Abstract
Salinization is recognized as a threat to soil fertility worldwide. A challenge in understanding the effects of salinity on soil microbial communities is the fact that it can be difficult to disentangle the effects of salinity from those of other variables that may co-vary with salinity. Here we use a trait-based approach to identify direct effects of salinity on soil bacterial communities across two salinity gradients. Through dose–response relationships between salinity and bacterial growth, we quantified distributions of the trait salt tolerance within the communities. Community salt tolerance was closely correlated with soil salinity, indicating a strong filtering effect of salinity on the bacterial communities. Accompanying the increases in salt tolerance were consistent shifts in bacterial community composition. We identified specific bacterial taxa that increased in relative abundances with community salt tolerance, which could be used as bioindicators for high community salt tolerance. A strong filtering effect was also observed for pH across the gradients, with pH tolerance of bacterial communities correlated to soil pH. We propose phenotypic trait distributions aggregated at the community level as a useful approach to study the role of environmental factors as filters of microbial community composition.
Similar content being viewed by others
Introduction
One of the major challenges in the field of microbial ecology is to move beyond descriptive reports of patterns in community composition and find a basis for predicting shifts in bacterial communities in response to environmental changes [1]. Unfortunately, our understanding of the underlying dynamics that structure bacterial communities remains limited. Differences in community composition have primarily been linked to edaphic factors based on correlative relationships [2]. In a global survey of bacterial communities from both terrestrial and aquatic environments, salinity emerged as the dominant factor linked to bacterial community composition [3] and has also been reported to be the factor most strongly correlated with community composition in aquatic systems [4, 5]. In soil, a large proportion of the variance in overall soil bacterial community composition has been shown to be associated with differences in soil pH [6, 7]. Despite the widespread occurrences of saline soils, much of our understanding of the role of salinity as a driver of bacterial communities has been derived from aquatic systems, with far fewer reports of salinity responses along soil salinity gradients.
Globally, it is estimated that around 900 million hectares of soil are affected to some degree by high ion concentrations [8]. Saline soils, commonly defined as soils with an electrical conductivity (EC) in a saturated soil extract of >4 dS m−1 [9], cover a large area in arid and semi-arid regions of the globe. ECs in saline soils can reach >200 dS m−1 [10], matching the EC of the most saline bodies of water. Considering that soil salinity can span several orders of magnitude (from <0.1 dS m−1 in non-saline soils to >100 dS m−1 in highly saline soil), it should be expected that salinity has a similarly strong effect on bacterial community composition in soil as has been observed in aquatic systems and that soil salinity could be another strong driver of microbial community composition in addition to soil pH. Indeed, studies on the composition of bacterial communities of saline soils found shifts in community composition associated with salinity, as well as a decline in phylogenetic diversity as salinity increased [11,12,13,14].
When identifying environmental drivers of bacterial community composition based on correlative relationships, the influence of confounding factors is difficult to tease apart from the direct effect of the environmental factor of interest. For instance, soil salinity is frequently correlated with reduced organic matter (OM) input [15, 16] and differences in soil pH [17]. To identify direct effects of environmental factors on the community, trait-based approaches have been proposed [18, 19]. However, information on phenotypic traits at the resolution of single bacterial taxa is often unavailable [20], especially in soil where most taxa remain undescribed [21]. Measuring traits aggregated at the community level can be used as an alternative to measure the distribution of traits within a community [18, 22]. This approach has been used to study microbial trait distributions including those of pH tolerance [23, 24], salt tolerance [25, 26], and heavy metal tolerance [27] among others. Tolerance to heavy metal pollution measured at the level of communities has been linked to shifts in community composition [28, 29], which makes it possible to infer a causal connection between shifts in the abundance of microbial taxa and changes in community tolerance. Shifts in environmental community tolerance indicate that the environmental factor in question effectively filtered the bacterial community. Changes in bacterial community composition that correlate with increases in community salt tolerance are thus far stronger indicators of a causal link between soil salinity and community composition than correlations between salinity and bacterial community compositions alone.
In this study, we tested the associations between community composition and the distribution of the trait salt tolerance along gradients of soil salinity. We used two salinity gradients, of which one was confounded by a pH gradient. We also established pH optima for the communities to investigate whether soil pH had selected for a shift in the trait distribution of pH tolerance and thus also investigated if pH had filtered bacterial community structure. We hypothesized (i) that the local environment would have selected for a community with matched trait distributions for salt tolerance and pH tolerance, (ii) that differences in salt tolerance and pH tolerance would be reflected in differences in the community composition, and (iii) that changes in bacterial community composition observed with increasing salt tolerance of the community would be consistent across both gradients.
Material and methods
Sites
Bulk soil samples were collected along two salinity gradients located along Lake O’Connor in Western Australia. Each gradient was sampled in three transects of eight sampling points. At each sampling point, 6 soil cores (5 cm diameter, 5 cm depth) within a 2 m radius were combined into a composite sample and stored in polyethylene Ziploc bags. This resulted in a total of 24 samples collected for each gradient (Fig. S1). Along the gradients, vegetation changed with distance from the lakeshore. On the northern shore gradient (henceforth the agricultural [AG] gradient; 32°28’ S, 119°13’ E), the first sampling point of each transect was located at the edge of the salt lake. The vegetation at sampling point 1 consisted of small halophilic shrubs with little ground cover. At sampling points 2–4, halophilic species were gradually replaced by grasses and small trees (Allocasuarina sp. and Eucalytpus sp.), while ground cover increased with distance from the lakeshore. Sampling points 5–8 were located in agricultural land used for wheat production. The distance covered by each transect of the AG gradient was ca. 200 m. The distance between transect A and B was ca. 130 m and between B and C ca. 560 m.
On the southern shore gradient (henceforth the natural vegetation [NV] gradient; 32°30’ S, 119°13’ E), sampling point 1 was also located at the lakeshore, with a vegetation consisting of mostly halophilic shrubs. At sampling point 2, vegetation consisted of grasses and Allocasuarina sp. From samplings points 3–6, Eucalyptus sp. became more common, with increasing ground cover. Sampling points 7–8 were covered with non-saline Eucalyptus woodland. Each transect of the NV gradient covered a distance of ca. 300 m from the lakeshore to the last sampling point. The distance between transects was ca. 300 m. The AG and NV gradients were located about 2.5 km apart.
Soil EC and pH were determined in a 1:5 soil: water mixture. OM contents were obtained as loss-on-ignition (600 °C, 12 h). The amount of water-soluble cations (Ca2+, K+, Mg2+, Na+) was analyzed in water extractions from soil with inductively coupled plasma–optical emission spectroscopy (Optima 8300, Perkin Elmer).
Determination of trait distributions of salt and pH tolerance
Prior to measuring community salt and pH tolerance, soils were incubated with 5 mg g−1 alfalfa–straw mixture for 3 weeks at room temperature to boost bacterial growth rates. After this pre-incubation period, assays to determine the trait distributions of salt and pH tolerance were performed. Soil suspensions were created by mixing 1 g of soil with 20 ml of water. After homogenization and centrifugation (1000 × g), aliquots (1.35 ml) of bacterial suspension were transferred to 2 ml microcentrifugation tubes and were then adjusted to either different electrical conductivities (suspension EC) or different pH values (suspension pH).
To change suspension EC, the soil suspensions were mixed with a range of solutions of different concentrations of NaCl dissolved in water to create a gradient of 8 different salt concentrations (including one control level without salt addition) in a final volume of 1.5 ml. Target concentrations of NaCl additions ranged from 5.5 to 0.007 mol l−1, with the achieved suspension EC depending on the initial EC of the soil suspension. To adjust suspension pH, suspensions were mixed with 0.15 ml of a citrate–phosphate pH buffer including 11 different levels ranging from pH 3.0 (final concentration 1.1 mM K2HPO4 and 0.5 mM citric acid) to 8.0 (final concentration 0.25 mM KH2PO4 and 6.4 mM K2HPO4) or distilled H2O, resulting in a range of 12 different pHs for each sample. The suspension pH was validated with a pH meter. The used buffer concentrations did not affect bacterial Leu incorporation rates within the short time frame studied [30]. Following the adjustment of salinity or pH in the soil suspensions, bacterial growth was measured as the incorporation of 3H-labeled leucine into bacterial protein [31, 32]. Briefly, 2 µl of radioactively labeled leucine, ([3H]Leu, 185 MBq ml−1, 2 TBq mmol−1, Perkin Elmer) were added together with non-labeled leucine to the samples, resulting in a total concentration of 280 nM leucine. After 1 h incubation at room temperature, growth was terminated by the addition of trichloroacetic acid (TCA). After a series of washing steps using TCA and ethanol, the amount of incorporated 3H-label was determined through liquid scintillation [32].
DNA extraction, amplification, and sequencing
DNA was extracted from all soil samples before the addition of plant material used to boost growth rates for tolerance measurements (see above) and from two thirds of samples after they had been incubated with plant material for 3 weeks to check if the incubation period had resulted in community shifts. Subsamples of each soil sample were freeze-dried and ground. DNA was extracted from portions of 250 mg of homogenized ground soil using the MoBio PowerSoil DNA Isolation Kit (Carlsbad, CA, USA) according to the manufacturer’s recommendations. Extracted DNA was amplified using the 16S rRNA gene primer pair 515-F (5′-GTGCCAGCMGCCGCGGTAA-3′) and 806-R (5′-GGACTACHVGGGTWTCTAAT-3′) targeting the V4 region of the 16 S rRNA gene, which included Illumina adapters and unique barcode sequences for each sample. PCR was performed with GoTaq® Hot Start PCR Master Mix (Promega, Madison, WI, USA) in a 25 μl reaction. Thermal cycling consisted of an initial denaturation step at 94 °C for 3 min, followed by 35 cycles of denaturation at 94 °C (45 s), annealing at 50 °C (30 s), extension at 70 °C (90 s), and a final extension at 72 °C for 10 min. The amplified DNA was sequenced using a Illumina MiSeq platform (Illumina, San Diego, CA, USA).
Sequences were processed using the UPARSE pipeline [33] as described in Ramirez et al. [34]. Sequences were quality filtered and clustered de novo into operational taxonomic units (OTUs) at a 97% similarity level. Taxonomic information was assigned to OTUs using the 16S rRNA Greengenes database [35]. To correct for differences in sequencing depth, samples were rarefied to 10,000 reads. Samples with <10,000 reads and OTUs that were observed <10 times across all samples were excluded from downstream analyses. These criteria resulted in the removal of 4 out of the 48 samples and 8928 of the 12,326 OTUs.
Data analysis
Growth rates in the salt and pH tolerance assays were normalized to growth rates measured at the optimum EC or pH for each sample. In samples in which bacterial growth was inhibited only by increasing salinity, dose–response relationships were established using a logistic model, Y = c/[1 + eb(x−a)], where Y is the leucine incorporation rate, x is the logarithm of the suspension EC, a is the logIC50, c is the bacterial growth rate in the control without added salt, and b is a slope parameter indicating the rate of inhibition. In samples in which growth was inhibited by both increasing and decreasing salinity from its growth optimum, a double-logistic model [23] was used: Y = copt/(1 + exp[blow-EC(x – a(low-EC))]) + copt/(1 + exp[bhigh-EC(x – ahigh-EC)]) − copt, with copt the growth rate at optimal suspension EC, b the slope indicating the rate of decrease toward higher or lower suspension EC, and a the logIC50 toward higher and lower suspension EC. For comparison of salt tolerance between curves, the logIC50 toward higher suspension EC was used. To estimate the pH tolerance of the communities, the same double-logistic model was used, with suspension pH replacing suspension EC as the predictor variable. The suspension pH at which bacterial growth reached its optimum (pHopt) in each sample was used as an indicator of the community-level trait distribution of pH tolerance. Kaleidagraph 4.5.0 for Mac (Synergy software) was used to fit the logistic and double-logistic models. Linear regression models were used to test for significant correlations between community-level trait indicators (logIC50 and pHopt) and soil EC and pH.
The diversity of each sample was determined by calculating the Shannon diversity index. Multivariate statistics were performed in the R environment version 3.3.1 (R [36]) using the ‘vegan’ package [37]. The differences in overall community composition between samples were calculated using the Bray–Curtis dissimilarity index after Hellinger transformation [38]. To highlight the relationship between changes in community tolerance and shifts in community composition, a constrained ordination was performed by distance-based redundancy analysis using capscale (vegan) with logIC50, pHopt, and gradient as constraining variables. In addition, patterns in bacterial community compositions were also visualized by applying an unconstrained ordination method (principal coordinate analysis). The significance of constraining variables was tested with a permutation test (number of permutations = 10,000) using the anova function of the vegan package. Correlations between community composition and distributions of salt and pH tolerance were tested using Mantel tests between Bray–Curtis distance matrices of community composition and Euclidean distance matrices of trait distributions. To identify important OTUs correlated with trait distributions, we selected OTUs with a relative abundance of ≥1% in at least one sample. For these OTUs, we calculated Spearman’s rank correlations with both logIC50 and pHopt. OTUs with a Spearman’s rank correlation coefficient (ρ) of ≥0.5 or ≤−0.5 were selected as being positively or negatively correlated with a certain trait. To test for significant correlations between diversity and environmental variables, multiple linear regressions (α = 0.05) were performed for each gradient followed by analysis of variance using type II Sums of Squares, with Shannon diversity as the dependent variable and the logarithm of EC and pH as independent variables.
Results
Characterization of gradients
The AG gradient encompassed soil ECs ranging from 0.1 to 3 dS m−1 measured in a 1:5 soil:water mixture, while the soil pH along the gradient was between ca. 5.5 and ca. 7.0 for all but one sample, which had a pH > 8 (Fig. S2). The NV gradient encompassed soil ECs ranging from 0.2 to 9 dS m−1. Soil pH along the gradient ranged from ca. 4.5 to ca. 8.5. Along the NV gradient, there was a significant negative linear correlation between soil EC and soil pH (R2 = 0.50, p < 0.001), whereas along the AG gradient soil EC and soil pH were not significantly correlated (Fig. S2). OM content along the AG gradient ranged from 2.8 to 4.8% dry weight (dw) and was positively correlated with soil pH (R2 = 0.26, p < 0.01; Fig. S3) but not with soil EC (not shown). Along the NV gradient, OM content ranged from 7.6 to 10.0% dw and was neither correlated with soil EC nor soil pH (Fig. S3). Na+ was the predominant cation in sites from both gradients (Fig. S4).
Community trait distributions
In samples from sites with lower salinity, bacterial growth was inhibited with increasing salinity (Fig. 1a). The sigmoidal relationship between growth and the logarithm of the suspension EC could be modeled with a logistic function (R2 from 0.95 to 0.99). In communities from high-salinity samples, bacterial growth was inhibited by both low and high suspension EC (Fig. 1a). For these samples, a double-logistic function could fit the relationships between suspension EC and bacterial growth well (R2 from 0.91 to 0.99). In the most saline samples, the suspension EC at which communities had their growth optimum was about a factor of 10 higher than that measured in a 1:5 soil:water suspension.
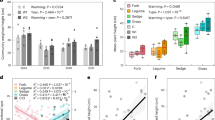
Response curves between bacterial growth and (a) suspension electrical conductivity (EC) and (b) suspension pH in the community tolerance assays. Depicted are representative examples of response curves of soils of different soil EC and soil pH. Values of soil EC (a) and soil pH (b) measured in a 1:5 soil:water mixture of the selected soils are given in the figure legends. Bacterial growth was normalized to the maximum growth rate in each sample. The relationships between bacterial growth and EC and pH were described with logistic or double logistic functions. To describe the community-level salt tolerance in a sample, the IC50 value (suspension EC at which bacterial growth was inhibited by 50% compared to the maximum) was used. To describe the community-level pH tolerance, the suspension pH at which growth reached its maximum was used (pHopt). (c) Relationship between soil EC and community-level salt tolerance (indicated by IC50) along both salinity gradients (AG and NV gradient). (d) Relationship between soil pH and community-level pH tolerance (indicated by pHopt). Samples with a pH < 5.5 (open symbols) were excluded from the regression analysis
We used the logIC50 as an indicator of the community-level distribution of the trait of salt tolerance. There was a strong positive relationship between the logarithm of the soil EC and the logIC50 (R2 = 0.83, p < 0.001; Fig. 1c). The difference between soil EC and IC50 decreased with increasing salinity. In the most saline samples, the IC50 was about 10 times higher than the soil EC, whereas in the least saline samples IC50 was 70 times higher than soil EC.
The relationships between bacterial growth and suspension pH could also be modeled with a double-logistic function (R2 from 0.46 to 0.99, with a mean R2 = 0.91; Fig. 1b). The indicator used for the distribution of the trait pH tolerance was the pHopt for bacterial growth. Above soil pH 5.5, there was a linear relationship between soil pH and pHopt (R2 = 0.63, p < 0.001; Fig. 1d). For every 1 unit increase in soil pH, the optimum pH increased by ca. 0.5 units. Consequently, around soil pH 5.5, pHopt was around 0.8 units higher than the soil pH, whereas around soil pH 8.5 pHopt was around 0.6 units lower than the soil pH.
Community composition
In total, 3035 OTUs occurred with a frequency of at least 10 reads in the dataset. In the samples from the AG gradient, we found 2020 different OTUs and in the NV gradient 1897 different OTUs. Of these OTUs, 882 were found in samples from both gradients, whereas the rest were unique to either gradient. In all, 97% of reads belonged to Bacteria, and 3% were assigned to Archaea. The Archaea found along the gradients primarily belonged to two groups, namely, the Thaumarcheota (74% of archaeal reads) and the Halobacteria (20% of archaeal reads). Thaumarcheota were found in samples of all salinities, but were less common in highly saline sites (47% of archaeal reads in samples of salinities >4 dS m−1), whereas Halobacteria increased in relative abundance with high salinity (35% of archaeal reads in samples of salinities >4 dS m−1). The phylum Proteobacteria made up 41% of reads, followed by Bacteroidetes (15%), Actinobacteria (14%), Gemmatimonadetes (6%), and Planctomycetes (5%). The most abundant family was the Sphingomonadaceae, a family of Alphaproteobacteria, which accounted for 7% of all reads. Other families that made up >3% of reads were the Chitinophagaceae (Bacteroidetes) and the Xanthomonadaceae (Gammaproteobacteria).
Community composition changed with increasing salt tolerance along both gradients (Fig. 2, Fig. S5A) and converged between gradients as community salt tolerance increased. Similar patterns were observed when both constrained (Fig. 2) and unconstrained ordination methods (Fig. S5A) were used to visualize patterns in bacterial community composition along the gradients. Both salt and pH tolerance explained significant amounts of community composition, with a higher proportion of variation being related to salt tolerance (Fig. 2). Minor changes in bacterial community composition occurred during the 3-week incubation with plant material used to boost growth rates for tolerance measurements, but the effect of the incubation with plant material was not significant (Fig. S5B). Community composition along both gradients was significantly correlated with salt tolerance (Mantel test; ρ = 0.35, p < 0.001) and pH tolerance (Mantel test; ρ = 0.56, p < 0.001), with shifts in community composition being better correlated with changes in the trait distribution of pH tolerance than salt tolerance (Fig. 3a, b). Along the AG gradient, community composition was more strongly correlated with salt tolerance (Mantel test; ρ = 0.54, p < 0.001) than pH tolerance (Mantel test; ρ = 0.34, p < 0.01) (Fig. 3c, d), whereas along the NV gradient, the correlation between community composition and salt tolerance (Mantel test; ρ = 0.29, p < 0.01) was weaker than between community composition and pH tolerance (Mantel test; ρ = 0.84, p < 0.001) (Fig. 3e, f).
Canonical analysis of principal coordinates (CAPSCALE) derived from Bray–Curtis dissimilarities of the community composition of sampling points based on 16S rRNA gene amplicon sequencing. Panel (a) shows samples from the AG gradient, (b) shows samples from the NV gradient, and (c) shows samples from both gradients. Numbers in parentheses in the axis labels give the percentage of variance accounted for by the principal coordinates. Community salt tolerance (IC50), community pH tolerance (pHopt), and a dummy variable for gradient were used as constraining variables. The significance of constraining variables was tested with PERMANOVA and is indicated by asterisks next to the variable names.
Relationships between pairwise differences in community composition (Bray–Curtis dissimilarities) and community trait distributions (Euclidean distances). The fitted lines show linear regression curves between pairwise community distances and tolerance trait distances. Statistics (ρ) were calculated through Mantel tests (Spearman rank correlation). Panels (a, b) show the correlation between community dissimilarities and salt (a) and pH (b) tolerance distances for samples from both gradients. Panels (c, d) include samples from the AG gradient only, while (e, f) those from the NV gradient only
Overall, an increase in pH tolerance by 1 pH unit was connected to a shift in community composition of a similar magnitude as the change in community composition associated with an increase of logIC50 by 0.5 log (dS m−1), i.e., a 3.5-fold increase in salt tolerance (Fig. 3a, b). Increases in community salt tolerance were accompanied by larger shifts in community composition along the AG gradient than along the NV gradient. An increase in salt tolerance by 1 log (dS m−1) (i.e., a 10-fold increase) along the AG gradient (Fig. S7 C) was accompanied by a community shift of the same magnitude as an increase in salt tolerance by ca. 1.7 log (dS m−1) (i.e. a 50-fold increase) along the NV gradient (Fig. 3e).
In the AG gradient, 46 OTUs with a relative abundance ≥1% were found to be positively correlated with salt tolerance, while 33 OTUs were negatively correlated (Table S1). Fifteen of the 46 positively correlated OTUs belonged to the Gammaproteobacteria and 17 to the phylum Bacteroidetes, making them the taxonomic groups most strongly associated with increased community-level salt tolerance. Among these Gammaproteobacteria were OTUs classified as belonging to the genera Salinisphaera and Alkanibacter and several OTUs belonging to the families Xanthomonadaceae and Alteromonadaceae (including OTUs classified as Marinobacter). The Bacteroidetes associated with high salt tolerance included several Flavobacteriaceae (including OTUs classified as Gillisia, Gramella, and Salinimicrobium), Rhodothermaceae (including Rubricoccus), and Flammeovirgaceae, as well as OTUs classified as Balneola. In the NV gradient, 19 OTUs were positively correlated with salt tolerance. The class Gammaproteobacteria made up 10 of these 19 OTUs, which were classified as belonging to the families Salinisphaeraceae, Xanthomonadaceae, and Alteromonadacaea. Ten OTUs positively correlated with salinity in the NV gradient were also found to be positively correlated with salt tolerance in the AG gradient (Table S1). Of these 10 OTUs, 9 were only observed in communities with an IC50 >1.5 log (dS m−1).
In the NV gradient, 42 OTUs were positively correlated with pH optimum, while 24 were negatively correlated (Table S2). There was some overlap along the NV gradient between OTUs that responded to salt tolerance and pH optimum: 23 OTUs were positively correlated with pH optimum and negatively correlated with salt tolerance, while 17 OTUs were negatively correlated with pH optimum and positively with salt tolerance (Table S1, S2). OTUs associated with communities with high salt tolerance and low pH optima included mainly members of the phylum Gammaproteobacteria, whereas OTUs associated with communities with low salt tolerance and a high pH optima were more phylogenetically diverse.
Shannon diversity declined along the gradients with increasing salinity and decreasing pH (Fig. S6). The multiple linear regression models for Shannon diversity were significant for both the AG gradient (F(2,20) = 14.9, R2 = 0.56, p < 0.001) and the NV gradient (F(2,18) = 49.9, R2 = 0.83, p < 0.001). In the AG gradient, both logEC (p < 0.001) and pH (p < 0.05) significantly predicted Shannon diversity. In the NV gradient, only pH was found to significantly predict Shannon diversity (p < 0.001), whereas logEC did not predict a significant proportion of the variation in diversity beyond variation also attributed to pH.
Discussion
Trait–environment relationships
The two salinity gradients used in this study cover much of the range of salinities observed across soils globally [39], from non-saline to hypersaline soils, including sites where salt concentrations in the pore water approached saturated conditions. We hypothesized that a community inhabiting a more saline site would have been selected for higher salt tolerance, thereby increasing the amount of salinity required to inhibit growth of that community. We observed substantial and systematic variation in community tolerance to salt along the environmental gradients. In accordance with our hypothesis, community-level salt tolerance increased proportionally to the increase in soil salinity (Fig. 1b). As such, inconsistent links between soil salinity and bacterial salt tolerance that had previously been reported [25, 26] are likely at least partly explained by the small sample sizes and limited ranges in soil salinity considered.
Along the NV gradient, the span in pH ranged from ca. 4.5 to 8.5, encompassing most of the range of pH values that are commonly found in soils around the globe. Soil pH is the factor that was found to be the most strongly correlated with microbial community composition in continental-scale comparisons of soil communities [6, 40]. Similar to the observed increase in community salt tolerance with salinity, we expected community tolerance to pH to shift along the pH gradient. Consistent with our expectation, we found a significant correlation between soil pH and pHopt. The observed shifts in community salt tolerance as well as pH tolerance along the gradients (Fig. 1b, d) indicate that both environmental factors had shaped trait distributions and filtered the microbial communities [29, 41]. As such, this approach enabled us to infer a direct impact of the studied environmental factors on the bacterial community, which is not confounded with other factors changing along the gradients, since those would not have resulted in corresponding shifts in tolerance trait distributions [42,43,44].
Connecting phenotypic trait distributions and community compositions
One of the main mechanisms through which the trait distribution of tolerance in a community could change includes shifts in community composition, with better adapted species replacing less adapted ones [45, 46]. Changes in bacterial community composition along the gradients that correlate with the salt tolerance of the community are more likely to be a result of a direct influence of salinity, whereas changes in the community with salinity that are not accompanied with shifts in salt tolerance are more likely to be the result of indirect factors covarying with salinity. Compositional changes that accompany shifts in the distribution of traits that are selected upon by the environmental factor in question, in this case salinity, thus provide a far better basis to establish causal links between salinity and community composition than simply testing for correlations between soil salinity and community composition. However, an important caveat to consider is that not all of the observed changes in microbial community composition that were correlated with an increase in salt tolerance would necessarily be due to salinity alone.
As salinity increased along the studied gradients and became a more important constraint on the bacterial community, communities became more similar between samples from different gradients (Fig. 2c). It is likely that only a limited set of bacterial taxa could withstand the considerable stresses imposed on microbial cells in the highly saline soils, which was reflected also in a declining diversity of communities with increasing salinity (Fig. S6). Bacterial communities from different origins that are exposed to the same selection pressure frequently respond in a consistent manner, as has been reported, e.g., in response to heavy metal exposure [47, 48], addition of low molecular weight organic substrate [49], and periodic drying–rewetting events [50], as well as in response to macronutrient addition [51, 52]. As such, this suggests the existence of generalizable and predictable responses to environmental shifts in bacterial communities.
There are conflicting reports on which soil bacterial or archaeal taxa would be expected to increase in response to soil salinity. Previous studies identified a range of different taxa that were positively correlated with salinity [11, 13, 53, 54], which makes it difficult to derive meaningful a priori predictions of bacterial community responses to increasing salinity. In aquatic habitats, high salinity is often associated with a switch toward an archaeal-dominated community [55,56,57]. In some extremely saline soils, which included more extreme salinities than in this study, high relative abundances of archaea of up to 80% of the community were reported [10, 14]. However, in our study system we saw no change in the abundance of archaeal sequences along the salinity gradients and archaeal sequences never made up ca. >2% of the total number of reads. Canfora et al. [11] found no increase in Proteobacteria but an increasing abundance of Bacteroidetes, while in a study in wetland soils primarily Gamma- and Deltaproteobacteria were associated with higher salinity [54]. Studies in arid and saline soils in Antarctica found the phylum most strongly correlated with salinity to be Firmicutes [13, 53]. We found high community salt tolerance to be positively correlated primarily with OTUs belonging to two high-level phylogenetic groups, namely, the Bacteroidetes and the Gammaproteobacteria, indicating that the trait salt tolerance is represented in these taxonomic groups, with only a few other phyla among the OTUs associated with high salt tolerance. The Gammaproteobacteria that were found to be positively correlated with salinity included phylogenetic groups known to contain halophilic or marine bacteria, e.g., the families Alteromonadaceae [58, 59] and Salinisphaeraceae [60]. Also among the Bacteroidetes that were related to high salt tolerance were taxa belonging to the family Rhodothermaceae, which includes many halophilic bacteria [61], and the Flavobacteriaceae genera Gillisia, Gramella, and Salinimicrobium, which are frequently found in saline environments [62,63,64]. Taxa from the groups found in high salinity sites of both gradients could be targeted for the use as ‘bioindicator’ taxa for salt-affected communities to assess whether salinity had posed an important selective constraint on the communities. Investigations of these ‘bioindicator’ taxa could be useful in environments where salinity is variable due to irrigation or flooding and current soil salinities do not necessarily reflect the past impact of salinity [65]. In a similar vein, bacterial bioindicator taxa for heavy metal pollution have been put forward by looking for bacterial taxa correlated with high heavy metal pollution [66,67,68].
Comparing shifts in community salt and pH tolerance
In our study system, we were able to use the trait–environment relationships established between soil salinity and salt tolerance, as well as between soil pH and pH tolerance to compare the impact of salinity in a terrestrial system to that of pH (factors that are often difficult to disentangle). Salt tolerance accounted for a higher proportion of the total variation in community composition in the total dataset (Fig. 2c). In contrast, along the NV gradient, which was confounded with pH, a larger proportion of the variation in community composition was connected to differences in pH tolerance (Fig. 2b). Overall, bacterial communities whose salt tolerance differed by a factor of 3.5 were as dissimilar in their composition as communities whose pH tolerance differed by 1 pH unit (Fig. 3a, b).
Along the salinity gradients, a larger shift in the community composition per unit change in salt tolerance was observed in the AG gradient (Fig. 3c), which covered a narrower range of salinities than the NV gradient (Fig. 3d). It is possible that, as salinity in the soil solution approached more extreme values in the highly saline sites of the NV gradient, further increases in salinity from already saline conditions resulted in smaller changes in community composition. With salinity becoming the main factor constraining community composition, the pool of species that would have been capable of surviving these extreme conditions and able to outcompete and replace less adapted ones could have decreased [69]. This would imply that increases in salinity in non- or low-saline habitats would result in larger shifts in the bacterial community composition than increases in salinity in already highly saline soils. Similarly, along gradients of heavy metal pollution progressively smaller changes in community composition as levels of heavy metal pollution increased have been reported [67, 70]. In contrast, changes in pH tolerance were connected to shifts in community composition of similar magnitude along both gradients, despite the AG gradient covering a much smaller range of pH values. However, pH values between ca. 5 and 8 are common in soils [6, 40]. Therefore the span of soil pH values covered by the gradients in this study arguably represented less extreme conditions for bacteria than the upper limit of the range of salinities found along the gradients.
Conclusions
We propose a framework of using bacterial tolerance trait distributions along environmental gradients to identify environmental factors that constitute important filters on bacterial communities and thus infer a causal link . Accompanying these shifts in bacterial tolerance trait distributions along environmental gradients of pH and salinity, we could document large bacterial community differences. As bacterial communities were filtered by salinity, the community compositions grew similar between the two gradients. We identified a high degree of overlap between the two gradients in terms of which taxa were positively correlated with increased salt tolerance, indicative of consistent changes in community composition as bacterial communities became more salt tolerant. These taxa could be used as bioindicators to infer the distribution of salt tolerance in communities derived from environmental samples currently analyzed as part of large-scale survey efforts (including e.g. the Earth Microbiome Project [39]). Identifying such bioindicator taxa makes it possible to use community composition data to predict phenotypic traits (in this case, salt tolerance) of highly diverse soil microbial communities .
Disclaimer
The submitted material is original research, has not been published previously, and has not been submitted for publication elsewhere while under consideration.
References
Antwis RE, Griffiths SM, Harrison XA, Aranega-Bou P, Arce A, Bettridge AS, et al. Fifty important research questions in microbial ecology. FEMS Microbiol Ecol. 2017;93:fix044.
Lauber CL, Strickland MS, Bradford MA, Fierer N. The influence of soil properties on the structure of bacterial and fungal communities across land-use types. Soil Biol Biochem. 2008;40:2407–15.
Lozupone CA, Knight R. Global patterns in bacterial diversity. Proc Natl Acad Sci USA. 2007;104:11436–40.
Herlemann DPR, Labrenz M, Jurgens K, Bertilsson S, Waniek JJ, Andersson AF. Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea. ISME J. 2011;5:1571–9.
Logares R, Lindstrom ES, Langenheder S, Logue JB, Paterson H, Laybourn-Parry J, et al. Biogeography of bacterial communities exposed to progressive long-term environmental change. ISME J. 2013;7:937–48.
Lauber CL, Hamady M, Knight R, Fierer N. Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl Environ Microbiol. 2009;75:5111–20.
Rousk J, Bååth E, Brookes PC, Lauber CL, Lozupone C, Caporaso JG, et al. Soil bacterial and fungal communities across a pH gradient in an arable soil. ISME J. 2010;4:1340–51.
Szabolcs I. Salt-affected soils. Boca Raton: CRC Press LLC; 1989.
Richards LA. Diagnosis and improvement of saline and alkali soils. Washington: United States Department Of Agriculture; 1969.
Pandit AS, Joshi MN, Bhargava P, Shaikh I, Ayachit GN, Raj SR, et al. A snapshot of microbial communities from the Kutch: one of the largest salt deserts in the World. Extremophiles. 2015;19:973–87.
Canfora L, Bacci G, Pinzari F, Lo Papa G, Dazzi C, Benedetti A. Salinity and bacterial diversity: to what extent does the concentration of salt affect the bacterial community in a saline soil? PLos ONE. 2014;9:e106662.
Hollister EB, Engledow AS, Hammett AJM, Provin TL, Wilkinson HH, Gentry TJ. Shifts in microbial community structure along an ecological gradient of hypersaline soils and sediments. ISME J. 2010;4:829–38.
Van Horn DJ, Okie JG, Buelow HN, Gooseff MN, Barrett JE, Takacs-Vesbach CD. Soil microbial responses to increased moisture and organic resources along a salinity gradient in a polar desert. Appl Environ Microbiol. 2014;80:3034–43.
Xie KH, Deng Y, Zhang SC, Zhang WH, Liu JR, Xie YL, et al. Prokaryotic community distribution along an ecological gradient of salinity in surface and subsurface saline soils. Sci Rep. 2017;7:13332.
Setia R, Gottschalk P, Smith P, Marschner P, Baldock J, Setia D, et al. Soil salinity decreases global soil organic carbon stocks. Sci Total Environ. 2013;465:267–72.
Wong VNL, Dalal RC, Greene RSB. Salinity and sodicity effects on respiration and microbial biomass of soil. Biol Fert Soils. 2008;44:943–53.
Rengasamy P. Soil processes affecting crop production in salt-affected soils. Funct Plant Biol. 2010;37:613–20.
Wallenstein MD, Hall EK. A trait-based framework for predicting when and where microbial adaptation to climate change will affect ecosystem functioning. Biogeochemistry. 2012;109:35–47.
Webb CT, Hoeting JA, Ames GM, Pyne MI, Poff NL. A structured and dynamic framework to advance traits-based theory and prediction in ecology. Ecol Lett. 2010;13:267–83.
Green JL, Bohannan BJM, Whitaker RJ. Microbial biogeography: from taxonomy to traits. Science. 2008;320:1039–43.
Delgado-Baquerizo M, Oliverio AM, Brewer TE, Benavent-Gonzalez A, Eldridge DJ, Bardgett RD, et al. A global atlas of the dominant bacteria found in soil. Science. 2018;359:320–5.
Fierer N, Barberan A, Laughlin DC. Seeing the forest for the genes: using metagenomics to infer the aggregated traits of microbial communities. Front Microbiol. 2014;5:614.
Bååth E, Kritzberg E. pH tolerance in freshwater bacterioplankton: trait variation of the community as measured by leucine incorporation. Appl Environ Microbiol. 2015;81:7411–9.
Bárcenas-Moreno G, Bååth E, Rousk J. Functional implications of the pH-trait distribution of the microbial community in a re-inoculation experiment across a pH gradient. Soil Biol Biochem. 2016;93:69–78.
Kamble PN, Gaikwad VB, Kuchekar SR, Bååth E. Microbial growth, biomass, community structure and nutrient limitation in high pH and salinity soils from Pravaranagar (India). Eur J Soil Biol. 2014;65:87–95.
Rousk J, Elyaagubi FK, Jones DL, Godbold DL. Bacterial salt tolerance is unrelated to soil salinity across an arid agroecosystem salinity gradient. Soil Biol Biochem. 2011;43:1881–7.
Fernández-Calviño D, Arias-Estévez M, Díaz-Raviña M, Bååth E. Assessing the effects of Cu and pH on microorganisms in highly acidic vineyard soils. Eur J Soil Sci. 2012;63:571–8.
Wakelin S, Gerard E, Black A, Hamonts K, Condron L, Yuan T, et al. Mechanisms of pollution induced community tolerance in a soil microbial community exposed to Cu. Environ Pollut. 2014;190:1–9.
Berg J, Brandt KK, Al-Soud WA, Holm PE, Hansen LH, Sørensen SJ, et al. Selection for Cu-tolerant bacterial communities with altered composition, but unaltered richness, via long-term Cu exposure. Appl Environ Microbiol. 2012;78:7438–46.
Fernández-Calviño D, Rousk J, Brookes PC, Bååth E. Bacterial pH-optima for growth track soil pH, but are higher than expected at low pH. Soil Biol Biochem. 2011;43:1569–75.
Bååth E. Thymidine and leucine incorporation in soil bacteria with different cell size. Microb Ecol. 1994;27:267–78.
Bååth E, Pettersson M, Söderberg KH. Adaptation of a rapid and economical microcentrifugation method to measure thymidine and leucine incorporation by soil bacteria. Soil Biol Biochem. 2001;33:1571–4.
Edgar RC. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods. 2013;10:996–8.
Ramirez KS, Leff JW, Barberan A, Bates ST, Betley J, Crowther TW, et al. Biogeographic patterns in below-ground diversity in New York City’s Central Park are similar to those observed globally. P Roy Soc B Biol Sci. 2014;281:1988.
McDonald D, Price MN, Goodrich J, Nawrocki EP, DeSantis TZ, Probst A, et al. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 2012;6:610–8.
R Core Team. R: a language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing; 2016.
Oksanen J, Blanchet FG, Friendly M, Kindt R, Legendre P, McGlinn D, et al. vegan: Community ecology package. R package version 2.4-1. 2016. https://CRAN.R-project.org/package=vegan
Legendre P, Gallagher ED. Ecologically meaningful transformations for ordination of species data. Oecologia. 2001;129:271–80.
Thompson LR, Sanders JG, McDonald D, Amir A, Ladau J, Locey KJ, et al. A communal catalogue reveals Earth’s multiscale microbial diversity. Nature. 2017;551:457–63.
Griffiths RI, Thomson BC, James P, Bell T, Bailey M, Whiteley AS. The bacterial biogeography of British soils. Environ Microbiol. 2011;13:1642–54.
Schmitt H, Van Beelen P, Tolls J, Van Leeuwen CL. Pollution-induced community tolerance of soil microbial communities caused by the antibiotic sulfachloropyridazine. Environ Sci Technol. 2004;38:1148–53.
Blanck H. A critical review of procedures and approaches used for assessing pollution-induced community tolerance (PICT) in biotic communities. Hum Ecol Risk Assess. 2002;8:1003–34.
Ogilvie LA, Grant A. Linking pollution induced community tolerance (PICT) and microbial community structure in chronically metal polluted estuarine sediments. Mar Environ Res. 2008;65:187–98.
Tlili A, Hollender J, Kienle C, Behra R. Micropollutant-induced tolerance of in situ periphyton: establishing causality in wastewater-impacted streams. Water Res. 2017;111:185–94.
Lau JA, Lennon JT. Rapid responses of soil microorganisms improve plant fitness in novel environments. Proc Natl Acad Sci USA. 2012;109:14058–62.
Shade A, Peter H, Allison SD, Baho DL, Berga M, Burgmann H, et al. Fundamentals of microbial community resistance and resilience. Front Microbiol. 2012;3:417.
Laplante K, Derome N. Parallel changes in the taxonomical structure of bacterial communities exposed to a similar environmental disturbance. Ecol Evol. 2011;1:643–59.
Macdonald CA, Clark IM, Zhao FJ, Hirsch PR, Singh BK, McGrath SP. Long-term impacts of zinc and copper enriched sewage sludge additions on bacterial, archaeal and fungal communities in arable and grassland soils. Soil Biol Biochem. 2011;43:932–41.
Eilers KG, Lauber CL, Knight R, Fierer N. Shifts in bacterial community structure associated with inputs of low molecular weight carbon compounds to soil. Soil Biol Biochem. 2010;42:896–903.
Barnard RL, Osborne CA, Firestone MK. Responses of soil bacterial and fungal communities to extreme desiccation and rewetting. ISME J. 2013;7:2229.
Leff JW, Jones SE, Prober SM, Barberan A, Borer ET, Firn JL, et al. Consistent responses of soil microbial communities to elevated nutrient inputs in grasslands across the globe. Proc Natl Acad Sci USA. 2015;112:10967–72.
Ramirez KS, Craine JM, Fierer N. Consistent effects of nitrogen amendments on soil microbial communities and processes across biomes. Glob Change Biol. 2012;18:1918–27.
Geyer KM, Altrichter AE, Takacs-Vesbach CD, Van Horn DJ, Gooseff MN, Barrett JE. Bacterial community composition of divergent soil habitats in a polar desert. FEMS Microbiol Ecol. 2014;89:490–4.
Morrissey EM, Franklin RB. Evolutionary history influences the salinity preference of bacterial taxa in wetland soils. Front Microbiol. 2015;6:1013.
Vavourakis CD, Ghai R, Rodriguez-Valera F, Sorokin DY, Tringe SG, Hugenholtz P, et al. Metagenomic insights into the uncultured diversity and physiology of microbes in four hypersaline soda lake brines. Front Microbiol. 2016;7:211.
Ventosa A, Fernandez AB, Leon MJ, Sanchez-Porro C, Rodriguez-Valera F. The Santa Pola saltern as a model for studying the microbiota of hypersaline environments. Extremophiles. 2014;18:811–24.
Zhong ZP, Liu Y, Miao LL, Wang F, Chu LM, Wang JL, et al. Prokaryotic community structure driven by salinity and ionic concentrations in plateau lakes of the Tibetan Plateau. Appl Environ Microbiol. 2016;82:1846–58.
Ivanova EP, Mikhailov VV. A new family, Alteromonadaceae fam. nov., including marine proteobacteria of the genera Alteromonas, Pseudoalteromonas, Idiomarina, and Colwellia. Microbiology. 2001;70:10–17.
López-Pérez M, Rodriguez-Valera F. The family Alteromonadaceae. In: Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F, editors. The Prokaryotes: Gammaproteobacteria. Berlin, Heidelberg: Springer Berlin Heidelberg; 2014. p. 69–92.
Antunes A, Alam I, Bajic VB, Stingl U. Genome sequence of Salinisphaera shabanensis, a gammaproteobacterium from the harsh, variable environment of the brine-seawater interface of the Shaban Deep in the Red Sea. J Bacteriol. 2011;193:4555–6.
Park S, Akira Y, Kogure K. The family Rhodothermaceae. In: Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F, editors. The Prokaryotes: Other Major Lineages of Bacteria and The Archaea. Berlin, Heidelberg: Springer Berlin Heidelberg; 2014. p. 849–56.
Chen YG, Cui XL, Zhang YQ, Li WJ, Wang YX, Kim CJ, et al. Salinimicrobium terrae sp nov., isolated from saline soil, and emended description of the genus Salinimicrobium. Int J Syst Evol Microbiol. 2008;58:2501–4.
Joung Y, Kim H, Jang T, Ahn TS, Joh K. Gramella jeungdoensis sp nov., isolated from a solar saltern in Korea. J Microbiol. 2011;49:1022–6.
Nedashkovskaya OI, Kim SB, Lee KH, Mikhailov VV, Bae KS. Gillisia mitskevichiae sp nov., a novel bacterium of the family Flavobacteriaceae, isolated from sea water. Int J Syst Evol Microbiol. 2005;55:321–3.
Walsh DA, Papke RT, Doolittle WF. Archaeal diversity along a soil salinity gradient prone to disturbance. Environ Microbiol. 2005;7:1655–66.
Bier RL, Voss KA, Bernhardt ES. Bacterial community responses to a gradient of alkaline mountaintop mine drainage in Central Appalachian streams. ISME J. 2015;9:1378–90.
Pei YX, Yu ZS, Ji J, Khan A, Li XK. Microbial community structure and function indicate the severity of chromium contamination of the Yellow River. Front Microbiol. 2018;9:38.
Schneider AR, Gommeaux M, Duclercq J, Fanin N, Conreux A, Alahmad A, et al. Response of bacterial communities to Pb smelter pollution in contrasting soils. Sci Total Environ. 2017;605:436–44.
Li H, Colica G, Wu PP, Li DH, Rossi F, De Philippis R, et al. Shifting species interaction in soil microbial community and its influence on ecosystem functions modulating. Microb Ecol. 2013;65:700–8.
Epelde L, Lanzen A, Blanco F, Urich T, Garbisu C. Adaptation of soil microbial community structure and function to chronic metal contamination at an abandoned Pb-Zn mine. FEMS Microbiol Ecol. 2015;91:1–11.
Acknowledgements
The authors thank Andrew Wherrett for help with soil sampling and Dr. Ellen Kandeler for comments on an earlier version of the manuscript. We acknowledge funding from the Swedish research council (grant no. 2015-04942), the Swedish research council Formas (grant no. 942-2015-270), and the Knut and Alice Wallenberg Foundation (grant no. KAW 2017.0171).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rath, K.M., Fierer, N., Murphy, D.V. et al. Linking bacterial community composition to soil salinity along environmental gradients. ISME J 13, 836–846 (2019). https://doi.org/10.1038/s41396-018-0313-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-018-0313-8
This article is cited by
-
Nutrient-induced acidification modulates soil biodiversity-function relationships
Nature Communications (2024)
-
Effects of soil type and salinity levels on the performance and bacteriome of the halophyte Atriplex nummularia (old man saltbush)
Plant and Soil (2024)
-
Organic fertilizer and hydrothermal conditions change the distribution of Medicago sativa L. productivity and soil bacterial diversity in coastal saline soil
Chemical and Biological Technologies in Agriculture (2023)
-
Metagenomics reveals the habitat specificity of biosynthetic potential of secondary metabolites in global food fermentations
Microbiome (2023)
-
Microbial life in 25-m-deep boreholes in ancient permafrost illuminated by metagenomics
Environmental Microbiome (2023)