Abstract
ATP-dependent chromatin remodeling is an essential process required for the dynamic organization of chromatin structure. Here we describe the genome-wide location and activity of three remodeler proteins with diverse physiological functions in the mouse genome: Brg1, Chd4 and Snf2h. The localization patterns of all three proteins substantially overlap with one another and with regions of accessible chromatin. Furthermore, using inducible mutant variants, we demonstrate that the catalytic activity of these proteins contributes to the remodeling of chromatin genome wide and that each of these remodelers can independently regulate chromatin reorganization at distinct sites. Many regions require the activity of more than one remodeler to regulate accessibility. These findings provide a dynamic view of chromatin organization and highlight the differential contributions of remodelers to chromatin maintenance in higher eukaryotes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
Change history
29 January 2014
In the version of this article initially published, the ChIP-seq data were not available in a public repository. The error has been corrected in the HTML and PDF versions of the article.
References
Bork, P. & Koonin, E.V. An expanding family of helicases within the 'DEAD/H' superfamily. Nucleic Acids Res. 21, 751–752 (1993).
Xi, H. et al. Identification and characterization of cell type-specific and ubiquitous chromatin regulatory structures in the human genome. PLoS Genet. 3, e136 (2007).
Boyle, A.P. et al. High-resolution mapping and characterization of open chromatin across the genome. Cell 132, 311–322 (2008).
Hesselberth, J.R. et al. Global mapping of protein-DNA interactions in vivo by digital genomic footprinting. Nat. Methods 6, 283–289 (2009).
John, S. et al. Chromatin accessibility pre-determines glucocorticoid receptor binding patterns. Nat. Genet. 43, 264–268 (2011).
Narlikar, G.J., Phelan, M.L. & Kingston, R.E. Generation and interconversion of multiple distinct nucleosomal states as a mechanism for catalyzing chromatin fluidity. Mol. Cell 8, 1219–1230 (2001).
Rippe, K. et al. DNA sequence- and conformation-directed positioning of nucleosomes by chromatin-remodeling complexes. Proc. Natl. Acad. Sci. USA 104, 15635–15640 (2007).
Blosser, T.R., Yang, J.G., Stone, M.D., Narlikar, G.J. & Zhuang, X. Dynamics of nucleosome remodelling by individual ACF complexes. Nature 462, 1022–1027 (2009).
van Vugt, J.J. et al. Multiple aspects of ATP-dependent nucleosome translocation by RSC and Mi-2 are directed by the underlying DNA sequence. PLoS ONE 4, e6345 (2009).
Boyer, L.A. et al. Functional delineation of three groups of the ATP-dependent family of chromatin remodeling enzymes. J. Biol. Chem. 275, 18864–18870 (2000).
Agalioti, T. et al. Ordered recruitment of chromatin modifying and general transcription factors to the IFN-β promoter. Cell 103, 667–678 (2000).
Bottomley, M.J. Structures of protein domains that create or recognize histone modifications. EMBO Rep. 5, 464–469 (2004).
Alenghat, T., Yu, J. & Lazar, M.A. The N-CoR complex enables chromatin remodeler SNF2H to enhance repression by thyroid hormone receptor. EMBO J. 25, 3966–3974 (2006).
Hogan, C. & Varga-Weisz, P. The regulation of ATP-dependent nucleosome remodelling factors. Mutat. Res. 618, 41–51 (2007).
Ho, L. et al. An embryonic stem cell chromatin remodeling complex, esBAF, is an essential component of the core pluripotency transcriptional network. Proc. Natl. Acad. Sci. USA 106, 5187–5191 (2009).
Schnetz, M.P. et al. CHD7 targets active gene enhancer elements to modulate ES cell-specific gene expression. PLoS Genet. 6, e1001023 (2010).
Sala, A. et al. Genome-wide characterization of chromatin binding and nucleosome spacing activity of the nucleosome remodelling ATPase ISWI. EMBO J. 30, 1766–1777 (2011).
Clapier, C.R. & Cairns, B.R. The biology of chromatin remodeling complexes. Annu. Rev. Biochem. 78, 273–304 (2009).
Jothi, R., Cuddapah, S., Barski, A., Cui, K. & Zhao, K. Genome-wide identification of in vivo protein-DNA binding sites from ChIP-Seq data. Nucleic Acids Res. 36, 5221–5231 (2008).
Boyle, A.P. et al. High-resolution genome-wide in vivo footprinting of diverse transcription factors in human cells. Genome Res. 21, 456–464 (2011).
Peterson, C.L. & Workman, J.L. Promoter targeting and chromatin remodeling by the SWI/SNF complex. Curr. Opin. Genet. Dev. 10, 187–192 (2000).
Goldmark, J.P., Fazzio, T.G., Estep, P.W., Church, G.M. & Tsukiyama, T. The Isw2 chromatin remodeling complex represses early meiotic genes upon recruitment by Ume6p. Cell 103, 423–433 (2000).
Schultz, D.C., Friedman, J.R. & Rauscher, F.J. III. Targeting histone deacetylase complexes via KRAB-zinc finger proteins: the PHD and bromodomains of KAP-1 form a cooperative unit that recruits a novel isoform of the Mi-2α subunit of NuRD. Genes Dev. 15, 428–443 (2001).
Bailey, T.L., Williams, N., Misleh, C. & Li, W.W. MEME: discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res. 34, W369–W373 (2006).
Rao, M. et al. Inhibition of cyclin D1 gene transcription by Brg-1. Cell Cycle 7, 647–655 (2008).
Biddie, S.C. et al. Transcription factor AP1 potentiates chromatin accessibility and glucocorticoid receptor binding. Mol. Cell 43, 145–155 (2011).
Biddie, S.C., John, S. & Hager, G.L. Genome-wide mechanisms of nuclear receptor action. Trends Endocrinol. Metab. 21, 3–9 (2010).
Ishihara, K., Oshimura, M. & Nakao, M. CTCF-dependent chromatin insulator is linked to epigenetic remodeling. Mol. Cell 23, 733–742 (2006).
Richmond, E. & Peterson, C.L. Functional analysis of the DNA-stimulated ATPase domain of yeast SWI2/SNF2. Nucleic Acids Res. 24, 3685–3692 (1996).
Corona, D.F. et al. ISWI is an ATP-dependent nucleosome remodeling factor. Mol. Cell 3, 239–245 (1999).
de la Serna, I.L., Carlson, K.A. & Imbalzano, A.N. Mammalian SWI/SNF complexes promote MyoD-mediated muscle differentiation. Nat. Genet. 27, 187–190 (2001).
Hakimi, M.A. et al. A chromatin remodelling complex that loads cohesin onto human chromosomes. Nature 418, 994–998 (2002).
Dirscherl, S.S., Henry, J.J. & Krebs, J.E. Neural and eye-specific defects associated with loss of the imitation switch (ISWI) chromatin remodeler in Xenopus laevis. Mech. Dev. 122, 1157–1170 (2005).
Srinivasan, R., Mager, G.M., Ward, R.M., Mayer, J. & Svaren, J. NAB2 represses transcription by interacting with the CHD4 subunit of the nucleosome remodeling and deacetylase (NuRD) complex. J. Biol. Chem. 281, 15129–15137 (2006).
Schnitzler, G., Sif, S. & Kingston, R.E. Human SWI/SNF interconverts a nucleosome between its base state and a stable remodeled state. Cell 94, 17–27 (1998).
Moshkin, Y.M. et al. Remodelers organize cellular chromatin by counteracting intrinsic histone-DNA sequence preferences in a class-specific manner. Mol. Cell. Biol. 32, 675–688 (2012).
Gkikopoulos, T. et al. A role for Snf2-related nucleosome-spacing enzymes in genome-wide nucleosome organization. Science 333, 1758–1760 (2011).
Yen, K., Vinayachandran, V., Batta, K., Koerber, R.T. & Pugh, B.F. Genome-wide nucleosome specificity and directionality of chromatin remodelers. Cell 149, 1461–1473 (2012).
Ramirez-Carrozzi, V.R. et al. Selective and antagonistic functions of SWI/SNF and Mi-2β nucleosome remodeling complexes during an inflammatory response. Genes Dev. 20, 282–296 (2006).
Gao, H. et al. Opposing effects of SWI/SNF and Mi-2/NuRD chromatin remodeling complexes on epigenetic reprogramming by EBF and Pax5. Proc. Natl. Acad. Sci. USA 106, 11258–11263 (2009).
Yildirim, O. et al. Mbd3/NURD complex regulates expression of 5-hydroxymethylcytosine marked genes in embryonic stem cells. Cell 147, 1498–1510 (2011).
Curtis, C.D. & Griffin, C.T. The chromatin-remodeling enzymes BRG1 and CHD4 antagonistically regulate vascular Wnt signaling. Mol. Cell. Biol. 32, 1312–1320 (2012).
Kassabov, S.R., Henry, N.M., Zofall, M., Tsukiyama, T. & Bartholomew, B. High-resolution mapping of changes in histone-DNA contacts of nucleosomes remodeled by ISW2. Mol. Cell. Biol. 22, 7524–7534 (2002).
Nagaich, A.K., Walker, D.A., Wolford, R.G. & Hager, G.L. Rapid periodic binding and displacement of the glucocorticoid receptor during chromatin remodeling. Mol. Cell 14, 163–174 (2004).
Boeger, H., Griesenbeck, J. & Kornberg, R.D. Nucleosome retention and the stochastic nature of promoter chromatin remodeling for transcription. Cell 133, 716–726 (2008).
Boeger, H., Griesenbeck, J., Strattan, J.S. & Kornberg, R.D. Removal of promoter nucleosomes by disassembly rather than sliding in vivo. Mol. Cell 14, 667–673 (2004).
Johnson, T.A., Elbi, C., Parekh, B.S., Hager, G.L. & John, S. Chromatin remodeling complexes interact dynamically with a glucocorticoid receptor regulated promoter. Mol. Biol. Cell 19, 3308–3322 (2008).
McKnight, J.N., Jenkins, K.R., Nodelman, I.M., Escobar, T. & Bowman, G.D. Extranucleosomal DNA binding directs nucleosome sliding by Chd1. Mol. Cell. Biol. 31, 4746–4759 (2011).
Rigaud, G., Roux, J., Pictet, R. & Grange, T. In vivo footprinting of rat TAT gene: dynamic interplay between the glucocorticoid receptor and a liver-specific factor. Cell 67, 977–986 (1991).
Voss, T.C. et al. Dynamic exchange at regulatory elements during chromatin remodeling underlies assisted loading mechanism. Cell 146, 544–554 (2011).
Won, K.J. et al. An integrated approach to identifying cis-regulatory modules in the human genome. PLoS ONE 4, e5501 (2009).
Bultman, S. et al. A Brg1 null mutation in the mouse reveals functional differences among mammalian SWI/SNF complexes. Mol. Cell 6, 1287–1295 (2000).
Siatecka, M., Xue, L. & Bieker, J.J. Sumoylation of EKLF promotes transcriptional repression and is involved in inhibition of megakaryopoiesis. Mol. Cell. Biol. 27, 8547–8560 (2007).
Hsiao, P.W., Fryer, C.J., Trotter, K.W., Wang, W. & Archer, T.K. BAF60a mediates critical interactions between nuclear receptors and the BRG1 chromatin-remodeling complex for transactivation. Mol. Cell. Biol. 23, 6210–6220 (2003).
Sims, R.J. III et al. Human but not yeast CHD1 binds directly and selectively to histone H3 methylated at lysine 4 via its tandem chromodomains. J. Biol. Chem. 280, 41789–41792 (2005).
Hassan, A.H. et al. Function and selectivity of bromodomains in anchoring chromatin-modifying complexes to promoter nucleosomes. Cell 111, 369–379 (2002).
Fragoso, G., Pennie, W.D., John, S. & Hager, G.L. The position and length of the steroid-dependent hypersensitive region in the mouse mammary tumor virus long terminal repeat are invariant despite multiple nucleosome B frames. Mol. Cell. Biol. 18, 3633–3644 (1998).
Voss, T.C. et al. Combinatorial probabilistic chromatin interactions produce transcriptional heterogeneity. J. Cell Sci. 122, 345–356 (2009).
John, S. et al. Interaction of the glucocorticoid receptor with the global chromatin landscape. Mol. Cell 29, 611–624 (2008).
Siersbæk, R. et al. Extensive chromatin remodelling and establishment of transcription factor 'hotspots' during early adipogenesis. EMBO J. 30, 1459–1472 (2011).
Baek, S., Sung, M.H. & Hager, G.L. Quantitative analysis of genome-wide chromatin remodeling. Methods Mol. Biol. 833, 433–441 (2012).
Bailey, T.L. & Gribskov, M. Combining evidence using p-values: application to sequence homology searches. Bioinformatics 14, 48–54 (1998).
Acknowledgements
The authors thank D. Picketts (University of Ottawa) and J. Svaren (University of Wisconsin) for the kind gift of remodeler cDNA constructs (hSNF2H and mChd4, respectively), A. Indrawan for technical assistance, the US National Cancer Institute Advanced Technology Program Sequencing Facility for sequencing services and Epitomics, Inc. for generation of the monoclonal rabbit antibody to BRG1. This research was supported by the Intramural Research Program of the US National Institutes of Health, National Cancer Institute Center for Cancer Research and by a UNCF/Merck Postdoctoral Science Research Fellowship to S.A.M.
Author information
Authors and Affiliations
Contributions
S.A.M. and G.L.H. conceived of and designed the study. S.A.M., M.W., T.A.J. and R.L.S. performed the experiments. S.J. provided technical advice. S.B. and M.-H.S. conducted the bioinformatics analysis. S.A.M. performed the experimental analysis and data interpretation. S.A.M. and G.L.H. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Detection and characterization of remodeler-binding sites.
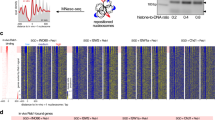
(a) Left-side, Western blot detection of Brg1 in nuclear extracts with monoclonal antibody. Right-side, top graph, ChIP-QPCR analysis of initial Brg1 antiserum at regions bound by Brg1 (Orm2 and Fabp4) and not bound by Brg1 (tRNA). Right-side, bottom graph, ChIP-QPCR analysis of final monoclonal Brg1 antibody at the same regions described above. (b) Specificity of Brg1, Chd4 and Snf2h antibodies by Western blot analysis of whole cell extracts. MW, molecular weight markers. Asterisk, unidentified band that does not appear following immunoprecipitation with the Chd4 antibody. (c) Additional examples of ChIP-seq genome browser views of Brg1, Chd4, and Snf2h occupancy. Images represent tag densities relative to genome coordinates. Input tracks of sonicated, genomic DNA are aligned below each ChIP-seq track. (d) Box plots of tag density enrichment (log2 scale) of remodeler binding sites located at the indicated genomic regions. Sites are classified as promoter (< 2.5 kb upstream or downstream from the nearest TSS), exon (> 2.5 kb downstream from the nearest TSS , overlapping an exon), intron (> 2.5 kb downstream from the nearest TSS , in an intron not overlapping an exon), distal upstream (> 2.5 kb upstream from the nearest TSS), or downstream (> 2.5 kb downstream from the nearest TSS).
Supplementary Figure 2 Co-occupancy of remodeler proteins.
(a) Additional example of ChIP-seq genome browser view of Brg1, Chd4, and Snf2h occupancy at the same genomic coordinates on chromosome 4. Tag density is indicated on the y-axis. (b) ChIP-QPCR analysis of Brg1, Chd4, and Snf2h at the indicated genomic regions. Bars represent the average of two replicates normalized to input DNA. No Ab, no antibody control; tRNA, negative control. Error bars represent the SEM of two independent replicates. (c-e) Genome browser view of regions analyzed in panel [(b)]. Sites amplified are highlighted in grey. Input track is included in the alignment.
Supplementary Figure 3 Interaction of remodeler proteins.
(a) To detect potential interactions of soluble complexes, Western blot analysis of co-immunoprecipitation (Co-IP) experiments was performed using nuclear extracts and Brg1, Snf2h, and Chd4 specific antibodies. As a control, known protein interactions were analyzed by Western (Baf155 (Brg1-associated), Wstf (Snf2h-associated), and Hdac1 (Chd4-associated). Input, 15 μg of IP nuclear extract; IP, immunoprecipitation; and control IP, lysate from 'no antibody' control sample. (b) Aggregate plot of the average tag density of Brg1 binding sites (hotspot) at regions occupied by: Brg1, Chd4, and Snf2h (black), Brg1 and Chd4 only (brown), Brg1 and Snf2h only (light blue), or Brg1 only (pink). (c) Same as in [(b)], except for Chd4 binding sites. (d) Same as in [(b and c)] for Snf2h binding sites. (e) Potential remodeler interactions at the template were examined by sequential ChIP (re-ChIP) analysis. Primary ChIP with anti-Brg1, anti-Snf2h, and anti-Chd4 was performed at the Orm2 and Ccl2 sites described in Supplementary Fig. 2b. (f) Positive control; secondary re-ChIP was performed with the anti-Brg1 IP from panel (e). (g) Secondary re-ChIP was performed with anti-Snf2h and anti-Chd4 on the anti-Brg1 IP from panel (e). Glul is a negative control site where none of the remodeler proteins were detected.
Supplementary Figure 4 De novo motif analysis of unique remodeler-binding sites.
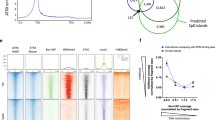
(a) Shown are the most significantly enriched motifs associated with Brg1 unique sites (not bound by Chd4 or Snf2h) identified by MEME analysis (P < 10-4) using the top 2,000 sites . The AP-1 motif was found to be the most highly enriched (MEME E value = 1.4e-1841). (b) Venn diagrams representing the overlap of binding sites for Chd4 or Snf2h with AP-1 sites. (c-d) Similar de novo motif analysis as above was performed on Chd4 and Snf2h unique sites. The CTCF motif was found to be the most highly enriched motif for Chd4 unique sites (MEME E value = 3.6e-815) and Snf2h unique sites (MEME E value = 1.1e-1026). (e) Venn diagrams of sites shared between remodelers and CTCF. Left-side, Venn diagram of the overlap between Brg1 and CTCF sites. Right-side, three-way Venn diagram representing the overlap between remodeler sites that specifically co-localize with CTCF sites. (f) Lost and gained DHS sites; Left-side, distribution at annotated genic regions of DHS sites lost following the expression of dnBrg1. Bottom; Shown are the most significantly enriched motifs identified by MEME analysis (P < 10-4) using the top 1,000 DHS sites (based on tag density) lost following dnBrg1 expression. The motif identified as AP-1 was found to be the most highly enriched motif (MEME E value = 1.8e-184). (g) Lost and gained DHS sites; Right-side, distribution at annotated genomic regions of DHS sites gained following the expression of dnChd4. Bottom, the most significantly enriched motifs identified using the top 1,000 DHS sites (based on tag density) gained following dnChd4 expression. The motif identified as Sp1 was found to be the most highly enriched motif (MEME E value = 6.5e-167).
Supplementary Figure 5 Enrichment levels of overlapping remodeler and DHS sites.
(a) Additional example of genome browser view displaying Brg1, Chd4, and Snf2h ChIP-seq occupancy and DNase I hypersensitivity patterns at a region on chromosome 6. (b-d) Box plots of tag density enrichment (log2 scale) of the indicated remodeler binding sites overlapping DNase I hypersensitive sites (DHS) and remodeler binding sites not associated with DHS sites. (e) Box plot of tag density enrichment (log2 scale) of DHS sites overlapping remodeler binding sites (Remodeler) and DHS sites not associated with remodeler binding. The number of each site type is indicated below the corresponding box plot. (f) Aggregate plot of the average tag density of DHS sites (hotspot) at regions occupied by: all remodelers (any/all combinations of remodeler binding, black), no remodelers (red), Brg1 only (light blue), Chd4 only (brown), or Snf2h only (pink).
Supplementary Figure 6 Characterization of the Inducible dn-remodeler system.
(a) Top, alignment of hBRG1, mouse Chd4, and hSNF2H ATPase domain region containing a conserved lysine (K) residue essential for catalytic activity. Bottom, schematic representation of dominant-negative remodeler constructs. Each remodeler contains either a single or triple FLAG tag at its N- or C-terminus for the monitoring of expression. Highlighted is the mutation of the conserved lysine within the ATPase domain to either arginine (R) or cysteine (C). (b) Schematic of tetracycline-regulated expression system. Each dominant-negative remodeler construct was stably transfected into cells containing the tetracycline transactivator (tTA) system. The tTA construct is positioned downstream of a tetracycline (Tet) response element (TRE) bound to a minimal CMV promoter and is itself regulated by the tTA protein (Vp16-TetR) providing autoinducible control. (c) Western blot analysis of dominant-negative (dn) protein expression derived from whole cell extracts of cultured cells in the presence or absence of tetracycline (Tet). Each remodeler was FLAG-tagged at either the N- or C-terminus and detected using a FLAG-specific antibody. As a positive control, the expression of tetracycline transactivator (tTA) was analyzed (only expressed in the absence of Tet). Actin was analyzed to determine equal loading. (d) mRNA expression of dominant-negative remodelers in the presence (+Tet, white bars) or absence of tetracycline (-Tet, black bars). Expression is shown relative to the +Tet condition (no expression) with normalization to Actin expression. Error bars represent the SEM of three independent replicates. (e-g) Effect of dn-remodeler expression on levels of alternate remodeling proteins. Western blot analysis of selected remodeler expression levels with activation of alternate dn factors. Nuclear extracts from cells either expressing (- tet), or not expressing (+tet, 48 hr.), FLAG-tagged versions of dnBrg1, dnSnf2h, and dnChd4 were tested for effects on the other remodeling proteins. Input, 15 μg of IP nuclear extract; western blots were developed with antibodies specific to FLAG, Brg1, Chd4, Snf2h, the tet regulator, or GAPDH as loading control. Black arrows indicate position of size markers; blue arrows indicated position of factor tested. (e) Cells expressing dnCHD4; Brg1 and Snf2h levels are unaffected. (f) Cells expressing dnBrg1; Chd4 and Snf2h levels are unaffected. Strong induction of the tet regulator is also shown at the top of the panel. (g) Cells expressing dn Snf2h; Brg1 and Chd4 levels are unaffected.
Supplementary Figure 7 Remodeler binding at affected DHS sites.
(a-c) Genome browser views of ChIP-seq remodeler binding at DNase I hypersensitive (DHS, DNaseI-seq) sites affected by dominant-negative remodeler expression. In each image tag densities are displayed relative to the indicated genomic coordinates. The left image is an example of an affected site bound only by the wild-type version of the mutant remodeler. The right image displays binding by all three remodelers at the affected DHS site. Changes in hypersensitivity in the absence (+Tet) or presence (-Tet) of the indicated dominant-negative remodeler are overlayed and highlighted by arrows. (d-f) Overlay of remodeler binding at DHS sites in the presence or absence of the indicated dominant-negative remodeler protein. Displayed are scatter plots of DHS site tag density in the presence of each dominant-negative protein (y-axis) compared to DHS site tag density in the absence of this protein. Inset, original scatter plots of DNase I hypersensitivity from Fig. 5.
Supplementary Figure 8 Control DHS data analysis for remodeler binding at affected DHS sites.
(a) The scatter plot compares the maximum tag density of -Tet versus +Tet DHS hotspots in the control 3134 dataset. (b) The venn diagram shows the overlap between -Tet DHS hotspots and +Tet DHS hotspots in the control 3134 dataset. The area of each subset is proportional to the number of sites.
Supplementary Figure 9 Positional distribution for trend classes in chromatin access at DHS sites.
Venn diagrams present the occurrence frequencies for the 27 trend classes expected for dnBrg1, dnSnf2h and dnChd4 interactions at DHS sites. For each of these classes, the positional distribution is shown here for categories with more than 50 sites genome-wide. Notation; Brg1-UP indicates sites increased by more than 2-fold in -Tet (with dnBrg1 expression); Brg1-DN indicates means sites decreased by more than 2-fold in -Tet; NC indicates no change. The same notation is used for the dnSnf2h and dnChd4 classes. Sites are classified as promoter (< 2.5 kb upstream or downstream from the nearest TSS), exon (> 2.5 kb downstream from the nearest TSS , overlapping an exon), intron (> 2.5 kb downstream from the nearest TSS , in an intron not overlapping an exon), distal upstream (> 2.5 kb upstream from the nearest TSS), or downstream (> 2.5 kb downstream from the nearest TSS).
Supplementary Figure 10 Effects of dn-remodeler expression on gene expression.
RNA levels were examined by microarray analysis in cells either expressing (- tet), or not expressing (+tet) dnBrg1, dnSnf2h, and dnChd4. (a) Example 1; Gipr locus is strongly up-regulated by dnCHD4. In a parallel cell line with tet regulated w.t. Chd4, strong induction of the w.t. protein has no effect of Gipr exression. (b) DHS track for Gipr locus; pronounced DHS element immediately downstream of TSS (arrow) is activated by dnCHD4. (c) Example 2; Kcne1 locus is up-regulated by dnCHD4. Again, w.t. Chd4 has no effect. (d) DHS track for Kcne1 locus; DHS element downstream of TSS (arrow) is activated by dnCHD4. (e) In parallel cell lines, -tet activation produces 30-35 fold increases in dnChd4 and w.t. Chd4 RNA levels. (f) Global effects on gene expression. The number of genes either up- or down-regulated (fc > 1.2; p<0.05) is shown.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 and Supplementary Tables 1 and 2 (PDF 19860 kb)
Rights and permissions
About this article
Cite this article
Morris, S., Baek, S., Sung, MH. et al. Overlapping chromatin-remodeling systems collaborate genome wide at dynamic chromatin transitions. Nat Struct Mol Biol 21, 73–81 (2014). https://doi.org/10.1038/nsmb.2718
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2718
This article is cited by
-
Heterochromatin rewiring and domain disruption-mediated chromatin compaction during erythropoiesis
Nature Structural & Molecular Biology (2023)
-
Histone exchange sensors reveal variant specific dynamics in mouse embryonic stem cells
Nature Communications (2023)
-
ARID1A-dependent maintenance of H3.3 is required for repressive CHD4-ZMYND8 chromatin interactions at super-enhancers
BMC Biology (2022)
-
Chromatin-directed proteomics-identified network of endogenous androgen receptor in prostate cancer cells
Oncogene (2021)
-
Overarching control of autophagy and DNA damage response by CHD6 revealed by modeling a rare human pathology
Nature Communications (2021)