Abstract
DNA polymerase ɛ (Pol ɛ) is a high-fidelity polymerase that has been shown to participate in leading-strand synthesis during DNA replication in eukaryotic cells. We present here a ternary structure of the catalytic core of Pol ɛ (142 kDa) from Saccharomyces cerevisiae in complex with DNA and an incoming nucleotide. This structure provides information about the selection of the correct nucleotide and the positions of amino acids that might be critical for proofreading activity. Pol ɛ has the highest fidelity among B-family polymerases despite the absence of an extended β-hairpin loop that is required for high-fidelity replication by other B-family polymerases. Moreover, the catalytic core has a new domain that allows Pol ɛ to encircle the nascent double-stranded DNA. Altogether, the structure provides an explanation for the high processivity and high fidelity of leading-strand DNA synthesis in eukaryotes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Burgers, P.M. Polymerase dynamics at the eukaryotic DNA replication fork. J. Biol. Chem. 284, 4041–4045 (2009).
Miyabe, I., Kunkel, T.A. & Carr, A.M. The major roles of DNA polymerases epsilon and delta at the eukaryotic replication fork are evolutionarily conserved. PLoS Genet. 7, e1002407 (2011).
Nick McElhinny, S.A., Gordenin, D.A., Stith, C.M., Burgers, P.M. & Kunkel, T.A. Division of labor at the eukaryotic replication fork. Mol. Cell 30, 137–144 (2008).
Larrea, A.A. et al. Genome-wide model for the normal eukaryotic DNA replication fork. Proc. Natl. Acad. Sci. USA 107, 17674–17679 (2010).
Pursell, Z.F., Isoz, I., Lundstrom, E.B., Johansson, E. & Kunkel, T.A. Yeast DNA polymerase ɛ participates in leading-strand DNA replication. Science 317, 127–130 (2007).
Hamatake, R.K. et al. Purification and characterization of DNA polymerase II from the yeast Saccharomyces cerevisiae: identification of the catalytic core and a possible holoenzyme form of the enzyme. J. Biol. Chem. 265, 4072–4083 (1990).
Araki, H., Hamatake, R.K., Johnston, L.H. & Sugino, A. DPB2, the gene encoding DNA polymerase II subunit B, is required for chromosome replication in Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 88, 4601–4605 (1991).
Isoz, I., Persson, U., Volkov, K. & Johansson, E. The C-terminus of Dpb2 is required for interaction with Pol2 and for cell viability. Nucleic Acids Res. 40, 11545–11553 (2012).
Ohya, T., Maki, S., Kawasaki, Y. & Sugino, A. Structure and function of the fourth subunit (Dpb4p) of DNA polymerase ɛ in Saccharomyces cerevisiae. Nucleic Acids Res. 28, 3846–3852 (2000).
Araki, H. et al. Cloning DPB3, the gene encoding the third subunit of DNA polymerase II of Saccharomyces cerevisiae. Nucleic Acids Res. 19, 4867–4872 (1991).
Tsubota, T., Maki, S., Kubota, H., Sugino, A. & Maki, H. Double-stranded DNA binding properties of Saccharomyces cerevisiae DNA polymerase ɛ and of the Dpb3p-Dpb4p subassembly. Genes Cells 8, 873–888 (2003).
Aksenova, A. et al. Mismatch repair-independent increase in spontaneous mutagenesis in yeast lacking non-essential subunits of DNA polymerase ɛ. PLoS Genet. 6, e1001209 (2010).
Shcherbakova, P.V. et al. Unique error signature of the four-subunit yeast DNA polymerase ɛ. J. Biol. Chem. 278, 43770–43780 (2003).
Chilkova, O. et al. The eukaryotic leading and lagging strand DNA polymerases are loaded onto primer-ends via separate mechanisms but have comparable processivity in the presence of PCNA. Nucleic Acids Res. 35, 6588–6597 (2007).
Núñez-Ramírez, R. et al. Flexible tethering of primase and DNA Pol α in the eukaryotic primosome. Nucleic Acids Res. 39, 8187–8199 (2011).
Palles, C. et al. Germline mutations affecting the proofreading domains of POLE and POLD1 predispose to colorectal adenomas and carcinomas. Nat. Genet. 45, 136–144 (2013).
Kandoth, C. et al. Integrated genomic characterization of endometrial carcinoma. Nature 497, 67–73 (2013).
Church, D.N. et al. DNA polymerase ɛ and δ exonuclease domain mutations in endometrial cancer. Hum. Mol. Genet. 22, 2820–2828 (2013).
Johansson, E. & Macneill, S.A. The eukaryotic replicative DNA polymerases take shape. Trends Biochem. Sci. 35, 339–347 (2010).
Swan, M.K., Johnson, R.E., Prakash, L., Prakash, S. & Aggarwal, A.K. Structural basis of high-fidelity DNA synthesis by yeast DNA polymerase δ. Nat. Struct. Mol. Biol. 16, 979–986 (2009).
Perera, R.L. et al. Mechanism for priming DNA synthesis by yeast DNA Polymerase α. eLife 2, e00482 (2013).
Franklin, M.C., Wang, J. & Steitz, T.A. Structure of the replicating complex of a pol α family DNA polymerase. Cell 105, 657–667 (2001).
Nuutinen, T. et al. The solution structure of the amino-terminal domain of human DNA polymerase ɛ subunit B is homologous to C-domains of AAA+ proteins. Nucleic Acids Res. 36, 5102–5110 (2008).
Asturias, F.J. et al. Structure of Saccharomyces cerevisiae DNA polymerase ɛ by cryo-electron microscopy. Nat. Struct. Mol. Biol. 13, 35–43 (2006).
Hartlepp, K.F. et al. The histone fold subunits of Drosophila CHRAC facilitate nucleosome sliding through dynamic DNA interactions. Mol. Cell. Biol. 25, 9886–9896 (2005).
Liu, S. et al. Crystal structure of the herpes simplex virus 1 DNA polymerase. J. Biol. Chem. 281, 18193–18200 (2006).
Yang, W. Nucleases: diversity of structure, function and mechanism. Q. Rev. Biophys. 44, 1–93 (2011).
Morrison, A. & Sugino, A. The 3′→5′ exonucleases of both DNA polymerases δ and ɛ participate in correcting errors of DNA replication in Saccharomyces cerevisiae. Mol. Gen. Genet. 242, 289–296 (1994).
Savino, C. et al. Insights into DNA replication: the crystal structure of DNA polymerase B1 from the archaeon Sulfolobus solfataricus. Structure 12, 2001–2008 (2004).
Holm, L. & Rosenstrom, P. Dali server: conservation mapping in 3D. Nucleic Acids Res. 38, W545–W549 (2010).
Garcia-Diaz, M., Bebenek, K., Krahn, J.M., Kunkel, T.A. & Pedersen, L.C. A closed conformation for the Pol λ catalytic cycle. Nat. Struct. Mol. Biol. 12, 97–98 (2005).
Garcia-Diaz, M., Bebenek, K., Krahn, J.M., Pedersen, L.C. & Kunkel, T.A. Role of the catalytic metal during polymerization by DNA polymerase λ. DNA Repair (Amst.) 6, 1333–1340 (2007).
Fortune, J.M. et al. Saccharomyces cerevisiae DNA polymerase δ: high fidelity for base substitutions but lower fidelity for single- and multi-base deletions. J. Biol. Chem. 280, 29980–29987 (2005).
Xia, S., Christian, T.D., Wang, J. & Konigsberg, W.H. Probing minor groove hydrogen bonding interactions between RB69 DNA polymerase and DNA. Biochemistry 51, 4343–4353 (2012).
Wang, M. et al. Insights into base selectivity from the 1.8 A resolution structure of an RB69 DNA polymerase ternary complex. Biochemistry 50, 581–590 (2011).
Pata, J.D. Structural diversity of the Y-family DNA polymerases. Biochim. Biophys. Acta 1804, 1124–1135 (2010).
Stocki, S.A., Nonay, R.L. & Reha-Krantz, L.J. Dynamics of bacteriophage T4 DNA polymerase function: identification of amino acid residues that affect switching between polymerase and 3′ → 5′ exonuclease activities. J. Mol. Biol. 254, 15–28 (1995).
Hogg, M., Aller, P., Konigsberg, W., Wallace, S.S. & Doublie, S. Structural and biochemical investigation of the role in proofreading of a β hairpin loop found in the exonuclease domain of a replicative DNA polymerase of the B family. J. Biol. Chem. 282, 1432–1444 (2007).
Williams, L.N., Herr, A.J. & Preston, B.D. Emergence of DNA Polymerase ɛ antimutators that escape error-induced extinction in yeast. Genetics 193, 751–770 (2013).
Doublié, S., Tabor, S., Long, A.M., Richardson, C.C. & Ellenberger, T. Crystal structure of a bacteriophage T7 DNA replication complex at 2.2 A resolution. Nature 391, 251–258 (1998).
Ling, H., Boudsocq, F., Woodgate, R. & Yang, W. Crystal structure of a Y-family DNA polymerase in action: a mechanism for error-prone and lesion-bypass replication. Cell 107, 91–102 (2001).
Nakamura, T., Zhao, Y., Yamagata, Y., Hua, Y.J. & Yang, W. Watching DNA polymerase η make a phosphodiester bond. Nature 487, 196–201 (2012).
Sabouri, N. & Johansson, E. Translesion synthesis of abasic sites by yeast DNA polymerase ɛ. J. Biol. Chem. 284, 31555–31563 (2009).
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Crystallogr. 26, 795–800 (1993).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Terwilliger, T. SOLVE and RESOLVE: automated structure solution, density modification and model building. J. Synchrotron Radiat. 11, 49–52 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Chilkova, O., Jonsson, B.H. & Johansson, E. The quaternary structure of DNA polymerase ɛ from Saccharomyces cerevisiae. J. Biol. Chem. 278, 14082–14086 (2003).
Hogg, M., Sauer-Eriksson, A.E. & Johansson, E. Promiscuous DNA synthesis by human DNA polymerase θ. Nucleic Acids Res. 40, 2611–2622 (2012).
Potterton, L. et al. Developments in the CCP4 molecular-graphics project. Acta Crystallogr. D Biol. Crystallogr. 60, 2288–2294 (2004).
Acknowledgements
Data were collected at beamline ID23-2 of the European Synchrotron Radiation Facility. We thank U.H. Sauer (Umeå University) for help and advice with crystal handling and diffraction-data collection, S. Huang for help with refinement, P. Burgers (Washington University School of Medicine) for the kind gift of purified DNA polymerase δ, the Protein Production Platform of Umeå University for help with protein production and the High Performance Computing Center North (HPC2N) in the Swedish National Strategic e-Science Research Program eSSENCE for their technical support. This research was supported by the Swedish Research Council (E.J. and A.E.S.-E.), Swedish Cancer Society (E.J.), Kempe Foundations (E.J. and A.E.S.-E.), Knut and Alice Wallenberg Foundation (E.J.), Insamlingstiftelsen vid den medicinska fakulteten vid Umeå Universitet (E.J.), Karriärbidrag, Umeå University (E.J.) and Umeå Centre for Microbial Research (A.E.S.-E.).
Author information
Authors and Affiliations
Contributions
E.-B.L. overexpressed and purified the protein. M.H. and P.O. grew and handled the crystals. M.H., A.E.S.-E. and E.J. collected the diffraction data. A.E.S.-E. determined and refined the structure. P.O., G.O.B. and R.A.G. constructed the mutant proteins and performed the enzymatic assays. M.H., A.E.S.-E. and E.J. designed the experiments and prepared the manuscript. All authors discussed and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
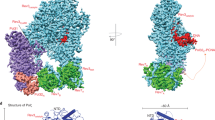
Supplementary Figure 1 Localization of insertions in the structure of Pol ɛ.
(a) The ribbon structure of Pol ɛ is colored in green with the insertions that are not present in Pol δ highlighted in gold. (b) To aid the reader, the same orientation of the Pol ɛ structure in (a) is shown but now colored according to its respective domains (see Figure 1 for color coding). (c) A close-up view of the N-terminal domains from four DNA polymerases (Pol ɛ, Pol δ (PDB entry 3iay), Herpes Simplex Virus-I (PDB entry 2gv9) and RB69 gp43 (PDB entry 1ig9)).
Supplementary Figure 2 Schematic diagram showing the interactions between Pol ɛ and the 11–16 primer template DNA.
Residues interacting with DNA are color coded according to the domains they belong to (see Figure 1 for color coding). Residues in brackets indicate that these residues have side chains positioned in the vicinity of the DNA with the potential to form direct salt-bridges to the phosphodiester backbone. The first iodo-cytosine in the template DNA strand is not modeled in the structure and is shaded in white. Three base-specific contacts are indicated: the Nζ atom of K967 makes one base-specific hydrogen bond with the O2 atom of iodo-uracil 10, and the Nη1 and Nη2 atoms of R988 make base-specific hydrogen bonds to the N3 atom of guanine 8 and the O2 atom of iodo-uracil 7, respectively. The three nucleotides mentioned are all located in the primer strand. Water-mediated interactions between amino acids and the template bases are indicated with a red line and shown in Figure 5. The main-chain amino group of Y645 forms a hydrogen bond to the 3'-hydroxyl group on the deoxyribose moiety of the dATP
Supplementary Figure 3 Composition of the polymerase active sites in Pol ɛ, Pol δ (PDB 3IAY) and RB69 gp43 (PDB 1IG9).
Metals and carbon atoms are shown in green, blue, and yellow in Pol ɛ, Pol δ, and RB69 gp43, respectively. Water molecules are shown as red spheres. A superimposition of the three active sites is also shown. The corresponding residues D877 in Pol ɛ, D764 in Pol δ, and D623 in RB69 gp43 are in almost identical positions in all three structures, but the orientations of the corresponding residues D640 in Pol ɛ, D411 in RB69 gp43, and D608 in Pol δ vary.
Supplementary Figure 4 Composition of the P domain and metal-binding sites
(a) Close-up view of the P-domain (dark blue), primer template dsDNA (orange) and dATP (red) in two orientations. (b) Close-up view of Mg2+ ion coordination in Pol ɛ. The Mg2+ ion, bound in site B, is octahedrally coordinated by D640, D877, the main chain carbonyl oxygen of V641, and three oxygen atoms from the triphosphate moiety of dATP. For clarity, the carbon atoms of ddC are shown in orange whereas other carbon atoms are shown in yellow. Some distances in Å, including metal- and hydrogen bonds, and the distance between the 3'-C of the primer-end and the α-phosphate, are shown as dashed lines. (c) The anomalous map showed electron density over the metal site of ~6 sigma over the background noise. The density is contoured at 4 sigma level in the figure.
Supplementary Figure 5 Distribution of cancer-associated mutations in Pol ɛ.
(a-d) Cancer-associated mutations in human POLE have previously been reported. We have indicated the corresponding amino acids in red in Pol ɛ.
Supplementary Figure 6 Antimutator mutations in Pol ɛ previously identified in a genetic screen.
The five amino acid changes isolated in that study are listed in the table and the targeted amino acids are shown in blue in the structure.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1–3 (PDF 8027 kb)
Rights and permissions
About this article
Cite this article
Hogg, M., Osterman, P., Bylund, G. et al. Structural basis for processive DNA synthesis by yeast DNA polymerase ɛ. Nat Struct Mol Biol 21, 49–55 (2014). https://doi.org/10.1038/nsmb.2712
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2712
This article is cited by
-
A mechanistic model of primer synthesis from catalytic structures of DNA polymerase α–primase
Nature Structural & Molecular Biology (2024)
-
Efficient discrimination against RNA-containing primers by human DNA polymerase ε
Scientific Reports (2022)
-
A conserved mechanism for regulating replisome disassembly in eukaryotes
Nature (2021)
-
Structural understanding of non-nucleoside inhibition in an elongating herpesvirus polymerase
Nature Communications (2021)
-
How asymmetric DNA replication achieves symmetrical fidelity
Nature Structural & Molecular Biology (2021)