Abstract
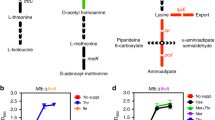
Several major pathogens, including Mycobacterium tuberculosis, parasitize host cells and exploit host-derived nutrients to sustain their own metabolism. Although the carbon sources that are used by M. tuberculosis have been extensively studied, the mechanisms by which mycobacteria capture and metabolize nitrogen, which is another essential constituent of biomolecules, have only recently been revisited. In this Progress article, we discuss central nitrogen metabolism in M. tuberculosis, the mechanisms that are used by this pathogen to obtain nitrogen from its host and the potential role of nitrogen capture and metabolism in virulence.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Zumla, A. George, A., Sharma, V., Herbert, N. & Baroness Masham of Ilton. WHO's 2013 global report on tuberculosis: successes, threats, and opportunities. Lancet 382, 1765–1767 (2013).
Ehrt, S. & Schnappinger, D. Mycobacterial survival strategies in the phagosome: defence against host stresses. Cell. Microbiol. 11, 1170–1178 (2009).
Honer zu Bentrup, K. & Russell, D. G. Mycobacterial persistence: adaptation to a changing environment. Trends Microbiol. 9, 597–605 (2001).
Rustad, T. R., Sherrid, A. M., Minch, K. J. & Sherman, D. R. Hypoxia: a window into Mycobacterium tuberculosis latency. Cell. Microbiol. 11, 1151–1159 (2009).
Russell, D. G. Mycobacterium tuberculosis: here today, and here tomorrow. Nature Rev. Mol. Cell Biol. 2, 569–577 (2001).
Zhang, Y. J. & Rubin, E. J. Feast or famine: the host–pathogen battle over amino acids. Cell. Microbiol. 15, 1079–1087 (2013).
Eisenreich, W., Dandekar, T., Heesemann, J. & Goebel, W. Carbon metabolism of intracellular bacterial pathogens and possible links to virulence. Nature Rev. Microbiol. 8, 401–412 (2010).
Leigh, J. A. & Dodsworth, J. A. Nitrogen regulation in bacteria and archaea. Annu. Rev. Microbiol. 61, 349–377 (2007).
Amon, J., Titgemeyer, F. & Burkovski, A. A genomic view on nitrogen metabolism and nitrogen control in mycobacteria. J. Mol. Microbiol. Biotechnol. 17, 20–29 (2009).
Viljoen, A. J., Kirsten, C. J., Baker, B., van Helden, P. D. & Wiid, I. J. The role of glutamine oxoglutarate aminotransferase and glutamate dehydrogenase in nitrogen metabolism in Mycobacterium bovis BCG. PLoS ONE 8, e84452 (2013).
Cole, S. T. et al. Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 393, 537–544 (1998).
Tullius, M. V., Harth, G. & Horwitz, M. A. Glutamine synthetase GlnA1 is essential for growth of Mycobacterium tuberculosis in human THP-1 macrophages and guinea pigs. Infect. Immun. 71, 3927–3936 (2003).
Amon, J., Titgemeyer, F. & Burkovski, A. Common patterns — unique features: nitrogen metabolism and regulation in Gram-positive bacteria. FEMS Microbiol. Rev. 34, 588–605 (2010).
Arcondeguy, T., Jack, R. & Merrick, M. P(II) signal transduction proteins, pivotal players in microbial nitrogen control. Microbiol. Mol. Biol. Rev. 65, 80–105 (2001).
Parish, T. & Stoker, N. G. glnE is an essential gene in Mycobacterium tuberculosis. J. Bacteriol. 182, 5715–5720 (2000).
Shetty, N. D., Reddy, M. C., Palaninathan, S. K., Owen, J. L. & Sacchettini, J. C. Crystal structures of the apo and ATP bound Mycobacterium tuberculosis nitrogen regulatory PII protein. Protein Sci. 19, 1513–1524 (2010).
Williams, K. J., Bennett, M. H., Barton, G. R., Jenkins, V. A. & Robertson, B. D. Adenylylation of mycobacterial Glnk (PII) protein is induced by nitrogen limitation. Tuberculosis (Edinb.) 93, 198–206 (2013).
Read, R., Pashley, C. A., Smith, D. & Parish, T. The role of GlnD in ammonia assimilation in Mycobacterium tuberculosis. Tuberculosis (Edinb.) 87, 384–390 (2007).
Mehta, R. et al. Adenylylation and catalytic properties of Mycobacterium tuberculosis glutamine synthetase expressed in Escherichia coli versus mycobacteria. J. Biol. Chem. 279, 22477–22482 (2004).
Nott, T. J. et al. An intramolecular switch regulates phosphoindependent FHA domain interactions in Mycobacterium tuberculosis. Sci Signal 2, ra12 (2009).
O'Hare, H. M. et al. Regulation of glutamate metabolism by protein kinases in mycobacteria. Mol. Microbiol. 70, 1408–1423 (2008).
Villarino, A. et al. Proteomic identification of M. tuberculosis protein kinase substrates: PknB recruits GarA, a FHA domain-containing protein, through activation loop-mediated interactions. J. Mol. Biol. 350, 953–963 (2005).
Malm, S. et al. The roles of the nitrate reductase NarGHJI, the nitrite reductase NirBD and the response regulator GlnR in nitrate assimilation of Mycobacterium tuberculosis. Microbiology 155, 1332–1339 (2009).
Jenkins, V. A., Barton, G. R., Robertson, B. D. & Williams, K. J. Genome wide analysis of the complete GlnR nitrogen-response regulon in Mycobacterium smegmatis. BMC Genomics 14, 301 (2013).
Jessberger, N. et al. Nitrogen starvation-induced transcriptome alterations and influence of transcription regulator mutants in Mycobacterium smegmatis. BMC Res. Notes 6, 482 (2013).
Gouzy, A., Poquet, Y. & Neyrolles, O. Amino acid capture and utilization within the Mycobacterium tuberculosis phagosome. Future Microbiol. 9, 631–637 (2014).
Lyon, R. H., Hall, W. H. & Costas-Martinez, C. Utilization of amino acids during growth of Mycobacterium tuberculosis in rotary cultures. Infect. Immun. 1, 513–520 (1970).
Niederweis, M. Nutrient acquisition by mycobacteria. Microbiology 154, 679–692 (2008).
Khan, A. & Sarkar, D. Nitrate reduction pathways in mycobacteria and their implications during latency. Microbiology 158, 301–307 (2012).
Giffin, M. M., Raab, R. W., Morganstern, M. & Sohaskey, C. D. Mutational analysis of the respiratory nitrate transporter NarK2 of Mycobacterium tuberculosis. PLoS ONE 7, e45459 (2012).
Lofthouse, E. K. et al. Systems-based approaches to probing metabolic variation within the Mycobacterium tuberculosis complex. PLoS ONE 8, e75913 (2013).
Gouzy, A. et al. Mycobacterium tuberculosis exploits asparagine to assimilate nitrogen and resist acid stress during infection. PLoS Pathog. 10, e1003928 (2014).
Surken, M., Keller, C., Rohker, C., Ehlers, S. & Bange, F. C. Anaerobic arginine metabolism of Mycobacterium tuberculosis is mediated by arginine deiminase (arcA), but is not essential for chronic persistence in an aerogenic mouse model of infection. Int. J. Med. Microbiol. 298, 657–661 (2008).
Song, H. & Niederweis, M. Uptake of sulphate but not phosphate by Mycobacterium tuberculosis is slower than that for Mycobacterium smegmatis. J. Bacteriol. 194, 956–964 (2012).
Gouzy, A. et al. Mycobacterium tuberculosis nitrogen assimilation and host colonization require aspartate. Nature Chem. Biol. 9, 674–676 (2013).
Seth, A. & Connell, N. D. Amino acid transport and metabolism in mycobacteria: cloning, interruption, and characterization of an L-Arginine/γ-aminobutyric acid permease in Mycobacterium bovis BCG. J. Bacteriol. 182, 919–927 (2000).
Price, C. T., Bukka, A., Cynamon, M. & Graham, J. E. Glycine betaine uptake by the ProXVWZ ABC transporter contributes to the ability of Mycobacterium tuberculosis to initiate growth in human macrophages. J. Bacteriol. 190, 3955–3961 (2008).
Flores-Valdez, M. A., Morris, R. P., Laval, F., Daffe, M. & Schoolnik, G. K. Mycobacterium tuberculosis modulates its cell surface via an oligopeptide permease (Opp) transport system. FASEB J. 23, 4091–4104 (2009).
Green, R. M., Seth, A. & Connell, N. D. A peptide permease mutant of Mycobacterium bovis BCG resistant to the toxic peptides glutathione and S-nitrosoglutathione. Infect. Immun. 68, 429–436 (2000).
Braibant, M., Gilot, P. & Content, J. The ATP binding cassette (ABC) transport systems of Mycobacterium tuberculosis. FEMS Microbiol. Rev. 24, 449–467 (2000).
Lin, W. et al. Urease activity represents an alternative pathway for Mycobacterium tuberculosis nitrogen metabolism. Infect. Immun. 80, 2771–2779 (2012).
Clemens, D. L., Lee, B. Y. & Horwitz, M. A. Purification, characterization, and genetic analysis of Mycobacterium tuberculosis urease, a potentially critical determinant of host–pathogen interaction. J. Bacteriol. 177, 5644–5652 (1995).
Nathan, C. & Shiloh, M. U. Reactive oxygen and nitrogen intermediates in the relationship between mammalian hosts and microbial pathogens. Proc. Natl Acad. Sci. USA 97, 8841–8848 (2000).
Davidge, K. S. & Dikshit, K. L. Haemoglobins of mycobacteria: structural features and biological functions. Adv. Microb. Physiol. 63, 147–194 (2013).
Cunningham-Bussel, A., Zhang, T. & Nathan, C. F. Nitrite produced by Mycobacterium tuberculosis in human macrophages in physiologic oxygen impacts bacterial ATP consumption and gene expression. Proc. Natl Acad. Sci. USA 110, E4256–E4265 (2013).
Jung, J. Y. et al. The intracellular environment of human macrophages that produce nitric oxide promotes growth of mycobacteria. Infect. Immun. 81, 3198–3209 (2013).
Davis, A. S. et al. Mechanism of inducible nitric oxide synthase exclusion from mycobacterial phagosomes. PLoS Pathog. 3, e186 (2007).
MacMicking, J. D. et al. Identification of nitric oxide synthase as a protective locus against tuberculosis. Proc. Natl Acad. Sci. USA 94, 5243–5248 (1997).
Aly, S. et al. Oxygen status of lung granulomas in Mycobacterium tuberculosis-infected mice. J. Pathol. 210, 298–305 (2006).
Shin, J. H. et al. 1H NMR-based metabolomic profiling in mice infected with Mycobacterium tuberculosis. J. Proteome Res. 10, 2238–2247 (2011).
Somashekar, B. S. et al. Metabolic profiling of lung granuloma in Mycobacterium tuberculosis infected guinea pigs: ex vivo1H magic angle spinning NMR studies. J. Proteome Res. 10, 4186–4195 (2011).
Beste, D. J. et al. 13C.flux spectral analysis of host-pathogen metabolism reveals a mixed diet for intracellular tuberculosis. Chem. Biol. 20, 1012–1021 (2013).
Cole, S. T. et al. Massive gene decay in the leprosy bacillus. Nature 409, 1007–1011 (2001).
Elharar, Y. et al. Survival of mycobacteria depends on proteasome-mediated amino acid recycling under nutrient limitation. EMBO J. 33, 1802–1814 (2014).
Gandotra, S., Lebron, M. B. & Ehrt, S. The Mycobacterium tuberculosis proteasome active site threonine is essential for persistence yet dispensable for replication and resistance to nitric oxide. PLoS Pathog. 6, e1001040 (2010).
Nau, G. J. et al. Human macrophage activation programs induced by bacterial pathogens. Proc. Natl Acad. Sci. USA 99, 1503–1508 (2002).
Tailleux, L. et al. Probing host pathogen crosstalk by transcriptional profiling of both Mycobacterium tuberculosis and infected human dendritic cells and macrophages. PLoS ONE 3, e1403 (2008).
Campbell-Valois, F. X. et al. Quantitative proteomics reveals that only a subset of the endoplasmic reticulum contributes to the phagosome. Mol Cell Proteomics 11, M111 016378 (2012).
Barel, M., Meibom, K., Dubail, I., Botella, J. & Charbit, A. Francisella tularensis regulates the expression of the amino acid transporter SLC1A5 in infected THP-1 human monocytes. Cell. Microbiol. 14, 1769–1783 (2012).
Wieland, H., Ullrich, S., Lang, F. & Neumeister, B. Intracellular multiplication of Legionella pneumophila depends on host cell amino acid transporter SLC1A5. Mol. Microbiol. 55, 1528–1537 (2005).
Das, P. et al. Cationic amino acid transporters and Salmonella Typhimurium ArgT collectively regulate arginine availability towards intracellular Salmonella growth. PLoS ONE 5, e15466 (2010).
Talaue, M. T. et al. Arginine homeostasis in J774.1 macrophages in the context of Mycobacterium bovis BCG infection. J. Bacteriol. 188, 4830–4840 (2006).
Qualls, J. E. et al. Sustained generation of nitric oxide and control of mycobacterial infection requires argininosuccinate synthase 1. Cell Host Microbe 12, 313–323 (2012).
Zhang, Y. J. et al. Tryptophan biosynthesis protects mycobacteria from CD4 T-cell-mediated killing. Cell 155, 1296–1308 (2013).
Chu, P., Rodriguez, A. R., Arulanandam, B. P. & Klose, K. E. Tryptophan prototrophy contributes to Francisella tularensis evasion of gamma interferon-mediated host defense. Infect. Immun. 79, 2356–2361 (2011).
van der Wel, N. et al. M. tuberculosis and M. leprae translocate from the phagolysosome to the cytosol in myeloid cells. Cell 129, 1287–1298 (2007).
Fuchs, T. M., Eisenreich, W., Kern, T. & Dandekar, T. Toward a systemic understanding of Listeria monocytogenes metabolism during infection. Front. Microbiol. 3, 23 (2012).
Steele, S. et al. Francisella tularensis harvests nutrients derived via ATG5-independent autophagy to support intracellular growth. PLoS Pathog. 9, e1003562 (2013).
Bradfute, S. B. et al. Autophagy as an immune effector against tuberculosis. Curr. Opin. Microbiol. 16, 355–365 (2013).
Carroll, P., Pashley, C. A. & Parish, T. Functional analysis of GlnE, an essential adenylyl transferase in Mycobacterium tuberculosis. J. Bacteriol. 190, 4894–4902 (2008).
Nilsson, M. T. et al. Structural basis for the inhibition of Mycobacterium tuberculosis glutamine synthetase by novel ATP-competitive inhibitors. J. Mol. Biol. 393, 504–513 (2009).
Youmans, A. S. & Youmans, G. P. Studies on the metabolism of Mycobacterium tuberculosis. II. The effect of compounds related to the Kreb's tricarboxylic acid cycle on the growth of Mycobacterium tuberculosis var. hominis. J. Bacteriol. 65, 96–99 (1953).
Pandey, A. K. & Sassetti, C. M. Mycobacterial persistence requires the utilization of host cholesterol. Proc. Natl Acad. Sci. USA 105, 4376–4380 (2008).
Daniel, J., Maamar, H., Deb, C., Sirakova, T. D. & Kolattukudy, P. E. Mycobacterium tuberculosis uses host triacylglycerol to accumulate lipid droplets and acquires a dormancy-like phenotype in lipid-loaded macrophages. PLoS Pathog. 7, e1002093 (2011).
Caire-Brandli, I. et al. Reversible lipid accumulation and associated division arrest of Mycobacterium avium in lipoprotein-induced foamy macrophages may resemble key events during latency and reactivation of tuberculosis. Infect. Immun. 82, 476–490 (2014).
Peyron, P. et al. Foamy macrophages from tuberculous patients' granulomas constitute a nutrient-rich reservoir for M. tuberculosis persistence. PLoS Pathog. 4, e1000204 (2008).
Russell, D. G., Cardona, P. J., Kim, M. J., Allain, S. & Altare, F. Foamy macrophages and the progression of the human tuberculosis granuloma. Nature Immunol. 10, 943–948 (2009).
Singh, V. et al. Mycobacterium tuberculosis-driven targeted recalibration of macrophage lipid homeostasis promotes the foamy phenotype. Cell Host Microbe 12, 669–681 (2012).
Marrero, J., Trujillo, C., Rhee, K. Y. & Ehrt, S. Glucose phosphorylation is required for Mycobacterium tuberculosis persistence in mice. PLoS Pathog. 9, e1003116 (2013).
Rhee, K. Y. et al. Central carbon metabolism in Mycobacterium tuberculosis: an unexpected frontier. Trends Microbiol. 19, 307–314 (2011).
Beste, D. J. et al. 13C metabolic flux analysis identifies an unusual route for pyruvate dissimilation in mycobacteria which requires isocitrate lyase and carbon dioxide fixation. PLoS Pathog. 7, e1002091 (2011).
Barel, M. & Charbit, A. Francisella tularensis intracellular survival: to eat or to die. Microbes Infect. (2013).
Gesbert, G. et al. Asparagine assimilation is critical for intracellular replication and dissemination of Francisella. Cell. Microbiol. (2013).
Sauer, J. D., Bachman, M. A. & Swanson, M. S. The phagosomal transporter A couples threonine acquisition to differentiation and replication of Legionella pneumophila in macrophages. Proc. Natl Acad. Sci. USA 102, 9924–9929 (2005).
Price, C. T., Al-Quadan, T., Santic, M., Rosenshine, I. & Abu Kwaik, Y. Host proteasomal degradation generates amino acids essential for intracellular bacterial growth. Science 334, 1553–1557 (2011).
Acknowledgements
A.G. is a fellow of the Fondation pour la Recherche Médicale (FRM). The authors received no specific funding for this work. Research in the laboratory of O.N. is supported by the Centre National de la Recherche Scientifique (CNRS), the FRM, the Agence Nationale de la Recherche (ANR), the European Union, the Fondation Mérieux, and the Bettencourt–Schueller Foundation. The funding agencies had no role in the decision to publish this article or in its preparation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Gouzy, A., Poquet, Y. & Neyrolles, O. Nitrogen metabolism in Mycobacterium tuberculosis physiology and virulence. Nat Rev Microbiol 12, 729–737 (2014). https://doi.org/10.1038/nrmicro3349
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro3349
This article is cited by
-
Urinary markers of Mycobacterium tuberculosis and dysbiosis in paediatric tuberculous meningitis cases undergoing treatment
Gut Pathogens (2024)
-
Volatilomes of human infection
Analytical and Bioanalytical Chemistry (2024)
-
Novel gene similar to nitrite reductase (NO forming) plays potentially important role in the latency of tuberculosis
Scientific Reports (2021)
-
Novel therapeutic evaluation biomarkers of lipid metabolism targets in uncomplicated pulmonary tuberculosis patients
Signal Transduction and Targeted Therapy (2021)
-
Identification of serum biomarkers for active pulmonary tuberculosis using a targeted metabolomics approach
Scientific Reports (2020)