem Cell Rep. 2019;12:1190–200.
Howley BV, Mohanty B, Dalton A, Grelet S, Karam J, Dincman T, et al. The ubiquitin E3 ligase ARIH1 regulates hnRNP E1 protein stability, EMT and breast cancer progression. Oncogene. 2022;41:1679–90.
Godfrey DI, Stankovic S, Baxter AG. Raising the NKT cell family. Nat Immunol. 2010;11:197–206.
Nelson A, Lukacs JD, Johnston B. The current landscape of NKT cell immunotherapy and the hills ahead. Cancers. 2021;13:5174. https://doi.org/10.3390/cancers13205174.
Godfrey DI, Uldrich AP, McCluskey J, Rossjohn J, Moody DB. The burgeoning family of unconventional T cells. Nat Immunol. 2015;16:1114–23.
Shao XQ, Chen ZY, Wang M, Yang YP, Yu YF, Liu WJ, et al. Effects of long-acting somatostatin analogues on lipid metabolism in patients with newly diagnosed acromegaly: a retrospective study of 120 cases. Horm Metab Res. 2022;54:25–32.
Wang M, Guo S, He M, Shao X, Feng L, Yu Y, et al. High-performance liquid chromatography-mass spectrometry-based lipid metabolite profiling of acromegaly. J Clin Endocrinol Metab. 2020;105:e1075–e1084. https://doi.org/10.1210/clinem/dgaa014.
Pellicci DG, Koay HF, Berzins SP. Thymic development of unconventional T cells: how NKT cells, MAIT cells and gammadelta T cells emerge. Nat Rev Immunol. 2020;20:756–70.
Väyrynen JP, Haruki K, Lau MC, Väyrynen SA, Ugai T, Akimoto N, et al. Spatial organization and prognostic significance of NK and NKT-like cells via multimarker analysis of the colorectal cancer microenvironment. Cancer Immunol Res. 2022;10:215–27.
Le Bouteiller P, Barakonyi A, Giustiniani J, Lenfant F, Marie-Cardine A, Aguerre-Girr M, et al. Engagement of CD160 receptor by HLA-C is a triggering mechanism used by circulating natural killer (NK) cells to mediate cytotoxicity. Proc Natl Acad Sci USA. 2002;99:16963–8.
Kim TJ, Park G, Kim J, Lim SA, Kim J, Im K, et al. CD160 serves as a negative regulator of NKT cells in acute hepatic injury. Nat Commun. 2019;10:3258.
Derynck R, Turley SJ, Akhurst RJ. TGFβ biology in cancer progression and immunotherapy. Nat Rev Clin Oncol. 2021;18:9–34.
Wang B, Zhang S, Tong F, Wang Y, Wei L. HPV(+) HNSCC-derived exosomal miR-9-5p inhibits TGF-β signaling-mediated fibroblast phenotypic transformation through NOX4. Cancer Sci. 2022;113:1475–87. https://doi.org/10.1111/cas.15281.
Nan P, Dong X, Bai X, Lu H, Liu F, Sun Y, et al. Tumor-stroma TGF-β1-THBS2 feedback circuit drives pancreatic ductal adenocarcinoma progression via integrin α(v)β(3)/CD36-mediated activation of the MAPK pathway. Cancer Lett. 2022;528:59–75.
Stuelten CH, Zhang YE. Transforming growth factor-β: an agent of change in the tumor microenvironment. Front Cell Dev Biol. 2021;9:764727.
Grauel AL, Nguyen B, Ruddy D, Laszewski T, Schwartz S, Chang J, et al. TGFbeta-blockade uncovers stromal plasticity in tumors by revealing the existence of a subset of interferon-licensed fibroblasts. Nat Commun. 2020;11:6315.
Saénz-de-Santa-María I, Celada L, Chiara MD. The leader position of mesenchymal cells expressing n-cadherin in the collective migration of epithelial cancer. Cells. 2020;9:731. https://doi.org/10.3390/cells9030731.
Labernadie A, Kato T, Brugués A, Serra-Picamal X, Derzsi S, Arwert E, et al. A mechanically active heterotypic E-cadherin/N-cadherin adhesion enables fibroblasts to drive cancer cell invasion. Nat Cell Biol. 2017;19:224–37.
Wang MD, Xiang H, Zhang L, Wang C. Integration of OV6 expression and CD68(+) tumor-associated macrophages with clinical features better predicts the prognosis of patients with hepatocellular carcinoma. Transl Oncol. 2022;25:101509.
Courtney AN, Tian G, Metelitsa LS. Natural killer T cells and other innate-like T lymphocytes as emerging platforms for allogeneic cancer cell therapy. Blood. 2022:blood.2022016201. https://doi.org/10.1182/blood.2022016201.
Acknowledgements
We appreciated the kindly help from Dr. Tang Jie (West China Hospital of Sichuan University) for technical support in scRNA-seq experiments, and Prof. Lu Kefeng (West China Hospital of Sichuan University) for language editing. We also appreciated the help from Figdraw Group for providing images in Fig. 1a.
Funding
We appreciated the financial supports from the National Natural Science Foundation of China (Grants No. 82072582), Sichuan Science and Technology Program (Grants No. 2023NSFSC1869 and 2022YFS0322), Post-Doctor Research Project, West China Hospital, Sichuan University (Grants No. 2020HXBH157) and 1.3.5 project for disciplines of excellence, West China Hospital, Sichuan University (Grants No. 2019HXFH018).
Author information
Authors and Affiliations
Contributions
LL, SY, HL, SJ and PZ conceived the project. LL and SY wrote the manuscript with help from all authors. HL, XL and WM performed scRNA-seq. LL, HL and LL performed IF, IHC and imaging. LL conducted the bioinformatics analyses. LL and YJ performed cellular and molecular investigations. AS constructed some of the figures. All authors edited and proofread the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
All experimental procedures involved human tumour samples were approved by the Institutional Review Board of West China Hospital of Sichuan University. Signed informed consents were obtain from patients before surgery. The animal experiments were performed according to the Guidelines set forth by Chinese National Institutes of Health and institutional guidelines and approved by the Biomedical Research Ethics Committee of West China Hospital of Sichuan University.
Consent for publication
Not applicable.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lyu, L., Jiang, Y., Ma, W. et al. Single-cell sequencing of PIT1-positive pituitary adenoma highlights the pro-tumour microenvironment mediated by IFN-γ-induced tumour-associated fibroblasts remodelling. Br J Cancer 128, 1117–1133 (2023). https://doi.org/10.1038/s41416-022-02126-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-022-02126-5
This article is cited by
-
Single-cell transcriptomic analysis reveals tumor cell heterogeneity and immune microenvironment features of pituitary neuroendocrine tumors
Genome Medicine (2024)
-
Single-cell transcriptomics reveal distinct immune-infiltrating phenotypes and macrophage–tumor interaction axes among different lineages of pituitary neuroendocrine tumors
Genome Medicine (2024)
-
Tumour microenvironment and pituitary tumour behaviour
Journal of Endocrinological Investigation (2023)
-
Tumor immune microenvironment in pituitary neuroendocrine tumors (PitNETs): increased M2 macrophage infiltration and PD-L1 expression in PIT1-lineage subset
Journal of Neuro-Oncology (2023)