Abstract
Background
Bronchopulmonary dysplasia (BPD), a common morbidity among very preterm infants, is associated with chronic disease and neurodevelopmental impairments. A hypothesized mechanism for these outcomes lies in altered glucocorticoid (GC) activity. We hypothesized that BPD and its treatments may result in epigenetic differences in the hypothalamic-pituitary-adrenal (HPA) axis, which is modulated by GC, and could be ascertained using an established GC risk score and DNA methylation (DNAm) of HPA axis genes.
Methods
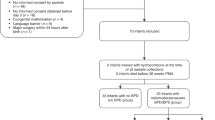
DNAm was quantified from buccal tissue (ECHO-NOVI) and from neonatal blood spots (ELGAN ECHO) via the EPIC microarray. Prenatal maternal characteristics, pregnancy complication, and neonatal medical complication data were collected from medical record review and maternal interviews.
Results
The GC score was not associated with steroid exposure or BPD. However, six HPA genes involved in stress response regulation demonstrated differential methylation with antenatal steroid exposure; two CpGs within FKBP5 and POMC were differentially methylated with BPD severity. These findings were sex-specific in both cohorts; males had greater magnitude of differential methylation within these genes.
Conclusions
These findings suggest that BPD severity and antenatal steroids are associated with DNAm at some HPA genes in very preterm infants and the effects appear to be sex-, tissue-, and age-specific.
Impact
-
This study addresses bronchopulmonary dysplasia (BPD), an important health outcome among preterm neonates, and interrogates a commonly studied pathway, the hypothalamic-pituitary-adrenal (HPA) axis.
-
The combination of BPD, the HPA axis, and epigenetic markers has not been previously reported.
-
In this study, we found that BPD itself was not associated with epigenetic responses in the HPA axis in infants born very preterm; however, antenatal treatment with steroids was associated with epigenetic responses.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 14 print issues and online access
$259.00 per year
only $18.50 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
Select de-identified data from the ECHO Program are available through NICHD’s Data and Specimen Hub (DASH). Information on study data not available on DASH, such as some Indigenous datasets, can be found on the ECHO study DASH webpage. The raw and processed DNAm data for NOVI are publicly accessible through NCBI Gene Expression Omnibus (GEO) via accession series GSE128821.
References
Martin, J. A., Hamilton, B. E. & Osterman, M. J. Births in the United States, 2013. NCHS Data Brief 175, 1–8 (2014).
Friedrich, M. J. Premature birth complications top cause of death in children younger than 5 years. JAMA 313, 235–235 (2015).
Aarnoudse-Moens, C. S., Weisglas-Kuperus, N., van Goudoever, J. B. & Oosterlaan, J. Meta-analysis of neurobehavioral outcomes in very preterm and/or very low birth weight children. Pediatrics 124, 717–728 (2009).
Hack, M. et al. Poor predictive validity of the Bayley Scales of Infant Development for cognitive function of extremely low birth weight children at school age. Pediatrics 116, 333–341 (2005).
Vohr, B. R., Wright, L. L., Poole, W. K. & McDonald, S. A. Neurodevelopmental outcomes of extremely low birth weight infants <32 weeks’ gestation between 1993 and 1998. Pediatrics 116, 635–643 (2005).
Aylward, G. P. Neurodevelopmental outcomes of infants born prematurely. J. Dev. Behav. Pediatr. 26, 427–440 (2005).
Grunau, R. E., Whitfield, M. F. & Davis, C. Pattern of learning disabilities in children with extremely low birth weight and broadly average intelligence. Arch. Pediatr. Adolesc. Med. 156, 615–620 (2002).
Hack, M. et al. Behavioral outcomes and evidence of psychopathology among very low birth weight infants at age 20 years. Pediatrics 114, 932–940 (2004).
Hack, M. et al. Behavioral outcomes of extremely low birth weight children at age 8 years. J. Dev. Behav. Pediatr. 30, 122–130 (2009).
Hille, E. T. et al. Social lifestyle, risk-taking behavior, and psychopathology in young adults born very preterm or with a very low birthweight. J. Pediatr. 152, 793–800, 800.e1–4 (2008).
Taylor, H. G., Klein, N. & Hack, M. School-age consequences of birth weight less than 750 g: a review and update. Dev. Neuropsychol. 17, 289–321 (2000).
Stephens, B. E. & Vohr, B. R. Neurodevelopmental outcome of the premature infant. Pediatr. Clin. North Am. 56, 631–646 (2009).
Allen, M. C. Neurodevelopmental outcomes of preterm infants. Curr. Opin. Neurol. 21, 123–128 (2008).
Schmidt, B. et al. Impact of bronchopulmonary dysplasia, brain injury, and severe retinopathy on the outcome of extremely low-birth-weight infants at 18 months: results from the trial of indomethacin prophylaxis in preterms. JAMA 289, 1124–1129 (2003).
Thébaud, B. et al. Bronchopulmonary dysplasia. Nat. Rev. Dis. Prim. 5, 1–23 (2019).
O’Reilly, M., Sozo, F. & Harding, R. Impact of preterm birth and bronchopulmonary dysplasia on the developing lung: long-term consequences for respiratory health. Clin. Exp. Pharm. Physiol. 40, 765–773 (2013).
Mowitz, M. E. et al. Health care burden of bronchopulmonary dysplasia among extremely preterm infants. Front. Pediatr. 7, 510 (2019).
Piyasena, C. et al. Dynamic changes in DNA methylation occur during the first year of life in preterm infants. Front. Endocrinol. 7, 158 (2016).
Schuster, J. et al. Effect of prematurity on genome wide methylation in the placenta. BMC Med. Genet. 20, 116 (2019).
Wang, X.-M. et al. Comparison of DNA methylation profiles associated with spontaneous preterm birth in placenta and cord blood. BMC Med. Genomics 12, 1 (2019).
Breton, C. V. et al. Small-magnitude effect sizes in epigenetic end points are important in children’s environmental health studies: the Children’s Environmental Health and Disease Prevention Research Center’s Epigenetics Working Group. Environ. Health Perspect. 125, 511–526 (2017).
Everson, T. M. et al. Epigenome-wide analysis identifies genes and pathways linked to neurobehavioral variation in preterm infants. Sci. Rep. 9, 6322 (2019).
Everson, T. M. et al. Serious neonatal morbidities are associated with differences in DNA methylation among very preterm infants. Clin. Epigenetics 12, 151 (2020).
Paniagua, U. et al. Epigenetic age acceleration, neonatal morbidities, and neurobehavioral profiles in infants born very preterm. Epigenetics 18, 2280738 (2023).
Giarraputo, J. et al. Medical morbidities and DNA methylation of NR3C1 in preterm infants. Pediatr. Res. 81, 68–74 (2017).
Knight, A. K. et al. Relationship between epigenetic maturity and respiratory morbidity in preterm infants. J. Pediatr. 198, 168–173.e2 (2018).
Cho, H. Y. et al. Prospective epigenome and transcriptome analyses of cord and peripheral blood from preterm infants at risk of bronchopulmonary dysplasia. Sci. Rep. 13, 12262 (2023).
Jackson, W. M. et al. Differential placental CpG methylation is associated with chronic lung disease of prematurity. Pediatr. Res. 91, 1428–1435 (2022).
Wang, X. et al. Epigenome-wide association study of bronchopulmonary dysplasia in preterm infants: results from the discovery-BPD program. Clin. Epigenetics 14, 57 (2022).
Cao-Lei, L., Laplante, D. P. & King, S. Prenatal maternal stress and epigenetics: review of the human research. Curr. Mol. Biol. Rep. 2, 16–25 (2016).
Parets, S. E. et al. Fetal DNA methylation associates with early spontaneous preterm birth and gestational age. PLoS ONE 8, e67489 (2013).
Provençal, N. et al. Glucocorticoid exposure during hippocampal neurogenesis primes future stress response by inducing changes in DNA methylation. Proc. Natl Acad. Sci. 117, 23280 (2020).
Wiechmann, T. et al. Identification of dynamic glucocorticoid-induced methylation changes at the FKBP5 locus. Clin. Epigenetics 11, 83 (2019).
Yang, R. et al. Longitudinal genome-wide methylation study of PTSD treatment using prolonged exposure and hydrocortisone. Transl. Psychiatry 11, 398 (2021).
Braun, P. R. et al. Genome‐wide DNA methylation investigation of glucocorticoid exposure within buccal samples. Psychiatry Clin. Neurosci. 73, 323–330 (2019).
Doyle, L. W. Postnatal corticosteroids to prevent or treat bronchopulmonary dysplasia. Neonatology 118, 244–251 (2021).
Godoy, L. D., Rossignoli, M. T., Delfino-Pereira, P., Garcia-Cairasco, N. & de Lima Umeoka, E. H. A comprehensive overview on stress neurobiology: basic concepts and clinical implications. Front. Behav. Neurosci. 12, 127 (2018).
Weaver, I. C. G. et al. Epigenetic programming by maternal behavior. Nat. Neurosci. 7, 847–854 (2004).
Weaver, I. C. G., Diorio, J., Seckl, J. R., Szyf, M. & Meaney, M. J. Early environmental regulation of hippocampal glucocorticoid receptor gene expression: characterization of intracellular mediators and potential genomic target sites. Ann. N. Y. Acad. Sci. 1024, 182–212 (2004).
Blackwell, C. K., Wakschlag, L. S., Gershon, R. C. & Cella, D., with the ECHO PRO Core. Measurement framework for the Environmental influences on Child Health Outcomes research program. Curr. Opin. Pediatr. 30, 276–284 (2018).
Gillman, M. W. & Blaisdell, C. J. Environmental influences on Child Health Outcomes, a research program of the National Institutes of Health. Curr. Opin. Pediatr. 30, 260–262 (2018).
Jacobson, L. P., Lau, B., Catellier, D. & Parker, C. B. An Environmental influences on Child Health Outcomes viewpoint of data analysis centers for collaborative study designs. Curr. Opin. Pediatr. 30, 269–275 (2018).
Hofheimer, J. A. et al. Psychosocial and medical adversity associated with neonatal neurobehavior in infants born before 30 weeks gestation. Pediatr. Res. 87, 721–729 (2020).
O’Shea, T. M. et al. The ELGAN study of the brain and related disorders in extremely low gestational age newborns. Early Hum. Dev. 85, 719–725 (2009).
Teschendorff, A. E. et al. A beta-mixture quantile normalization method for correcting probe design bias in Illumina Infinium 450 k DNA methylation data. Bioinformatics 29, 189–196 (2013).
Pidsley, R. et al. Critical evaluation of the Illumina MethylationEPIC BeadChip microarray for whole-genome DNA methylation profiling. Genome Biol. 17, 208 (2016).
Logue, M. W. et al. The correlation of methylation levels measured using Illumina 450K and EPIC BeadChips in blood samples. Epigenomics 9, 1363–1371 (2017).
Aryee, M. J. et al. Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 30, 1363–1369 (2014).
Fortin, J. P., Fertig, E. & Hansen, K. shinyMethyl: interactive quality control of Illumina 450k DNA methylation arrays in R. F1000Res 3, 175 (2014).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Martin, M. et al. Bronchopulmonary dysplasia and neurobehavioural outcomes at birth and 2 years in infants born before 30 weeks. Arch. Dis. Child. Fetal Neonatal Ed. fetalneonatal-2021-323405 https://doi.org/10.1136/archdischild-2021-323405 (2022).
Jensen, E. A. et al. The diagnosis of bronchopulmonary dysplasia in very preterm infants. An evidence-based approach. Am. J. Respir. Crit. Care Med. 200, 751–759 (2019).
Natarajan, G. et al. Effect of inborn vs. outborn delivery on neurodevelopmental outcomes in infants with hypoxic-ischemic encephalopathy: secondary analyses of the NICHD whole-body cooling trial. Pediatr. Res. 72, 414–419 (2012).
Zou, Z. et al. Associations of DNA methylation of HPA axis-related genes and neuroendocrine abnormalities in panic disorder. Psychoneuroendocrinology 142, 105777 (2022).
Kent, W. J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Zheng, S. C. et al. EpiDISH web server: epigenetic dissection of intra-sample-heterogeneity with online GUI. Bioinformatics https://doi.org/10.1093/bioinformatics/btz833 (2019).
Zheng, S. C. et al. A novel cell-type deconvolution algorithm reveals substantial contamination by immune cells in saliva, buccal and cervix. Epigenomics 10, 925–940 (2018).
Gervin, K. et al. Systematic evaluation and validation of reference and library selection methods for deconvolution of cord blood DNA methylation data. Clin. Epigenetics 11, 125 (2019).
Koestler, D. C. et al. Improving cell mixture deconvolution by identifying optimal DNA methylation libraries (IDOL). BMC Bioinform. 17, 120 (2016).
Salas, L. A. et al. An optimized library for reference-based deconvolution of whole-blood biospecimens assayed using the Illumina HumanMethylationEPIC BeadArray. Genome Biol. 19, 64 (2018).
Ramos-Navarro, C., Sanchez-Luna, M., Zeballos-Sarrato, S. & Pescador-Chamorro, I. Antenatal corticosteroids and the influence of sex on morbidity and mortality of preterm infants. J. Matern. Fetal Neonatal Med. 35, 3438–3445 (2022).
Suarez, A. et al. A polyepigenetic glucocorticoid exposure score at birth and childhood mental and behavioral disorders. Neurobiol. Stress 13, 100275 (2020).
McGowan, P. O. & Matthews, S. G. Prenatal stress, glucocorticoids, and developmental programming of the stress response. Endocrinology 159, 69–82 (2018).
Talge, N. M. et al. Antenatal maternal stress and long-term effects on child neurodevelopment: how and why? J. Child Psychol. Psychiatry 48, 245–261 (2007).
Syed, S. A. & Zannas, A. S. Epigenetics in Psychiatry 2nd edn (eds Peedicayil, J., Grayson, D. R. & Avramopoulos, D.) 701–709 (Academic Press, 2021).
Tsigos, C. & Chrousos, G. P. Hypothalamic–pituitary–adrenal axis, neuroendocrine factors and stress. J. Psychosom. Res. 53, 865–871 (2002).
Criado-Marrero, M. et al. Hsp90 and FKBP51: complex regulators of psychiatric diseases. Philos. Trans. R. Soc. Lond. B Biol. Sci. 373 https://doi.org/10.1098/rstb.2016.0532 (2018).
Somvanshi, P. R. et al. Role of enhanced glucocorticoid receptor sensitivity in inflammation in PTSD: insights from computational model for circadian-neuroendocrine-immune interactions. Am. J. Physiol. Endocrinol. Metab. 319, E48–E66 (2020).
Plieger, T., Felten, A., Splittgerber, H., Duke, É. & Reuter, M. The role of genetic variation in the glucocorticoid receptor (NR3C1) and mineralocorticoid receptor (NR3C2) in the association between cortisol response and cognition under acute stress. Psychoneuroendocrinology 87, 173–180 (2018).
Raffin-Sanson, M. L., de Keyzer, Y. & Bertagna, X. Proopiomelanocortin, a polypeptide precursor with multiple functions: from physiology to pathological conditions. Eur. J. Endocrinol. 149, 79–90 (2003).
DiPietro, J. A. & Voegtline, K. M. The gestational foundation of sex differences in development and vulnerability. Neuroscience 342, 4–20 (2017).
De Coster, S. et al. Gender-specific transcriptomic response to environmental exposure in Flemish adults. Environ. Mol. Mutagen 54, 574–588 (2013).
Gabory, A., Roseboom, T. J., Moore, T., Moore, L. G. & Junien, C. Placental contribution to the origins of sexual dimorphism in health and diseases: sex chromosomes and epigenetics. Biol. Sex Differ. 4, 5 (2013).
McCarthy, N. S. et al. Meta-analysis of human methylation data for evidence of sex-specific autosomal patterns. BMC Genomics 15, 981 (2014).
Rosenfeld, C. S. Sex-specific placental responses in fetal development. Endocrinology 156, 3422–3434 (2015).
Martin, E. et al. Sexual epigenetic dimorphism in the human placenta: implications for susceptibility during the prenatal period. Epigenomics 9, 267–278 (2017).
Clark, J. et al. Associations between placental CpG methylation of metastable epialleles and childhood body mass index across ages one, two and ten in the Extremely Low Gestational Age Newborns (ELGAN) cohort. Epigenetics 14, 1102–1111 (2019).
Watterberg, K. L., Scott, S. M. & Naeye, R. L. Chorioamnionitis, cortisol, and acute lung disease in very low birth weight infants. Pediatrics 99, E6 (1997).
Baud, O. & Watterberg, K. L. Prophylactic postnatal corticosteroids: early hydrocortisone. Semin. Fetal Neonatal Med. 24, 202–206 (2019).
Meakin, C. J. et al. Placental CpG methylation of HPA-axis genes is associated with cognitive impairment at age 10 among children born extremely preterm. Horm. Behav. 101, 29–35 (2018).
Acknowledgements
The authors wish to thank our ECHO Colleagues; the medical, nursing, and program staff; and the children and families participating in the ECHO cohorts. We also acknowledge the contribution of the following ECHO Program collaborators: ECHO Components—Coordinating Center: Duke Clinical Research Institute, Durham, North Carolina: Smith P.B., Newby L.K.; Data Analysis Center: Johns Hopkins University Bloomberg, School of Public Health, Baltimore, Maryland: Jacobson L.P.; Research Triangle Institute, Durham, North Carolina: Catellier D.J.; Person-Reported Outcomes Core: Northwestern University, Evanston, Illinois: Gershon R., Cella D. ECHO Awardees and Cohorts—Baystate Children’s Hospital, Springfield, MA: Vaidya R.; Beaumont Children’s Hospital, Royal Oak, MI: Obeid R.; Boston Children’s Hospital, Boston, MA: Rollins C.; East Carolina University, Brody School of Medicine, Greenville, NC: Bear K.; Michigan State University College of Human Medicine, East Lansing, MI: Lenski M.; Tufts University School of Medicine, Boston, MA: Singh R.; University of Chicago, Chicago, IL: Msall M.; University of Massachusetts Chan Medical School, Worcester, MA: Frazier J.; Atrium Health Wake Forest Baptist, Winston-Salem, NC: Gogcu S.; Yale School of Medicine, New Haven, CT: Montgomery A.; Boston Medical Center, Boston, MA: Kuban K., Douglass L., Jara H.; Boston University, Boston, MA: Joseph R.
Funding
Research reported in this publication was supported by the Environmental influences on Child Health Outcomes (ECHO) Program, Office of the Director, National Institutes of Health, under Award Numbers U2COD023375 (Coordinating Center), U24OD023382 (Data Analysis Center), U24OD023319 with co-funding from the Office of Behavioral and Social Science Research (PRO Core), UH3OD023347 (B.M.L. and C.J.M.) and UH3OD023348 (T.M.O. and R.C.F.). The Neonatal Neurobehavior and Outcomes in Very Preterm Infants (NOVI) cohort was also supported by R01HD072267 and R01HD084515.
Author information
Authors and Affiliations
Consortia
Contributions
T.M.E., C.J.M., T.M.O., R.C.F. and B.M.L. initiated, acquired the funding for, and, along with K.M.H., designed this investigation. K.M.H. and V.Z. performed the statistical analyses and interpreted the results. A.A.B., B.S.C., J.H., J.A.H., E.C.M., C.R.N., S.L.P., L.M.S., S.A.D., and L.M.D. coordinated data collection. K.M.H. and T.M.E. drafted the manuscript. All authors contributed to interpretation of the results and revisions to the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study was approved by local Institutional Review Boards, and participants gave informed consent.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hodge, K.M., Zhabotynsky, V., Burt, A.A. et al. Epigenetic associations in HPA axis genes related to bronchopulmonary dysplasia and antenatal steroids. Pediatr Res (2024). https://doi.org/10.1038/s41390-024-03116-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41390-024-03116-4
This article is cited by
-
Glucocorticosteroids and bronchopulmonary dysplasia : is epigenetics the missing link?
Pediatric Research (2024)