Abstract
Ubiquitin-specific protease 3 (USP3), a kind of cysteine protease, is a crucial family member of deubiquitinating enzymes. USP3 is aberrantly expressed in several tumors, which may contribute to cancer progression. However, the role of USP3 in gallbladder cancer (GBC) is still unknown. In the current study, we detected the expression of USP3 in GBC tissues, measured its contribution to the cell proliferation in GBC progression, and further studied the underlying mechanism of USP3 in GBC through pyruvate kinase L/R (PKLR; a kind of glycolytic enzyme). We found that the expression of USP3 in GBC tissues were higher than that of adjacent tissues, and the protein levels of USP3 and PKLR were positively correlated. Additionally, overexpressed USP3 significantly promoted cell proliferation in vitro and tumor growth in vivo, while the silencing of USP3 inhibited proliferation and tumor growth. Glycolysis in GBC cells ws promoted by the USP3 overexpression and inhibited bye USP3 downregulation. Moreover, the loss of USP3 promoted the ubiquitination and weakened the stability of PKLR. Results of the rescue assay confirmed that PKLR knockdown suppressed USP3-induced oncogenic activity in USP3 overexpressed GBC cells. These findings imply that USP3 is an essential positive regulator in GBC progression, and USP3-PKLR plays a vital role in the progression and metabolism of GBC.
Similar content being viewed by others
Introduction
Gallbladder cancer (GBC) is a rare, invasive, malignant tumor of the biliary tract with highly variable representation1,2,3. The etiology of this tumor is complex, but it has been reported to be related to gallstones4. Because of late diagnosis and, most patients detect symptoms and signs at an advanced stage, which makes GBC a significant threat to human health5. The lack of specific and sensitive diagnostic markers has significantly impacted the mortality of this disease6. Normally, radical resection is the most effective strategy to treat GBC7. Unfortunately, the number of patients involved in this operational course is hugely limited, and many patients can only receive non-surgical treatment8,9. Some clinical treatments, including conventional chemotherapy, radiotherapy, and molecular targeted therapy render GBC patients with time to relieve; whereas, in the past several years, overall survival has not significantly improved10. Thus, it is an urgent need for effective diagnostic biomarkers and potential therapeutic targets in GBC treatment.
The ubiquitin-proteasome pathway is a crucial intracellular protein degradation regulatory system11. Deubiquitination is an important anti-ubiquitination mechanism catalyzed by deubiquitination enzymes (DUBs). Ubiquitin-specific protease (USP) 3, a kind of cysteine protease belonging to the USP family, is a crucial family member of DUBs12. Recent studies have emphasized the significant meaning of USP3 in tumor progression. USP3 plays a significant role in many biological interactions13,14. It has been reported that USP3 overexpression facilitates cell migration and invasion in gastric cancer12, and USP3 could promote breast cancer cell proliferation by deubiquitinating KLF515. This research illustrates the potential of USP3 in cancer progression, but the role of GBC is not known
Pyruvate kinase L/R (PKLR), a kind of glycolytic enzyme, is considered a key regulator in glycolysis16. It catalyzes the production of pyruvate and ATP from phosphoenolpyruvate. Also, the alteration of energy metabolism is an important hallmark of cancer17. Some researchers have implied that the increment of glycolysis is related to the activation of oncogenes or mutated tumor suppressor genes18,19. Credibly, PKLR can act as a driver of tumor growth and metastasis20. Wang et al. showed that circFOXP1 promotes tumor progression in gallbladder cancer by regulating PKLR expression21, and Nguyen’s et al. found that PKLR promotes colorectal cancer liver colonization through the induction of glutathione synthesis22. These studies indicated the potential role of PKLR in cancer progression by regulating energy metabolism.
In this study, we aimed to investigate the effects of USP3 on the GBC progression. We found that USP3 exhibited a higher expression in GBC tissues than adjacent tissues. USP3 promoted the cell proliferation in vitro and the tumor growth in vivo. We also demonstrated that USP3 was a positive regulator of PKLR, and it deubiquitinated and stabilized PKLR. Further studies on the molecular mechanisms of energy metabolism and progression of GBC may provide new targets and therapeutic approaches for patients with advanced GBC.
Materials and Methods
Clinical samples
This study was approved by the Medical Research Ethics Committee of the First Affiliated Hospital of Zhengzhou University (No. 7, 2019) and carried out following the requirements of the Helsinki declaration. Sixteen pairs of GBC and adjacent specimens were collected in the First Affiliated Hospital of Zhengzhou University from August 2019 to January 2021. GBC tissues were harvested from the center of the tumor tissues. Adjacent tissues were harvested from 1 cm beyond the visible edge of the tumor tissues and further identified by pathological analysis. All participants involved in this study were aware and provided informed consent.
Cell culture and transfection
GBC-SD cells (iCell Bioscience Inc, Shanghai) were cultured in RPMI-1640 medium (Sigma, USA) supplemented with 10% fetal bovine serum (FBS, Sigma, USA) and 5% CO2 at 37 °C. NOZ cells (iCell Bioscience Inc, Shanghai) were cultured in DMEM medium (Sigma, USA) containing 10% FBS and 5% CO2 at 37 °C. After cells adhered to the glass surface, the medium was replaced every 6–8 h.
For transfection, Opti-MEM (Invitrogen, USA) was mixed with lipofectamine 2000 (Invitrogen, USA), and then Opti-MEM was mixed with shRNA-, OE-, or siRNA-, respectively. After 48 h, GBC-SD and NOZ cells were maintained in a completed culture medium containing 400 or 300 g/ml G418 (Sigma, USA)23, respectively. After 7 days of screening, the completed culture medium was replaced for 10 days, and stably transfected cell lines were obtained. In the additive group, 20 μM MG132 (Aladdin, Shanghai) was added15, and the cells were collected and detected 6 h later.
Real-time quantitative PCR
Total RNA was isolated from cultured GBC cells using an RNA extraction kit (BioTeke, Inc. Beijing) according to the manufacturer’s instructions, and RNA was quantified by Nano 2000 spectrophotometry (Thermo, USA). The synthesis of cDNA was performed with total RNA using Super M-MLV reverse transcriptase (BioTeke, Inc. Beijing). The level of mRNA was measured by real-time quantitative PCR (qRT-PCR) analysis based on SYBR Green kit (Sigma, USA) and quantified with the ExicyclerTM 96 apparatus (BIONEER, Korea). The thermal cycling steps were: 94 °C for 5 min, 94 °C for 15 s, 60 °C for 25 s, 72 °C for 30 s, and 40 cycles of 72 °C for 5.5 min, 40 °C for 2.5 min, 60 °C to 94 °C for 34 s, and 25 °C for 1–2 min. USP3 primers were listed as follows (GenScript, Nanjing): forward 5′ TTAGCCCAGAGTCCTTAT 3′and reverse 5′ AGCCGTGACAACAGTAGA 3′. The housekeeping gene β-actin was used as an internal standard.
Western blot
Total protein was extracted from the tumor tissues or whole cells using lysis buffer (Beyotime, Shanghai). Bicinchoninic acid (BCA) assay kit (Beyotime, Shanghai) was utilized for protein quantification. Protein about 20–40 μg was subjected to 10% SDS-PAGE and then transferred to polyvinylidene difluoride membranes. Membranes were blocked with 5% skimmed milk powder (Yili, China) in PBS for 1 h, then, incubated with primary antibodies (1:500, Aceptab, USA) overnight at 4 °C. After washing by TBST buffer, the membranes were incubated with corresponding HRP-linked secondary antibody (1:5000, Beyotime, Shanghai) for 45 min at 37 °C. Finally, the membranes were detected by the ECL Plus reagents (Beyotime, Shanghai) and visualized by Gel-Pro-Analyzer (Media Cybernetics, USA).
Cell proliferation assay
Cells (4 × 103 cell/well) were plated in 96-well plates and cultured at 37 °C overnight. Cell proliferation was guided by the Cell Counting Kit 8 (CCK8) method (Beyotime, Shanghai) after incubating cells for 0, 24, 48, 72, and 96 h. The optical density of each well was measured by a microplate reader (BIOTEK, USA) at 450 nm.
EdU assay
For cell incubation with EdU, cell proliferation was monitored with the cell proliferation EdU image kit (KeyGEN Biotech Corp, Jiangsu) according to the manufacturers’ instructions. After adding Click-iT solution, cells were incubated for 30 min at room temperature in the dark. DAPI solution was used for 5 min to stain the cells. Images were taken under an inverted fluorescent microscope (OLYMPUS, Japan).
Plate colony formation assay
For plate colony formation assay, stably transfected cells were seeded in 35 mm dishes. After 2 weeks, cells were immobilized with 4% polyoxymethylene solution (Sino Biological Inc, Beijing) for 20 min at room temperature. Then, cells were stained with Wright-Giemsa staining solution (Jiancheng Bioengineering Institute, Nanjing) for 5 min, and the colonies were recorded.
Cell cycle analysis
Cell cycle analysis was conducted to GBC-SD and NOZ cells by employing Cell Cycle Detection Kit (KeyGEN Biotech Corp) in accordance with the manufacturers’ instructions. Cells were fixed in 70% ethanol overnight, followed by three washes with PBS. Next, cells were stained with RNase/PI (1:9, v/v) for 30 min in the dark. Cells were subsequently analyzed with a NovoCyte flow cytometer (ACEA Biosciences, San Diego, California, USA).
In vivo tumor formation assay
In vivo tumor formation assay was performed after approval by the Ethical Committee for The First Affiliated Hospital of Zhengzhou University (No. 7, 2019). In this section, mice were fed and housed adaptively for a week. Stable expressed shRNA-USP3 or OE-USP3 GBC-SD cells (1 × 106 per mouse) were injected into subcutaneous tissue. The diameter of tumor tissue was measured, and the volume was calculated every 3 days. After 4 weeks, the mice were sacrificed with 200 mg/kg body weight sodium pentobarbital (Xiya Regent, China) and tumors were removed for further use.
Immunohistochemical method
All of the tissue samples were fixed and embedded in paraffin before being sliced into 5 μm thick sections, followed by deparaffinization and antigen retrieval. The tissue sections were incubated with 0.3% hydrogen peroxide for 15 min and blocked with 10% normal goat serum (Solarbio, Beijing). Then, the slides were incubated with primary antibody (1:100, USP3, Aceptab, USA; Ki67, ABclonal, China) at 4 °C overnight, and then HRP-labeled goat anti-rabbit antibody (1:500, Thermo Fisher, USA) was added at 37 °C for 1 h. After the incubation with DAB substrate solution (Beijing Solarbio), hematoxylin (Solarbio, Beijing) was used for counterstaining. All sections were observed and photographed with a microscope (OLYMPUS, Japan).
Immunofluorescence staining (IF)
Cells were washed with 1 × PBS, fixed with 4% paraformaldehyde, and incubated with 0.1% Triton X-100 (Solarbio, Shanghai, China) for 30 min at room temperature. After adding goat serum, the cells were incubated with primary antibody (USP3, 1:100; Aceptab, USA; PKLR, 1:50, Santa, China) at 4 °C overnight and then incubated with the appropriate secondary antibodies (Cy3 labeled goat anti-rabbit IgG, FITC-labeled goat anti-mouse IgG: 1:200, Beyotime, Shanghai) for 60 min at room temperature. The nuclei were counterstained with DAPI (Aladdin, Shanghai). The IF signals were detected and recorded using an OLYMPUS-BX53 microscope (OLYMPUS).
Co-immunoprecipitation and ubiquitin assay
For co-immunoprecipitation and ubiquitin experiments, total protein was extracted with a BCA kit (Beyotime, Shanghai). PBS was used as the pre-cleaning regent for Protein A/G plus agarose beads. Protein (1 μg/μL, 500 μL) was immunoprecipitated with the indicated IP antibodies (1 μL) at 4 °C overnight, followed by an incubation with 60 µL of protein A/G Plus agarose beads at 4 °C for 2 h. Immunoprecipitates were washed three times with PBS buffer and eluted with SDS sample loading buffer (Beyotime, Shanghai). Proteins were boiled for 5 min and then analyzed by western blotting with indicated antibodies. IP antibodies mentioned above referred to PKLR (IP mouse) antibody and USP3 (IP mouse) antibody. Anti-K48 and anti-K63 ubiquitin antibodies were purchased from Abclonal.
Protein stability analysis
Transfected GBC-SD cells were treated with 10 μg/ml cycloheximide (CHX; MCE, Shanghai, China), after which cells were harvested after 0, 2, 4, 8, and 24 h. Subsequently, cells were lysed by 1 mM PMSF buffer and then subjected to western blot with anti-PKLR as indicated.
Glucose consumption, ATP, pyruvic acid, and lactic acid assays
Glucose consumption was accessed using a glucose assays kit (Jiancheng Bioengineering Institute, Nanjing) according to the manufacturer’s protocol. The product of ATP, pyruvic acid, and lactic acid in the medium was measured respectively with corresponding kits (Jiancheng Bioengineering Institute, Nanjing) according to the specification of the product. BCA kit (Beyotime, Shanghai) was used in this section as the instrument to build the standard curve. The optical density value was measured and used as a standard curve to measure the concentration in the test samples.
Statistical analyzes
The results in this subject were subjected to statistical analysis using Graphpad software version 8.0. Clinical data was analyzed by paired t-test and the others were analyzed by one-way or two-way ANOVA. The correlation of the protein expression between USP3 and PKLR was analyzed by pearson’s correlation. Data were collected from at least three independent experiments, and p < 0.05 was considered to be statistically significant.
Results
USP3 is up-regulated in clinical GBC tissues
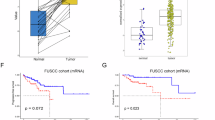
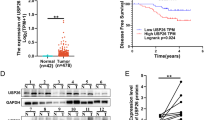
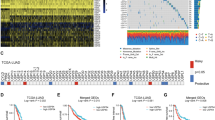
To explore whether USP3 was regulated in GBC, we first analyzed the mRNA and protein expression of USP3 in tumor and adjacent tissues from GBC patients. In Fig. 1A, the results of qRT-PCR showed that the mRNA expression level of USP3 was up-regulated in GBC tissues compared to adjacent tissues. The same results of the changes of the protein levels of USP3 and PKLR were obtained by western blot (Fig. 1B). Furthermore, the data from the Pearson correlation analysis revealed that the protein expression of USP3 and PKLR was positively correlated in GBC tissues and adjacent tissues (Fig. 1C).
A The mRNA expression level of USP3 was tested by qRT-PCR. ※※, p < 0.01. B Protein expression level of USP3 and PKLR was displayed by western blot. A, adjacent tissue. C, cancer tissue. C The correlation between the protein expression of USP3 and PKLR was analyzed by Pearson’s correlation analysis in GBC and adjacent samples.
USP3 affects the proliferation of GBC cells in vitro
Considering the significant association between USP3 expression and GBC tissues, two GBC cell lines, GBC-SD and NOZ cells were transfected with shRNA-USP3 and OE-USP3 to regulate USP3 expression, respectively. As shown in Fig. 2A, B, USP3 successfully realized overexpression and interference in GBC-SD and NOZ cells. We analyzed the role of USP3 in GBC cell proliferation. CCK8 assays showed that the down-regulation of USP3 significantly reduced the viability of GBC-SD and NOZ cells, while overexpression accomplished the opposite result (Fig. 2C). EdU assay was detected additionally. There was a trend that USP3 knockdown exhibited a lower proliferative activity, and the contrary trends were observed in the two cells by USP3 overexpression (Fig. 2D). Consistently, the plate colony formation assay showed that the knockdown of USP3 suppressed the growth capacity of GBC-SD and NOZ cells (Fig. 2E, F). In contrast, the upregulation of USP3 improved the plate colony formation ability of GBC-SD and NOZ cells (Fig. 2E, F). Furthermore, the cell cycle analysis in GBC cell lines were performed by flow cytometry. As revealed in supplemental Fig. 1A, C, the number of cells in G1 phase were increased, accompanied by the reduction of cells in S phase in GBC-SD and NOZ cells with USP3 knockdown. The opposite results were observed by the overexpression of USP3 (supplemental Fig. 1B, D). Therefore, USP3 promoted the proliferation ability of GBC cells.
GBC-SD and NOZ cells were stably transfected with shRNA-USP3-1/2 or OE-USP3, respectively. The overexpression or knockdown efficiency of USP3 in GBC-SD and NOZ cells was assessed by qRT-PCR (A) and western blot (B), respectively. Cell proliferation ability was performed by CCK8 assay (C), EdU (D), and colony formation assay (E, F). Data are expressed as mean ± SD. ns no significant, ※, p < 0.05; ※※, p < 0.01.Scale bars display 50 μm in (D).
USP3 regulates the tumor growth of GBC in vivo
To further test the effects of USP3 on tumorigenicity in vivo, we applied two sets of experiments. First, GBC-SD cells stably transfected with shRNA or overexpressed vector were injected into the nude mice subcutaneously. The volume and size of the tumors formed with USP3-downregulated cells were significantly decreased, while the parameters of the tumors that developed from USP3-overexpressed cells were significantly increased (Fig. 3A).
To further confirm USP3 expression in GBC xenograft tumor tissues, we examined USP3 expression in murine tumor samples using immunohistochemical staining. The results indicated that USP3 was downregulated in tumor of mice injected with USP3-knockdown-GBC-SD cells and was upregulated in tumor with USP3 overexpression (Fig. 3B). Moreover, the expression of tumor proliferation marker Ki67 was increased in tumors with USP3 upregulation and was decreased in tumors with USP3 downregulation (Fig. 3B). These findings revealed that USP3 promoted the in vivo growth of GBC tumors.
USP3 interacts with PKLR and induces deubiquitination of PKLR
We further explored the underlying mechanism of the effects of USP3 on GBC progression. The position of USP3 and PKLR was visualized by a double immunofluorescence staining assay according to a confocal laser scanning microscopy. Figure 4A illustrated that both USP3 (red fluorescence) and PKLR (green fluorescence) were localized in the same regions in GBC-SD and NOZ cells. Merged images in Fig. 4A revealed that the two proteins were co-localized mainly in the nucleus and slightly in the cytoplasm according to overlapping fluorescence signals. Moreover, the interaction between USP3 and PKLR was predicted by HitPredict database (http://www.hitpredict.org/) and Biogrid database (https://thebiogrid.org/). Then, the physical interaction between them was defined by Co-IP in GBC cells (Fig. 4B).
A Double fluorescence staining for USP3 and PKLR in GBC-SD and NOZ cells, with the nuclei counterstained with DAPI. Scale bars represent 25 μm. B The protein of GBC-SD and NOZ cells was immunoprecipitated by an anti-USP3 or anti-PKLR antibody. C OE-USP3 or shRNA-USP3-1/2 was transfected into GBC-SD cells for 48 h. After treatment with 20 mM/L MG132 for 6 h, PKLR was immunoprecipitated with anti-PKLR antibody, and the polyubiquitination of PKLR was examined by western blot using anti-ubiquitin (K48 and K63). D GBC-SD cells were pretreated with CHX 10 μg/mL for 0, 2, 4, 8, and 24 h. Cell lysates were analyzed by western blot. E The protein expression level of PKLR in GBC-SD and NOZ cells. (F) The protein expression level of PKLR and USP3 in tumor tissues of mice.
Since USP3 mediates the removal and processing of ubiquitin24, it was speculated that USP3 might regulate PKLR protein stability through deubiquitination of PKLR. To assess this possibility, the cells were treated with the proteasome inhibitor MG132 to block protein degradation15. Western blot results showed that the K48 and K63-linked ubiquitination of PKLR was suppressed by USP3 overexpression and was enhanced by USP3 knockdown (Fig. 4C). Then, we found that the half-life of PKLR protein was reduced by the loss of USP3 (Fig. 4D). These results clarified that the USP3 depletion promoted the ubiquitination of PKLR and the degradation of the PKLR protein.
To further confirm the regulation of USP3 on PKLR expression, the protein level of USP3 and PKLR was analyzed both in vitro and in vivo. As displayed in Fig. 4E, the protein expression level of PKLR was consistent with USP3. USP3 and PKLR were downregulated in GBC cells transfected with shRNA-USP3-1/2 and were upregulated in GBC cells with USP3 overexpression. Then, similar results were obtained in tumor tissues of xenograft mice (Fig. 4F).
USP3 regulates glycolysis of GBC cells
Considering PKLR has been regarded as a key regulator in the glycolysis procedure, we further determined the role of USP3 in GBC cells during glycolysis. USP3-downregulated GBC cells consumed less glucose (Fig. 5A) and exhibited lower cellular ATP levels (Fig. 5B), lactic acid production (Fig. 5C), and pyruvic acid production (Fig. 5D) than that of cells transfected with shRNA-NC. Adversely, USP3-overexpressed GBC cells caused a clear promotion in glucose consumption, cellular ATP level, lactate production, and pyruvic acid production in GBC cells (Fig. 5A–D). This phenomenon implied that USP3 was involved in glycolysis of GBC cells.
USP3 affected the malignant phenotypes and glycolysis of GBC cells by the regulation of PKLR
We further investigated the role of PKLR depletion in malignant phenotypes in GBC-SD cells with USP3 overexpression. As shown in Fig. 6A, down-regulation of PKLR suppressed the cell proliferation ability of cells with USP3 upregulation. Furthermore, the consumption of glucose and content of lactic acid were reduced by the loss of PKLR in GBC-SD cells transfected with OE-USP3 (Fig. 6B, C). These results suggested that USP3 affected the malignant phenotypes and glycolysis of GBC cells by regulating PKLR.
PKLR was silenced in USP3-overexpressed GBC-SD cells utilizing siRNA. Cell proliferation productivity of GBC-SD cells transfected with OE-USP3 and siRNA-PKLR-1/2 was assessed by CCK8 assay (A). The cellular glucose consumption (B) and lactic acid content (C) were further evaluated. Data are expressed as mean ± SD. ns no significant, ※, p < 0.05; ※※, p < 0.01.
Discussion
Deubiquitinase is a kind of protease that participates in various cellular processes25,26,27,28. Previous research has indicated its feasibility in the molecular pathophysiology of tumors. Li et al.29 found that USP4 inhibits the cell growth of breast cancer through the upregulation of PDCD4. Li et al.30 have shown that USP9X could promote tumor cell survival and confer chemoresistance through YAP1 stabilization. These results all show a positive or negative relationship between USP expression level and malignancy cancers. USP3, a member of the USP subfamily of deubiquitinase, also has been studied in recent years31,32. Wu et al.33 found an increased expression of USP3 in gastric carcinoma patients than that of non-tumor tissues. The similar results were obtained in the current study. Herein, we discovered that GBC tissues exhibited a higher expression level of USP3 compared to adjacent tissues. Our findings indicated a potential relationship between USP3 expression and GBC progression.
We found that many studies have examined the role of deubiquitinating enzymes in cell proliferation in recent years34. Zhang et al.35 discovered that suppression of USP17 inhibits the growth and invasion of lung cancer cells. Chen et al.36 showed that USP9X substrate TTK contributes to tumorigenesis in non-small-cell lung cancer, and the knockdown of USP9X or TTK inhibits cell proliferation, migration, and tumorigenesis. These results indicate the potential role of USP family in various cancer progressions. Therefore, USP3 may affect GBC progression. Fang et al.12 concluded that the overexpression of USP3 accelerates gastric carcinoma cell proliferation and cell cycle progression. USP3 knockdown suppressed breast cancer cell proliferation in HCC cell lines15. Herein, similar results were further elucidated by the promoted proliferation of USP3-upregulated GBC cells in vitro and the facilitated growth of USP3-overexpressed GBC tumor in vivo. The contrary results were observed in USP3-knockdown cells and tumors. Overall, these findings indicate that USP3 promoted the proliferation of GBC cells.
Aerobic glycolysis is a typical metabolic change involved in cancer phenomena, including rapid cell proliferation, invasion, and migration37. Pyruvate kinase family members are vital regulatory enzymes that play critical roles in various malignancies38 and interact with other glycolytic enzymes39. PKLR is a key factor of glycolytic reprogram regulation; for instance, suppression of PKLR affects metabolism40,41. High PKLR expression is correlated with enhanced glucose consumption, increased ATP, and lactic acid production, which attribute to the proliferation and growth of cancer cells. One study26 showed that improved PKLR expression promotes colorectal cancer liver colonization through induction of glutathione synthesis. A rencent paper20 demonstrated that NQO1/PKLR axis promotes the progress of breast cancer via modulating glycolytic reprogramming. PKLR also has been confirmed to play an important role in glycolytic metabolism in GBC cells21. In the present study, PKLR was assumed to interact with USP3, which was later identified by co-IP and fluorescence staining analysis. Interestingly, the K48 and K63-linked ubiquitination of PKLR were suppressed by USP3 overexpression and was enhanced by USP3 overexpression. As is known, K48-linked ubiquitin chains are the most prevalent proteasome-targeting signaling42. Although K63-linked ubiquitin chains have mainly been linked with altering protein complex formation, signaling, or trafficking, there is evidence stating that they can promote proteasomal targeting43,44. Thus, we concluded that the PKLR ubiquitination affected by USP3 was at leastassociated with K48 and K63-linked ubiquitin chains.
Our study revealed that PKLR was a downstream effector of USP3 and regulated glucose metabolism in GBC cells. Knockdown of USP3 caused less glucose consumption and lower cellular ATP levels, lactate production, and pyruvic acid production than the control cells. These results give evidence that the USP3/PKLR plays an important role in cancer progression by modulating glycolytic metabolism in GBC. Furthermore, we found that the loss of PKLR reduced the proliferated curves, content of glucose consumption, and lactic acid in GBC-SD cells with high expression of USP3. Though these in vitro results indicate that USP3 regulates the malignant phenotype of GBC partly by PKLR, in vivo confirmation is still needed, which will be explored in our further investigation.
In this work we concentrate on USP3, an important oncogene, which is overexpressed in various cancers, including GBC. USP3 can promote the cell proliferation of GBC cells in vitro and in vivo. In addition, USP3 is involved in the glycolysis process of GBC cells via PKLR regulation. Overexpressed USP3 stabilized PKLR protein by binding to and deubiquitinating PKLR. These findings indicate that USP3 is an important positive regulator in GBC progression, and USP3-PKLR plays a key role in the progression and metabolism of GBC.
In conclusion, USP3 is highly expressed in GBC tissues. USP3 promotes the GBC progression by regulating the glycolysis, which was mediated as least partly by the deubiquitination and the stability of PKLR.
Data availability
The datasets generated or analyzed during the study are available from the corresponding author upon reasonable request.
References
Wernberg, JA, Lucarelli, DD. Gallbladder cancer. Surg Clin N Am 94, 343–360 (2014).
Tsang ME, Coburn NG, Ridgway PF. Gallbladder Cancer. In: Wright FC, Escallon JE, Cukier M, Tsang ME, Haeed U, editors. Surgical Oncology Manual. New York, NY: Springer, 2016. 115–125.
Acharya, MR, Patkar, S, Parray, A, Goel, M. Management of gallbladder cancer in India. Chin Clin Oncol 8, 35 (2019).
Misra, S, Chaturvedi, A, Misra, NC, Sharma, ID. Carcinoma of the gallbladder. Lancet Oncol 4, 167–176 (2003).
Sharma, A, Sharma, KL, Gupta, A, Yadav, A, Kumar, A. Gallbladder cancer epidemiology, pathogenesis and molecular genetics: Recent update. World J Gastroenterol 23, 3978–3998 (2017).
Tiwari, PK. Epigenetic biomarkers in gallbladder cancer. Trends Cancer 6, 540–543 (2020).
Yadav, S, Tella, SH, Kommalapati, A, Mara, K, Prasai, K, Mady, MH, et al. A novel clinically based staging system for gallbladder cancer. J Natl Compr Canc Ne 18, 151–159 (2020).
Song, X, Hu, Y, Li, Y, Shao, R, Liu, F, Liu, Y. Overview of current targeted therapy in gallbladder cancer. Signal Transduct Tar 5, 230 (2020).
Baiu, I, Visser, B. Gallbladder cancer. JAMA 320, 1294 (2018).
Horgan, AM, Amir, E, Walter, T, Knox, JJ. Adjuvant therapy in the treatment of biliary tract cancer: a systematic review and meta-analysis. J Clin Oncol 30, 1934–1940 (2012).
Yan, C, Su, H, Song, X, Cao, H, Kong, L, Cui, W. Smad Ubiquitination Regulatory Factor 1 (Smurf1) promotes thyroid cancer cell proliferation and migration via ubiquitin-dependent degradation of Kisspeptin-1. Cell Physiol Biochem 49, 2047–2059 (2018).
Fang, CL, Lin, CC, Chen, HK, Hseu, YC, Hung, ST, Sun, DP. et al. Ubiquitin-specific protease 3 overexpression promotes gastric carcinogenesis and is predictive of poor patient prognosis. Cancer Sci 109, 3438–3449 (2018).
Cui, J, Song, Y, Li, Y, Zhu, Q, Tan, P, Qin, Y, et al. USP3 inhibits type I interferon signaling by deubiquitinating RIG-I-like receptors. Cell Res 24, 400–416 (2014).
Cheng, YC, Shieh, SY. Deubiquitinating enzyme USP3 controls CHK1 chromatin association and activation. Proc Natl Acad Sci USA 115, 5546–5551 (2018).
Wu, Y, Qin, J, Li, F, Yang, C, Li, Z, Zhou, Z, et al. USP3 promotes breast cancer cell proliferation by deubiquitinating KLF5. J Biol Chem 294, 17837–17847 (2019).
Nie, H, Li, J, Yang, XM, Cao, QZ, Feng, MX, Xue, F, et al. Mineralocorticoid receptor suppresses cancer progression and the Warburg effect by modulating the miR-338-3p-PKLR axis in hepatocellular carcinoma. Hepatology 62, 1145-1159 (2015).
Hanahan, D, Weinberg, RA. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Deberardinis, RJ, Lum, JJ, Hatzivassiliou, G, Thompson, CB. The biology of cancer: metabolic reprogramming fuels cell growth and proliferation. Cell Metab 7, 11–20 (2008).
Hitosugi, T, Chen, J. Post-translational modifications and the Warburg effect. Oncogene 33, 4279–4285 (2014).
Yang, Y, Zhu, G, Dong, B, Piao, J, Chen, L, Lin, Z. The NQO1/PKLR axis promotes lymph node metastasis and breast cancer progression by modulating glycolytic reprogramming. Cancer Lett 453, 170–183 (2019).
Wang, S, Zhang, Y, Cai, Q, Ma, M, Jin, LY, Weng, M, et al. Circular RNA FOXP1 promotes tumor progression and Warburg effect in gallbladder cancer by regulating PKLR expression. Mol Cancer 18, 145 (2019).
Nguyen, A, Loo, JM, Mital, R, Weinberg, EM, Man, FY, Zeng, Z, et al. PKLR promotes colorectal cancer liver colonization through induction of glutathione synthesis. J Clin Invest 126, 681–694 (2016).
Palmer, AE, Tsien, RY. Measuring calcium signaling using genetically targetable fluorescent indicators. Nat Protoc 1, 1057–1065 (2006).
Lancini, C, Vandenberk, PC, Vissers, JH, Gargiulo, G, Song, JY, Hulsman, D, et al. Tight regulation of ubiquitin-mediated DNA damage response by USP3 preserves the functional integrity of hematopoietic stem cells. J Exp Med 211, 1759–1777 (2014).
Schwickart, M, Huang, X, Lill, JR, Liu, J, Ferrando, R, French, DM, et al. Deubiquitinase USP9X stabilizes MCL1 and promotes tumour cell survival. Nature 463, 103–107 (2010).
Dupont, S, Mamidi, A, Cordenonsi, M, Montagner, M, Zacchigna, L, Adorno, M, et al. FAM/USP9x, a deubiquitinating enzyme essential for TGFbeta signaling, controls Smad4 monoubiquitination. Cell 136, 123–135 (2009).
Nagai, H, Noguchi, T, Homma, K, Katagiri, K, Takeda, K, Matsuzawa, A, et al. Ubiquitin-like sequence in ASK1 plays critical roles in the recognition and stabilization by USP9X and oxidative stress-induced cell death. Mol Cell 36, 805–818 (2009).
Savio, MG, Wollscheid, N, Cavallaro, E, Algisi, V, Difiore, PP, Sigismund, S, et al. USP9X controls EGFR fate by deubiquitinating the endocytic adaptor Eps15. Curr Biol 26, 173–183 (2016).
Li, Y, Jiang, D, Zhang, Q, Liu, X, Cai, Z. Ubiquitin-specific protease 4 inhibits breast cancer cell growth through the upregulation of PDCD4. Int J Mol Med 38, 803–811 (2016).
Li, L, Liu, T, Li, Y, Wu, C, Luo, K, Yin, Y, et al. The deubiquitinase USP9X promotes tumor cell survival and confers chemoresistance through YAP1 stabilization. Oncogene 37, 2422–2431 (2018).
Zhou, Q, Xiao, Z, Zhou, R, Zhou, Y, Fu, P, Li, X, et al. Ubiquitin-specific protease 3 targets TRAF6 for deubiquitination and suppresses IL-1β induced chondrocyte apoptosis. Biochem Bioph Res Co 514, 482–489 (2019).
Rhie BH, Antao AM, Karapurkar JK, Kim MS, Jo WJ, Ramakrishna S, et al. Ubiquitin-specific Protease 3 Deubiquitinates and stabilizes Oct4 protein in human embryonic stem cells. Int J Mol Sci 22, 5584 (2021).
Wu, X, Liu, M, Zhu, H, Wang, J, Dai, W, Li, J. et al. Ubiquitin-specific protease 3 promotes cell migration and invasion by interacting with and deubiquitinating SUZ12 in gastric cancer. J Exp Clin Cancer Res 38, 277 (2019).
McFarlane, C, Kelvin, AA, Delavega, M, Govender, U, Scott, CJ, Burrows, JF, et al. The deubiquitinating enzyme USP17 is highly expressed in tumor biopsies, is cell cycle regulated, and is required for G1-S progression. Cancer Res 70, 3329–3339 (2010).
Zhang, S, Yuan, J, Zheng, R. Suppression of Ubiquitin-Specific Peptidase 17 (USP17) inhibits tumorigenesis and invasion in non-small cell lung cancer cells. Oncol Res 24, 263–269 (2016).
Chen, X, Yu, C, Gao, J, Zhu, H, Cui, B, Zhang, T, et al. A novel USP9X substrate TTK contributes to tumorigenesis in non-small-cell lung cancer. Theranostics 8, 2348–2360 (2018).
Mathupala, SP, Ko, YH, Pedersen, PL. Hexokinase-2 bound to mitochondria: cancer’s stygian link to the “Warburg Effect” and a pivotal target for effective therapy. Semin Cancer Biol 19, 17–24 (2009).
Lim, SO, Li, CW, Xia, W, Lee, HH, Chang, SS, Shen, J, et al. EGFR signaling enhances aerobic glycolysis in triple-negative breast cancer cells to promote tumor growth and immune escape. Cancer Res 76, 1284–1296 (2016).
Fang, R, Xiao, T, Fang, Z, Sun, Y, Li, F, Gao, Y, et al. MicroRNA-143 (miR-143) regulates cancer glycolysis via targeting hexokinase 2 gene. J Biol Chem 287, 23227–23235 (2012).
Laplante, M, Sabatini, DM. mTOR signaling in growth control and disease. Cell 149, 274–293 (2012).
Cavazzoni, A, Bonelli, MA, Fumarola, C, Lamonica, S, Airoud, K, Bertoni, R, et al. Overcoming acquired resistance to letrozole by targeting the PI3K/AKT/mTOR pathway in breast cancer cell clones. Cancer Lett 323, 77–87 (2012).
Saeki, Y. Ubiquitin recognition by the proteasome. J Biochem 161, 113–124 (2017).
Ohtake, F, Tsuchiya, H, Saeki, Y, Tanaka, K. K63 ubiquitylation triggers proteasomal degradation by seeding branched ubiquitin chains. Proc Natl Acad Sci USA 115, e1401–e1408 (2018).
Vijayaraj, SL, Feltham, R, Rashidi, M, Frank, D, Liu, Z, Simpson, DS, et al. The ubiquitylation of IL-1β limits its cleavage by caspase-1 and targets it for proteasomal degradation. Nat Commun 12, 2713 (2021).
Acknowledgements
This research was supported by grants from the Co-operation Research Plan of Medical Science and Technology in Henan Province (Grant no. LHGJ20190149), the Key Scientific Research Projects of Universities in Henan Province (Grant no. 21A320052), and the National Natural Science Foundation of China (Grant no. 82172944).
Author information
Authors and Affiliations
Contributions
Yuling Sun and Ruopeng Liang conceived and founding the experiments; Jie Zhao, Rongtao Zhu, Weijie Wang and Qinwei Lu contributed sample collection/reagents/materials/analysis tools, performed all experiments. Xiaoxue Zhang analyzed the data and wrote the paper. All authors reviewed the manuscript for publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The study was conducted according to the guidelines of the declaration of Helsinki, and approved by the Medical Research Ethics Committee of the First Affiliated Hospital of Zhengzhou University (Approval No. 7, 2019).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liang, RP., Zhang, XX., Zhao, J. et al. Ubiquitin-specific protease 3 facilitates cell proliferation by deubiquitinating pyruvate kinase L/R in gallbladder cancer. Lab Invest 102, 1367–1376 (2022). https://doi.org/10.1038/s41374-022-00836-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-022-00836-1
This article is cited by
-
Ubiquitin specific peptidase 3: an emerging deubiquitinase that regulates physiology and diseases
Cell Death Discovery (2024)
-
USP3 promotes osteosarcoma progression via deubiquitinating EPHA2 and activating the PI3K/AKT signaling pathway
Cell Death & Disease (2024)
-
USP32 deubiquitinase: cellular functions, regulatory mechanisms, and potential as a cancer therapy target
Cell Death Discovery (2023)