Abstract
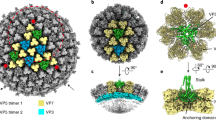
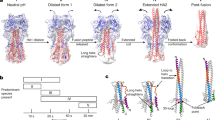
Viruses sense environmental cues such as pH to engage in membrane interactions for cell entry during infection, but how nonenveloped viruses sense pH is largely undefined. Here, we report both high- and low-pH structures of bluetongue virus (BTV), which enters cells via a two-stage endosomal process. The receptor-binding protein VP2 possesses a zinc finger that may function to maintain VP2 in a metastable state and a conserved His866, which senses early-endosomal pH. The membrane-penetration protein VP5 has three domains: dagger, unfurling and anchoring. Notably, the β-meander motif of the anchoring domain contains a histidine cluster that can sense late-endosomal pH and also possesses four putative membrane-interaction elements. Exposing BTV to low pH detaches VP2 and dramatically refolds the dagger and unfurling domains of VP5. Our biochemical and structure-guided-mutagenesis studies support these coordinated pH-sensing mechanisms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Ge, P. & Zhou, Z.H. Chaperone fusion proteins aid entropy-driven maturation of class II viral fusion proteins. Trends Microbiol. 22, 100–106 (2014).
Harrison, S.C. Viral membrane fusion. Nat. Struct. Mol. Biol. 15, 690–698 (2008).
Nibert, M.L., Schiff, L.A. & Field, B.N. in Fields Virology Vol. 2 (eds. Fields, B.N. et al.) 1679–1728 (Lippincott-Raven, 2001).
Settembre, E.C., Chen, J.Z., Dormitzer, P.R., Grigorieff, N. & Harrison, S.C. Atomic model of an infectious rotavirus particle. EMBO J. 30, 408–416 (2011).
Du, J., Bhattacharya, B., Ward, T.H. & Roy, P. Trafficking of bluetongue virus visualized by recovery of tetracysteine-tagged virion particles. J. Virol. 88, 12656–12668 (2014).
Forzan, M., Wirblich, C. & Roy, P. A capsid protein of nonenveloped Bluetongue virus exhibits membrane fusion activity. Proc. Natl. Acad. Sci. USA 101, 2100–2105 (2004).
Grimes, J.M. et al. The atomic structure of the bluetongue virus core. Nature 395, 470–478 (1998).
DeMaula, C.D., Bonneau, K.R. & MacLachlan, N.J. Changes in the outer capsid proteins of bluetongue virus serotype ten that abrogate neutralization by monoclonal antibodies. Virus Res. 67, 59–66 (2000).
Nunes, S.F. et al. A synthetic biology approach for a vaccine platform against known and newly emerging serotypes of bluetongue virus. J. Virol. 88, 12222–12232 (2014).
Yang, Y.-Y., Johnson, T.M., Mecham, J.O., Tam, J.P. & Li, J.K.-K. Epitopic mapping of linear and conformation-dependent antigenic determinants on GP5 of five U.S. bluetongue viruses. Virology 188, 530–536 (1992).
Niesen, F.H., Berglund, H. & Vedadi, M. The use of differential scanning fluorimetry to detect ligand interactions that promote protein stability. Nat. Protoc. 2, 2212–2221 (2007).
Roe, R.R. & Pang, Y.-P. Zinc’s exclusive tetrahedral coordination governed by its electronic structure. J. Mol. Model 5, 134–140 (1999).
Hanas, J.S., Larabee, J.L. & Hocker, J.R. in Zinc Finger Proteins 39–46 (Springer, 2005).
Hanas, J.S., Duke, A.L. & Gaskins, C.J. Conformation states of Xenopus transcription factor IIIA. Biochemistry 28, 4083–4088 (1989).
Huismans, H., van der Walt, N.T., Cloete, M. & Erasmus, B.J. Isolation of a capsid protein of bluetongue virus that induces a protective immune response in sheep. Virology 157, 172–179 (1987).
Esperante, S.A. et al. Fine modulation of the respiratory syncytial virus M2-1 protein quaternary structure by reversible zinc removal from its Cys(3)-His(1) motif. Biochemistry 52, 6779–6789 (2013).
Esperante, S.A., Chemes, L.B., Sánchez, I.E. & de Prat-Gay, G. The respiratory syncytial virus transcription antiterminator M(2-1) is a highly stable, zinc binding tetramer with strong pH-dependent dissociation and a monomeric unfolding intermediate. Biochemistry 50, 8529–8539 (2011).
Zhang, X. et al. Bluetongue virus coat protein VP2 contains sialic acid-binding domains, and VP5 resembles enveloped virus fusion proteins. Proc. Natl. Acad. Sci. USA 107, 6292–6297 (2010).
Hassan, S.H., Wirblich, C., Forzan, M. & Roy, P. Expression and functional characterization of bluetongue virus VP5 protein: role in cellular permeabilization. J. Virol. 75, 8356–8367 (2001).
Bhattacharya, B. & Roy, P. Bluetongue virus outer capsid protein VP5 interacts with membrane lipid rafts via a SNARE domain. J. Virol. 82, 10600–10612 (2008).
Selkrig, J., Leyton, D.L., Webb, C.T. & Lithgow, T. Assembly of β-barrel proteins into bacterial outer membranes. Biochim. Biophys. Acta 1843, 1542–1550 (2014).
Wimley, W.C. The versatile beta-barrel membrane protein. Curr. Opin. Struct. Biol. 13, 404–411 (2003).
Hong, H., Park, S., Jiménez, R.H., Rinehart, D. & Tamm, L.K. Role of aromatic side chains in the folding and thermodynamic stability of integral membrane proteins. J. Am. Chem. Soc. 129, 8320–8327 (2007).
Hayat, S. & Elofsson, A. BOCTOPUS: improved topology prediction of transmembrane β barrel proteins. Bioinformatics 28, 516–522 (2012).
Elaid, S. et al. A peptide derived from the rotavirus outer capsid protein VP7 permeabilizes artificial membranes. Biochim. Biophys. Acta 1838, 2026–2035 (2014).
Biancalana, M., Makabe, K., Koide, A. & Koide, S. Aromatic cross-strand ladders control the structure and stability of β-rich peptide self-assembly mimics. J. Mol. Biol. 383, 205–213 (2008).
Liemann, S., Chandran, K., Baker, T.S., Nibert, M.L. & Harrison, S.C. Structure of the reovirus membrane-penetration protein, Mu1, in a complex with is protector protein, Sigma3. Cell 108, 283–295 (2002).
Zhang, X., Jin, L., Fang, Q., Hui, W.H. & Zhou, Z.H. 3.3 Å cryo-EM structure of a nonenveloped virus reveals a priming mechanism for cell entry. Cell 141, 472–482 (2010).
Yoder, J.D. et al. VP5* rearranges when rotavirus uncoats. J. Virol. 83, 11372–11377 (2009).
Bergelson, J. & Coyne, C. in Viral Entry into Host Cells Vol. 790 (eds. Pöhlmann, S. & Simmons, G.) 24–41 (Springer, 2013).
Suomalainen, M. & Greber, U.F. Uncoating of non-enveloped viruses. Curr. Opin. Virol. 3, 27–33 (2013).
Day, P.M. & Schelhaas, M. Concepts of papillomavirus entry into host cells. Curr. Opin. Virol. 4, 24–31 (2014).
Zhang, X. et al. Cryo-EM structure of the mature dengue virus at 3.5-Å resolution. Nat. Struct. Mol. Biol. 20, 105–110 (2013).
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 10, 584–590 (2013).
Mindell, J.A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003).
Zhang, X., Zhang, X. & Zhou, Z.H. Low cost, high performance GPU computing solution for atomic resolution cryoEM single-particle reconstruction. J. Struct. Biol. 172, 400–406 (2010).
Grigorieff, N. FREALIGN: high-resolution refinement of single particle structures. J. Struct. Biol. 157, 117–125 (2007).
Zhang, X. et al. Near-atomic resolution s electron cryomicroscopy and single-particle reconstruction. Proc. Natl. Acad. Sci. USA 105, 1867–1872 (2008).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Pettersen, E.F. et al. UCSF Chimera: a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Acknowledgements
We thank S. Shivakoti and J. Sun for assistance in cell culture and virus isolation, M. Boyce for advice in sample purification and I. Jones, L. Nguyen and L. Wang for reading the manuscript. This project was supported in part by grants from the US National Institutes of Health (AI094386 to Z.H.Z.) and the Wellcome Trust UK (to P.R.). We acknowledge the use of instruments at the Electron Imaging Center for Nanomachines supported by UCLA and by instrumentation grants from the US National Institutes of Health (1S10OD018111) and the US National Science Foundation (DBI-1338135).
Author information
Authors and Affiliations
Contributions
Z.H.Z., P.R. and X.Z. designed the experiments. A.P. expressed proteins and performed the mutagenesis and biochemical experiments. C.C.C. and X.Y. grew and isolated viruses. X.Z. collected and analyzed the cryoEM data and built the atomic models. Z.H.Z., X.Z. and P.R. interpreted the structures and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 CryoEM imaging, reconstruction and validation.
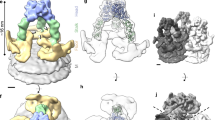
(a) Representative cryoEM DED image of BTV virions. (b) Fourier shell correlation coefficients (FSC) as a function of spatial frequency between maps from half datasets. (c-d) Representative regions of the virion reconstructions showing matching of side-chain features of VP3 (c) and VP7 (d) between the cryoEM density (grey) and x-ray models (ribbons). (e-f) CryoEM reconstructions of the BTV virion at pH3.4 (e) and pH5.5 (f) conditions, both low-pass filtered to 25Å resolution, showing similar structural features. The radial color scheme is the same as that in the Figure 1a.
Supplementary Figure 2 Sequences and secondary structures of VP2 and VP5.
Sequences and secondary structures of VP2 (a) and VP5 (b). α-Helices are marked by cylinders, β-strands by arrows, loops by thin lines. The flexible tip domain (P191-K405) of VP2 are indicated by a dashed line in (a).
Supplementary Figure 3 Structure of VP5.
(a) Conservation of VP5. The bar indicates the color scale of the conservation from most conserved (1, blue) to no conservation (0, white). (b) The internal surface of the β-meander core shown as a ribbon, with the side chains represented as sticks. (c) Hydrophilicity surface of the β-meander core of the VP5 anchoring domain. Gray: hydrophobic; red: oxygen atoms; blue: nitrogen atoms. (d) Possible transmembrane β-strands of VP5 anchoring domain. BOCTOPUS server predicts six transmembrane β-strands in the anchoring domain. HMM: Hidden Markov model; SVM: Support Vector Machines.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 (PDF 721 kb)
Structure of the BTV virion
Radially colored surface representation of the cryoEM reconstruction of the BTV virion at 3.5 Å resolution (MP4 10643 kb)
Structure of VP2
Surface representation of the density map of a VP2 monomer colored by domains: body (red), hairpin (green), and hub (blue). At the relatively high density threshold used, the flexible tip domain is invisible (MP4 10845 kb)
Zinc finger of VP2
Part of VP2 density map (mesh) containing the zinc-finger motif. The atomic model is superimposed into the map as sticks (MP4 5684 kb)
Structure of VP5
Surface representation of the density map of a VP5 monomer colored by domains: dagger (blue), unfurling (red) and anchoring (green). The density of the stem helix α3 in the unfurling domain is marked cyan as in Figure 3 (MP4 7201 kb)
Density map of a helix in VP5
Validation of the cryoEM map. Density map of the α17 helix of the VP5 is shown as semi-transparent surface (cyan) and is superimposed with the atomic model (sticks). Note that the side chains of amino acid residues match the densities of the cryoEM map (MP4 7227 kb)
Histidine cluster of VP5
The histidine cluster located at the interface between the anchoring and unfurling domains of VP5. Density map is shown as semi-transparent surface (grey) and is superimposed with the atomic model (ribbons and sticks) (MP4 5639 kb)
Rights and permissions
About this article
Cite this article
Zhang, X., Patel, A., Celma, C. et al. Atomic model of a nonenveloped virus reveals pH sensors for a coordinated process of cell entry. Nat Struct Mol Biol 23, 74–80 (2016). https://doi.org/10.1038/nsmb.3134
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3134
This article is cited by
-
Penetration of non-enveloped viruses
Nature Microbiology (2021)
-
The native structure of the assembled matrix protein 1 of influenza A virus
Nature (2020)
-
Assembly intermediates of orthoreovirus captured in the cell
Nature Communications (2020)
-
Vector competence is strongly affected by a small deletion or point mutations in bluetongue virus
Parasites & Vectors (2019)
-
Double-stranded RNA virus outer shell assembly by bona fide domain-swapping
Nature Communications (2017)