Abstract
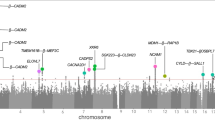
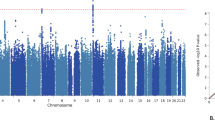
Despite the substantial heritability of antisocial behavior (ASB), specific genetic variants robustly associated with the trait have not been identified. The present study by the Broad Antisocial Behavior Consortium (BroadABC) meta-analyzed data from 28 discovery samples (N = 85,359) and five independent replication samples (N = 8058) with genotypic data and broad measures of ASB. We identified the first significant genetic associations with broad ASB, involving common intronic variants in the forkhead box protein P2 (FOXP2) gene (lead SNP rs12536335, p = 6.32 × 10−10). Furthermore, we observed intronic variation in Foxp2 and one of its targets (Cntnap2) distinguishing a mouse model of pathological aggression (BALB/cJ strain) from controls (BALB/cByJ strain). Polygenic risk score (PRS) analyses in independent samples revealed that the genetic risk for ASB was associated with several antisocial outcomes across the lifespan, including diagnosis of conduct disorder, official criminal convictions, and trajectories of antisocial development. We found substantial genetic correlations of ASB with mental health (depression rg = 0.63, insomnia rg = 0.47), physical health (overweight rg = 0.19, waist-to-hip ratio rg = 0.32), smoking (rg = 0.54), cognitive ability (intelligence rg = −0.40), educational attainment (years of schooling rg = −0.46) and reproductive traits (age at first birth rg = −0.58, father’s age at death rg = −0.54). Our findings provide a starting point toward identifying critical biosocial risk mechanisms for the development of ASB.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Allen JJ, Anderson CA. Aggression and violence: definitions and distinctions. In: The Wiley handbook of violence and aggression, Peter Sturmey (Editor-in-Chief), John Wiley & Sons, New Jersey. American Cancer Society; 2017. p. 1–14.
Rivenbark JG, Odgers CL, Caspi A, Harrington H, Hogan S, Houts RM, et al. The high societal costs of childhood conduct problems: evidence from administrative records up to age 38 in a longitudinal birth cohort. J Child Psychol Psychiatry. 2018;59:703–10.
Brewin CR, Andrews B, Rose S, Kirk M. Acute stress disorder and posttraumatic stress disorder in victims of violent crime. Am J Psychiatry. 1999;156:360–6.
Eme R. Male life-course-persistent antisocial behavior: the most important pediatric mental health problem. Arch Pediatrics Adolesc Med. 2010;164:486–7.
Abram KM, Zwecker NA, Welty LJ, Hershfield JA, Dulcan MK, Teplin LA. Comorbidity and continuity of psychiatric disorders in youth after detention: a prospective longitudinal study. JAMA Psychiatry. 2015;72:84–93.
Loeber R. The stability of antisocial and delinquent child behavior: a review. Child Dev. 1982;53:1431–46.
Jones SE, Miller JD, Lynam DR. Personality, antisocial behavior, and aggression: a meta-analytic review. J Crim Justice. 2011;39:329–37.
Miller JD, Lynam D. Structural models of personality and their relation to antisocial behavior: a meta-analytic review*. Criminology. 2001;39:765–98.
Polderman TJC, Benyamin B, de Leeuw CA, Sullivan PF, van Bochoven A, Visscher PM, et al. Meta-analysis of the heritability of human traits based on fifty years of twin studies. Nat Genet. 2015;47:702–9.
Burt SA. Are there meaningful etiological differences within antisocial behavior? Results of a meta-analysis. Clin Psychol Rev. 2009;29:163–78.
Burt SA, Klump KL. Etiological distinctions between aggressive and non-aggressive antisocial behavior: results from a nuclear twin family model. J Abnorm Child Psychol. 2012;40:1059–71.
Smoller JW, Andreassen OA, Edenberg HJ, Faraone SV, Glatt SJ, Kendler KS. Psychiatric genetics and the structure of psychopathology. Mol Psychiatry. 2019;24:409–20.
Lee PH, Anttila V, Won H, Feng Y-CA, Rosenthal J, Zhu Z, et al. Genomic relationships, novel loci, and pleiotropic mechanisms across eight psychiatric disorders. Cell. 2019;179:1469–82.e11.
Abdellaoui A, Smit DJA, van den Brink W, Denys D, Verweij KJH. Genomic relationships across psychiatric disorders including substance use disorders. Drug Alcohol Depend. 2021;220:108535.
Linnér RK, Mallard TT, Barr PB, Sanchez-Roige S, Madole JW, Driver MN, et al. Multivariate genomic analysis of 1.5 million people identifies genes related to addiction, antisocial behavior, and health. BioRxiv. 2020:2020.10.16.342501.
Tielbeek JJ, Johansson A, Polderman TJC, Rautiainen M-R, Jansen P, Taylor M, et al. Genome-wide association studies of a broad spectrum of antisocial behavior. JAMA Psychiatry. 2017;74:1242–50.
Ip H, Laan C, Brikell I, Sánchez-Mora C, Nolte I, St. Pourcain B, et al. Genetic association study of childhood aggression across raters, instruments and age. 2019.
Pappa I, Pourcain BS, Benke K, Cavadino A, Hakulinen C, Nivard MG, et al. A genome-wide approach to children’s aggressive behavior: the EAGLE consortium. Am J Med Genet Part B Neuropsychiatr Genet. 2016;171:562–72.
Demontis D, Walters RK, Rajagopal VM, Waldman ID, Grove J, Als TD, et al. Risk variants and polygenic architecture of disruptive behavior disorders in the context of attention-deficit/hyperactivity disorder. Nat Commun. 2021;12:576.
Jackson DB, Vaughn MG. Promoting health equity to prevent crime. Prev Med. 2018;113:91–4.
Burt SA, Donnellan MB. Personality correlates of aggressive and non-aggressive antisocial behavior. Pers Individ Differ. 2008;44:53–63.
Piquero AR, Daigle LE, Gibson C, Piquero NL, Tibbetts SG. Research note: Are life-course-persistent offenders at risk for adverse health outcomes? J Res Crime Delinq. 2007;44:185–207.
Tielbeek JJ, Boutwell BB. Exploring the genomic architectures of health, physical traits and antisocial behavioral outcomes: a brief report. Front Psychiatry. 2020;11:539.
Willer CJ, Li Y, Abecasis GR. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics. 2010;26:2190–1.
MacDermot KD, Bonora E, Sykes N, Coupe A-M, Lai CSL, Vernes SC, et al. Identification of FOXP2 truncation as a novel cause of developmental speech and language deficits. Am J Hum Genet. 2005;76:1074–80.
Becker M, Devanna P, Fisher SE, Vernes SC. Mapping of human FOXP2 enhancers reveals complex regulation. Front Mol Neurosci. 2018;11:47.
de Leeuw CA, Mooij JM, Heskes T, Posthuma D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput Biol. 2015;11:e1004219.
den Hoed J, Fisher SE. Genetic pathways involved in human speech disorders. Curr Opin Genet Dev. 2020;65:103–11.
Watanabe K, Stringer S, Frei O, Umićević Mirkov M, de Leeuw C, Polderman TJC, et al. A global overview of pleiotropy and genetic architecture in complex traits. Nat Genet. 2019;51:1339–48.
Bulik-Sullivan BK, Loh P-R, Finucane HK, Ripke S, Yang J, Patterson N, et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet. 2015;47:291–5.
Poulton R, Moffitt TE, Silva PA. The Dunedin multidisciplinary health and development study: overview of the first 40 years, with an eye to the future. Soc Psychiatry Psychiatr Epidemiol. 2015;50:679–93.
Odgers CL, Moffitt TE, Broadbent JM, Dickson N, Hancox RJ, Harrington H, et al. Female and male antisocial trajectories: from childhood origins to adult outcomes. Dev Psychopathol. 2008;20:673–716.
den Hoed J, Devaraju K, Fisher SE. Molecular networks of the FOXP2 transcription factor in the brain. EMBO Rep. 2021;22:e52803.
Lai CSL, Fisher SE, Hurst JA, Vargha-Khadem F, Monaco AP. A forkhead-domain gene is mutated in a severe speech and language disorder. Nature. 2001;413:519–23.
Lai CSL, Gerrelli D, Monaco AP, Fisher SE, Copp AJ. FOXP2 expression during brain development coincides with adult sites of pathology in a severe speech and language disorder. Brain. 2003;126:2455–62.
Uddén J, Hultén A, Bendtz K, Mineroff Z, Kucera KS, Vino A, et al. Toward robust functional neuroimaging genetics of cognition. J Neurosci. 2019;39:8778–87.
Deriziotis P, Fisher SE. Speech and language: translating the genome. Trends Genet. 2017;33:642–56.
Warr-Leeper G, Wright NA, Mack A. Language disabilities of antisocial boys in residential treatment. Behav Disord. 1994;19:159–69.
Cohen NJ, Menna R, Vallance DD, Barwick MA, Im N, Horodezky NB. Language, social cognitive processing, and behavioral characteristics of psychiatrically disturbed children with previously identified and unsuspected language impairments. J Child Psychol Psychiatry. 1998;39:853–64.
Brownlie E, Beitchman J, Escobar M, Young A, Atkinson L, Johnson C, et al. Early language impairment and young adult delinquent and aggressive behavior. J Abnorm Child Psychol. 2004;32:453–67.
Demontis D, Walters RK, Martin J, Mattheisen M, Als TD, Agerbo E, et al. Discovery of the first genome-wide significant risk loci for attention deficit/hyperactivity disorder. Nat Genet. 2019;51:63–75.
Johnson EC, Demontis D, Thorgeirsson TE, Walters, RK, Polimanti, R, et al. A large-scale genome-wide association study meta-analysis of cannabis use disorder. Lancet Psychiatry. 2020;7:1032–45.
Karlsson Linnér R, Biroli P, Kong E, Meddens SFW, Wedow R, Fontana MA, et al. Genome-wide association analyses of risk tolerance and risky behaviors in over 1 million individuals identify hundreds of loci and shared genetic influences. Nat Genet. 2019;51:245–57.
Vernes SC, Newbury DF, Abrahams BS, Winchester L, Nicod J, Groszer M, et al. A functional genetic link between distinct developmental language disorders. N Engl J Med. 2008;359:2337–45.
Medvedeva VP, Rieger MA, Vieth B, Mombereau C, Ziegenhain C, Ghosh T, et al. Altered social behavior in mice carrying a cortical Foxp2 deletion. Hum Mol Genet. 2019;28:701–17.
Caspi A, Houts RM, Ambler A, Danese A, Elliott ML, Hariri A, et al. Longitudinal assessment of mental health disorders and comorbidities across 4 decades among participants in the Dunedin Birth Cohort Study. JAMA Netw Open. 2020;3:e203221.
Turley P, Walters RK, Maghzian O, Okbay A, Lee JJ, Fontana MA, et al. Multi-trait analysis of genome-wide association summary statistics using MTAG. Nat Genet. 2018;50:229–37.
Maier RM, Zhu Z, Lee SH, Trzaskowski M, Ruderfer DM, Stahl EA, et al. Improving genetic prediction by leveraging genetic correlations among human diseases and traits. Nat Commun. 2018;9:989.
Whose genomics? Nat Hum Behav. 2019;3:409–10.
Martin AR, Kanai M, Kamatani Y, Okada Y, Neale BM, Daly MJ. Clinical use of current polygenic risk scores may exacerbate health disparities. Nat Genet. 2019;51:584–91.
Moffitt TE. Adolescence-limited and life-course-persistent antisocial behavior: a developmental taxonomy. 1993.
Moffitt TE. Male antisocial behaviour in adolescence and beyond. Nat Hum Behav.2018;2:177–86.
Jennings WG, Reingle JM. On the number and shape of developmental/life-course violence, aggression, and delinquency trajectories: a state-of-the-art review. J Crim Justice. 2012;40:472–89.
Barnes JC, Beaver K, Boutwell B. Examining the genetic underpinnings of Moffitt’s developmental taxonomy: a behavioral genetic analysis. Criminology. 2011;49:923–54.
Motz RT, Barnes JC, Caspi A, Arseneault L, Cullen FT, Houts R, et al. Does contact with the justice system deter or promote future delinquency? Results from a longitudinal study of British adolescent twins. Criminology. 2020;58:307–35.
Carlisi CO, Moffitt TE, Knodt AR, Harrington H, Ireland D, Melzer TR, et al. Associations between life-course-persistent antisocial behaviour and brain structure in a population-representative longitudinal birth cohort. Lancet Psychiatry. 2020;7:245–53.
Wertz J, Caspi A, Belsky DW, Beckley AL, Arseneault L, Barnes JC, et al. Genetics and crime: integrating new genomic discoveries into psychological research about antisocial behavior. Psychol Sci. 2018;29:791–803.
Tielbeek JJ, Barnes JC, Popma A, Polderman TJC, Lee JJ, Perry JRB, et al. Exploring the genetic correlations of antisocial behaviour and life history traits. BJPsych Open. 2018;4:467–70.
Martin J, Taylor MJ, Lichtenstein P. Assessing the evidence for shared genetic risks across psychiatric disorders and traits. Psychol Med. 2018;48:1759–74.
Harris KM, Halpern CT, Whitsel EA, Hussey JM, Killeya-Jones LA, Tabor J, et al. Cohort profile: the national longitudinal study of adolescent to adult health (add health). Int J Epidemiol. 2019;48:1415–1415k.
Boyd A, Golding J, Macleod J, Lawlor DA, Fraser A, Henderson J, et al. Cohort profile: the ‘children of the 90s’-the index offspring of the Avon longitudinal study of parents and children. Int J Epidemiol. 2013;42:111–27.
Northstone K, Lewcock M, Groom A, Boyd A, Macleod J, Timpson N, et al. The Avon longitudinal study of parents and children (ALSPAC): an update on the enrolled sample of index children in 2019. Wellcome Open Res. 2019;4:51.
Fraser A, Macdonald-Wallis C, Tilling K, Boyd A, Golding J, Davey Smith G, et al. Cohort profile: the Avon longitudinal study of parents and children: ALSPAC mothers cohort. Int J Epidemiol. 2013;42:97–110.
Franke B, Vasquez AA, Veltman JA, Brunner HG, Rijpkema M, Fernández G. Genetic variation in CACNA1C, a gene associated with bipolar disorder, influences brainstem rather than gray matter volume in healthy individuals. Biol Psychiatry. 2010;68:586–8.
Preisig M, Waeber G, Vollenweider P, Bovet P, Rothen S, Vandeleur C, et al. The PsyCoLaus study: methodology and characteristics of the sample of a population-based survey on psychiatric disorders and their association with genetic and cardiovascular risk factors. BMC Psychiatry. 2009;9:9.
The collaborative study on the genetics of alcoholism. Alcohol Health Res World. 1995;19:228–36.
Kaprio J. The Finnish Twin cohort study: an update. Twin Res Hum Genet. 2013;16:157–62.
Johansson A, Jern P, Santtila P, von der Pahlen B, Eriksson E, Westberg L, et al. The genetics of sexuality and aggression (GSA) twin samples in Finland. Twin Res Hum Genet. 2013;16:150–6.
Miller MB, Basu S, Cunningham J, Eskin E, Malone SM, Oetting WS, et al. The Minnesota center for twin and family research genome-wide association study. Twin Res Hum Genet. 2012;15:767–74.
Rujescu D, Hartmann AM, Giegling I, Konte B, Herrling M, Himmelein S, et al. Genome-wide association study in vestibular neuritis: involvement of the host factor for HSV-1 replication. Front Neurol. 2018;9:591.
Mitchell BL, Kirk KM, McAloney K, Wright MJ, Davenport TA, Hermens DF, et al. 16Up: outline of a study investigating wellbeing and information and communication technology use in adolescent twins. Twin Res Hum Genet. 2020;23:345–57.
Mitchell BL, Campos AI, Rentería ME, Parker R, Sullivan L, McAloney K, et al. Twenty-Five and Up (25Up) study: a new wave of the Brisbane longitudinal twin study. Twin Res Hum Genet. 2019;22:154–63.
Morosoli JJ, Colodro-Conde L, Barlow FK, Medland SE. Investigating perceived heritability of mental health disorders and attitudes toward genetic testing in the United States, United Kingdom, and Australia. Am J Med Genet B Neuropsychiatr Genet. 2021. https://doi.org/10.1002/ajmg.b.32875.
Lupton MK, Robinson GA, Adam RJ, Rose S, Byrne GJ, Salvado O, et al. A prospective cohort study of prodromal Alzheimer’s disease: prospective imaging study of ageing: genes, brain and behaviour (PISA). Neuroimage Clin. 2021;29:102527.
Madden PA, Bucholz KK, Dinwiddie SH, Slutske WS, Bierut LJ, Statham DJ, et al. Nicotine withdrawal in women. Addiction. 1997;92:889–902.
Slutske WS, Meier MH, Zhu G, Statham DJ, Blaszczynski A, Martin NG. The Australian Twin Study of Gambling (OZ-GAM): rationale, sample description, predictors of participation, and a first look at sources of individual differences in gambling involvement. Twin Res Hum Genet. 2009;12:63–78.
Knopik VS, Heath AC, Madden PAF, Bucholz KK, Slutske WS, Nelson EC, et al. Genetic effects on alcohol dependence risk: re-evaluating the importance of psychiatric and other heritable risk factors. Psychol Med. 2004;34:1519–30.
Saccone SF, Pergadia ML, Loukola A, Broms U, Montgomery GW, Wang JC, et al. Genetic linkage to chromosome 22q12 for a heavy-smoking quantitative trait in two independent samples. Am J Hum Genet. 2007;80:856–66.
Dick DM, Nasim A, Edwards AC, Salvatore JE, Cho SB, Adkins A, et al. Spit for Science: launching a longitudinal study of genetic and environmental influences on substance use and emotional health at a large US university. Front Genet. 2014;5:47.
McAra L, McVie S. Youth crime and justice: key messages from the Edinburgh study of youth transitions and crime. Criminol Crim Justice. 2010;10:179–209.
Oldehinkel AJ, Rosmalen JG, Buitelaar JK, Hoek HW, Ormel J, Raven D, et al. Cohort profile update: the TRacking Adolescents’ Individual Lives Survey (TRAILS). Int J Epidemiol. 2015;44:76–76n.
Jager A, Amiri H, Bielczyk N, van Heukelum S, Heerschap A, Aschrafi A, et al. Cortical control of aggression: GABA signalling in the anterior cingulate cortex. Eur Neuropsychopharmacol. 2020;30:5–16.
Raczy C, Petrovski R, Saunders CT, Chorny I, Kruglyak S, Margulies EH, et al. Isaac: ultra-fast whole-genome secondary analysis on Illumina sequencing platforms. Bioinformatics. 2013;29:2041–3.
Cingolani P, Platts A, Wang LL, Coon M, Nguyen T, Wang L, et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff. Fly. 2012;6:80–92.
Choi SW, Mak TS-H, O’Reilly PF. Tutorial: a guide to performing polygenic risk score analyses. Nat Protoc. 2020;15:2759–72.
Wray NR, Lin T, Austin J, McGrath JJ, Hickie IB, Murray GK, et al. From basic science to clinical application of polygenic risk scores: a primer. JAMA Psychiatry. 2020. https://doi.org/10.1001/jamapsychiatry.2020.3049.
Wertz J, Agnew-Blais J, Caspi A, Danese A, Fisher HL, Goldman-Mellor S, et al. From childhood conduct problems to poor functioning at age 18 years: examining explanations in a longitudinal cohort study. J Am Acad Child Adolesc Psychiatry. 2018;57:54–60.e4.
Satterthwaite TD, Connolly JJ, Ruparel K, Calkins ME, Jackson C, Elliott MA, et al. The Philadelphia neurodevelopmental cohort: a publicly available resource for the study of normal and abnormal brain development in youth. Neuroimage. 2016;124:1115–9.
Orri M, Boivin M, Chen C, Ahun MN, Geoffroy M-C, Ouellet-Morin I, et al. Cohort profile: Quebec longitudinal study of child development (QLSCD). Soc Psychiatry Psychiatr Epidemiol. 2021;56:883–94.
Boivin M, Brendgen M, Dionne G, Ouellet-Morin I, Dubois L, Pérusse D, et al. The Quebec Newborn Twin Study at 21. Twin Res Hum Genet. 2019;22:475–81.
Price AL, Weale ME, Patterson N, Myers SR, Need AC, Shianna KV, et al. Long-range LD can confound genome scans in admixed populations. Am J Hum Genet. 2008;83:132–5.
Chang CC, Chow CC, Tellier LC, Vattikuti S, Purcell SM, Lee JJ. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience. 2015;4:7.
Zheng J, Erzurumluoglu AM, Elsworth BL, Kemp JP, Howe L, Haycock PC, et al. LD Hub: a centralized database and web interface to perform LD score regression that maximizes the potential of summary level GWAS data for SNP heritability and genetic correlation analysis. Bioinformatics. 2017;33:272–9.
de Moor MHM, Costa PT, Terracciano A, Krueger RF, de Geus EJC, Toshiko T, et al. Meta-analysis of genome-wide association studies for personality. Mol Psychiatry. 2012;17:337–49.
Nagel M, Jansen PR, Stringer S, Watanabe K, de Leeuw CA, Bryois J, et al. Meta-analysis of genome-wide association studies for neuroticism in 449,484 individuals identifies novel genetic loci and pathways. Nat Genet. 2018;50:920–7.
Acknowledgements
GM-V was supported by a Doctoral Research Scholarship from the FRQSC. MB, IO-M, and J-PG are supported by the Canada Research Chair Program. KR is supported by a Sir Henry Wellcome Postdoctoral Fellowship. ADB and DD were supported by grants from the Lundbeck Foundation (R102-A9118, R155-2014-1724 and R248-2017-2003), the EU FP7 Program (Grant No. 602805, “Aggressotype”) and H2020 Program (Grant No. 667302, “CoCA”), NIMH (1U01MH109514-01. JL, AM, and KR acknowledge the following funders: Academy of Finland, the Signe and Ane Gyllenberg foundation, Juho Vainio foundation, Yrjö Jahnsson foundation, Jalmari and Rauha Ahokas foundation, Sigrid Juselius Foundation and The Finnish Society of Sciences and Letters. DD is supported by NIH R01 AA015416 (Finnish Twin Study), P50 AA022537 (Alcohol Research Center), R25 AA027402 (VCU GREAT), R34 AA027347 (Personalized Risk Assessment), R01 AA028064 (Parental Marital Discord, PI: JS), and U10 AA008401 (COGA) from the National Institute on Alcohol Abuse and Alcoholism (NIAAA), and by R01 DA050721 (Externalizing Consortium) from the National Institute on Drug Abuse (NIDA). BF, MK, and NRM were also supported by funding from the European Community’s Horizon 2020 Programme (H2020/2014–2020) under grant agreements no. 728018 (Eat2beNICE) and no. 847879 (PRIME). They also received relevant funding from the Netherlands Organization for Scientific Research (NWO) for the Dutch National Science Agenda NeurolabNL project (grant 400-17-602). HMS and MRM are members of the Medical Research Council (MRC) Integrative Epidemiology Unit at the University of Bristol (MC_UU_00011/7). This work is also supported by the NIHR Biomedical Research Centre at University Hospitals Bristol NHS Foundation Trust and the University of Bristol. AW has been supported by funding from the European Union Seventh Framework Programme (FP7/2007-2013) under grant agreement no. 602768 for the ACTION consortium. JK has been supported by the Academy of Finland Academy professorship program (grants 265240 and 263278). LC-C was supported by a QIMR Berghofer Research Fellowship. JJM was supported by a QIMR Berghofer International PhD Scholarship. ML was funded by an NHMRC Boosting Dementia Leadership Fellowship (APP1140441). SEM and NGM were funded by NHMRC investigator grants (APP1172917 and APP1172990). PJ has been supported by Academy of Finland (Grants no. 319403, 284385 and 274521) and the Stiftelsen för Åbo Akademi Foundation. MK was supported by a personal Rubicon grant (no. 45219212) from the Dutch Research Council. Additional acknowledgements for each study cohort are described in Supplementary Table 1. We acknowledge the work of all members of the Spit for Science Working Group. Spit for Science Director: Danielle M. Dick. Registry management: Kimberly Pedersen, Zoe Neale, Nathaniel Thomas. Data cleaning and management: Amy E. Adkins, Nathaniel Thomas, Zoe Neale, Kimberly Pedersen, Thomas Bannard and Seung B. Cho. Data collection: Amy E. Adkins, Peter Barr, Holly Byers, Erin C. Berenz, Erin Caraway, Seung B. Cho, James S. Clifford, Megan Cooke, Elizabeth Do, Alexis C. Edwards, Neeru Goyal, Laura M. Hack, Lisa J. Halberstadt, Sage Hawn, Sally Kuo, Emily Lasko, Jennifer Lend, Mackenzie Lind, Elizabeth Long, Alexandra Martelli, Jacquelyn L. Meyers, Kerry Mitchell, Ashlee Moore, Arden Moscati, Aashir Nasim, Zoe Neale, Jill Opalesky, Cassie Overstreet, A. Christian Pais, Kimberly Pedersen, Tarah Raldiris, Jessica Salvatore, Jeanne Savage, Rebecca Smith, David Sosnowski, Jinni Su, Nathaniel Thomas, Chloe Walker, Marcie Walsh, Teresa Willoughby, Madison Woodroof and Jia Yan. Genotypical data processing and cleaning: Cuie Sun, Brandon Wormley, Brien Riley, Fazil Aliev, Roseann Peterson and Bradley T. Webb.
Author information
Authors and Affiliations
Consortia
Contributions
Study concept and design: JJT, TJCP, and DP. Acquisition, analysis, or interpretation of data: EU, BSW, LC-C, EG, TTM, BEL, PRJ, AJ, HMS, GP, GRBS, AGA, KR, BK, MK, AMH, JES, IMN, DD, ALKM, SAB, JES, KS, RP, KMH, SV, MM, WGI, NRM, JM, JFV, BLM, JJM, TFMA, IO-M, RET, SMC, JPG, MRB, GD, FV, and MKL. Drafting of the manuscript: JJT, TJCP, PRJ, BBB, TEM, AC, and DP. Study supervision: TJCP, NGM, EC, KR, JGE, JL, CAH, AJO, HS, HL, MP, AW, EV, PJ, YL, DR, IG, TP, JK, KPH, MRM, GM-V, RP, EV, BBB, FA, DD, AP, SVF, ADB, SEM, BF, MB, JBP, JCG, JCB, SEF, TEM, AC, and DP. All authors critically evaluated and revised the manuscript. JJT had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Corresponding author
Ethics declarations
Competing interests
BF has received educational speaking fees from Medice.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tielbeek, J.J., Uffelmann, E., Williams, B.S. et al. Uncovering the genetic architecture of broad antisocial behavior through a genome-wide association study meta-analysis. Mol Psychiatry 27, 4453–4463 (2022). https://doi.org/10.1038/s41380-022-01793-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-022-01793-3
This article is cited by
-
Examining intergenerational risk factors for conduct problems using polygenic scores in the Norwegian Mother, Father and Child Cohort Study
Molecular Psychiatry (2024)
-
Hypothalamic subunit volumes and relations to violence and psychopathy in male offenders with or without a psychotic disorder
European Archives of Psychiatry and Clinical Neuroscience (2024)
-
Dissecting the polygenic contribution of attention-deficit/hyperactivity disorder and autism spectrum disorder on school performance by their relationship with educational attainment
Molecular Psychiatry (2024)
-
Resting heart rate and antisocial behaviour: a Mendelian randomisation study
Scientific Reports (2023)
-
Investigating genetically stratified subgroups to better understand the etiology of alcohol misuse
Molecular Psychiatry (2023)