Abstract
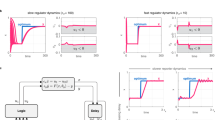
The integration of synthetic and cell-free biology has made tremendous strides towards creating artificial cellular nanosystems using concepts from solution-based chemistry, where only the concentrations of reacting species modulate gene expression rates. However, it is known that macromolecular crowding, a key feature in natural cells, can dramatically influence biochemical kinetics via volume exclusion effects, which reduce diffusion rates and enhance binding rates of macromolecules. Here, we demonstrate that macromolecular crowding can increase the robustness of gene expression by integrating synthetic cellular components of biological circuits and artificial cellular nanosystems. Furthermore, we reveal how ubiquitous cellular modules, including genetic components, a negative feedback loop and the size of the crowding molecules can fine-tune gene circuit response to molecular crowding. By bridging a key gap between artificial and living cells, our work has implications for efficient and robust control of both synthetic and natural cellular circuits.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Gibson, D. G. et al. Complete chemical synthesis, assembly, and cloning of a Mycoplasma genitalium genome. Science 319, 1215–1220 (2008).
Pinheiro, V. B. et al. Synthetic genetic polymers capable of heredity and evolution. Science 336, 341–344 (2012).
Nawroth, J. C. et al. A tissue-engineered jellyfish with biomimetic propulsion. Nature Biotechnol. 30, 792–797 (2012).
Kim, J. & Winfree, E. Synthetic in vitro transcriptional oscillators. Mol. Syst. Biol. 7, 465 (2011).
Fernandes, R., Roy, V., Wu, H. C. & Bentley, W. E. Engineered biological nanofactories trigger quorum sensing response in targeted bacteria. Nature Nanotech. 5, 213–217 (2010).
Murtas, G., Kuruma, Y., Bianchini, P., Diaspro, A. & Luisi, P. L. Protein synthesis in liposomes with a minimal set of enzymes. Biochem. Biophys. Res. Commun. 363, 12–17 (2007).
Schroeder, A. et al. Remotely activated protein-producing nanoparticles. Nano Lett. 12, 2685–2689 (2012).
Martino, C. et al. Protein expression, aggregation, and triggered release from polymersomes as artificial cell-like structures. Angew. Chem. Int. Ed. 51, 6416–6420 (2012).
Ishikawa, K., Sato, K., Shima, Y., Urabe, I. & Yomo, T. Expression of a cascading genetic network within liposomes. FEBS Lett. 576, 387–390 (2004).
Leduc, P. R. et al. Towards an in vivo biologically inspired nanofactory. Nature Nanotech. 2, 3–7 (2007).
Gardner, P. M., Winzer, K. & Davis, B. G. Sugar synthesis in a protocellular model leads to a cell signalling response in bacteria. Nature Chem. 1, 377–383 (2009).
Mansy, S. S. et al. Template-directed synthesis of a genetic polymer in a model protocell. Nature 454, 122–125 (2008).
Noireaux, V., Maeda, Y. T. & Libchaber, A. Development of an artificial cell, from self-organization to computation and self-reproduction. Proc. Natl Acad. Sci. USA 108, 3473–3480 (2011).
Chang, T. M. Therapeutic applications of polymeric artificial cells. Nature Rev. Drug Discov. 4, 221–235 (2005).
Jewett, M. C., Calhoun, K. A., Voloshin, A., Wuu, J. J. & Swartz, J. R. An integrated cell-free metabolic platform for protein production and synthetic biology. Mol. Syst. Biol. 4, 220 (2008).
Ellis, R. J. Macromolecular crowding: obvious but underappreciated. Trends Biochem. Sci. 26, 597–604 (2001).
Morelli, M. J., Allen, R. J. & Wolde, P. R. Effects of macromolecular crowding on genetic networks. Biophys. J. 101, 2882–2891 (2011).
Zimmerman, S. B. & Trach, S. O. Estimation of macromolecule concentrations and excluded volume effects for the cytoplasm of Escherichia coli. J. Mol. Biol. 222, 599–620 (1991).
Minton, A. P. The influence of macromolecular crowding and macromolecular confinement on biochemical reactions in physiological media. J. Biol. Chem. 276, 10577–10580 (2001).
Minton, A. P. The effect of volume occupancy upon the thermodynamic activity of proteins – some biochemical consequences. Mol. Cell. Biochem. 55, 119–140 (1983).
Li, G-W., Berg, O. G. & Elf, J. Effects of macromolecular crowding and DNA looping on gene regulation kinetics. Nature Phys. 5, 294–297 (2009).
Beg, Q. K. et al. Intracellular crowding defines the mode and sequence of substrate uptake by Escherichia coli and constrains its metabolic activity. Proc. Natl Acad. Sci. USA 104, 12663–12668 (2007).
Richter, K., Nessling, M. & Lichter, P. Experimental evidence for the influence of molecular crowding on nuclear architecture. J. Cell Sci. 120, 1673–1680 (2007).
Elcock, A. H. Models of macromolecular crowding effects and the need for quantitative comparisons with experiment. Curr. Opin. Struct. Biol. 20, 196–206 (2010).
Schoen, I., Krammer, H. & Braun, D. Hybridization kinetics is different inside cells. Proc. Natl Acad. Sci. USA 106, 21649–21654 (2009).
Phillip, Y., Sherman, E., Haran, G. & Schreiber, G. Common crowding agents have only a small effect on protein–protein interactions. Biophys. J. 97, 875–885 (2009).
Mika, J. T. & Poolman, B. Macromolecule diffusion and confinement in prokaryotic cells. Curr. Opin. Biotechnol. 22, 117–126 (2011).
Laurent, T. C. The interaction between polysaccharides and other macromolecules. 5. The solubility of proteins in the presence of dextran. Biochem. J. 89, 253–257 (1963).
Feder, T. J., Brust-Mascher, I., Slattery, J. P., Baird, B. & Webb, W. W. Constrained diffusion or immobile fraction on cell surfaces: a new interpretation. Biophys. J. 70, 2767–2773 (1996).
Friedman, L. J. & Gelles, J. Mechanism of transcription initiation at an activator-dependent promoter defined by single-molecule observation. Cell 148, 679–689 (2012).
Wang, Y., Guo, L., Golding, I., Cox, E. C. & Ong, N. P. Quantitative transcription factor binding kinetics at the single-molecule level. Biophys. J. 96, 609–620 (2009).
Bancaud, A. et al. Molecular crowding affects diffusion and binding of nuclear proteins in heterochromatin and reveals the fractal organization of chromatin. EMBO J. 28, 3785–3798 (2009).
Burg, M. B., Kwon, E. D. & Kultz, D. Osmotic regulation of gene expression. FASEB J. 10, 1598–1606 (1996).
Martin, C. T. & Coleman, J. E. Kinetic analysis of T7 RNA polymerase–promoter interactions with small synthetic promoters. Biochemistry 26, 2690–2696 (1987).
Gardner, T. S., Cantor, C. R. & Collins, J. J. Construction of a genetic toggle switch in Escherichia coli. Nature 403, 339–342 (2000).
Spitzer, J. & Poolman, B. The role of biomacromolecular crowding, ionic strength, and physicochemical gradients in the complexities of life's emergence. Microbiol. Mol. Biol. Rev. 73, 371–388 (2009).
Chen, H. Z. & Zubay, G. Prokaryotic coupled transcription-translation. Methods Enzymol. 101, 674–690 (1983).
Milo, R. et al. Network motifs: simple building blocks of complex networks. Science 298, 824–827 (2002).
Sunami, T. et al. Femtoliter compartment in liposomes for in vitro selection of proteins. Anal. Biochem. 357, 128–136 (2006).
Harris, D. C. & Jewett, M. C. Cell-free biology: exploiting the interface between synthetic biology and synthetic chemistry. Curr. Opin. Biotechnol. 23, 672–678 (2012).
Acknowledgements
The authors thank members of the LeDuc and Schwartz laboratories, the group of the Center for Mechanical Technology and Automation at University of Aveiro in Portugal, Dr. Shuqiang Huang, Dr. Gang Bao, and Dr. Lingchong You for discussions and comments, and R. Murphy, A. Mitchell, B. Armitage, T. Lee, F. Lanni and the Molecular Biosensor and Imaging Center for providing access to equipment. This work was partially supported by a Lane Postdoctoral Fellowship (C.T.), a Society in Science – Branco Weiss Fellowship (C.T.), NIH 1R01GM086237 (M.B. & S.S.), NIH 8U54GM103529 (M.B. & S.S.), NIH 1R01AI076318 (R.S), NIH 1R01CA140214 (R.S), NSF CMMI-1100430 (P.L.), NSF CMMI-0856187 (P.L.), NSF CMMI-1160840 (P.L.), ONR N000140910215 (P.L.), and NSF CPS-1135850 (P.L.).
Author information
Authors and Affiliations
Contributions
C.T., S.S., R.S. and P.L. conceived the research and designed the experiments. C.T. and S.S. performed the experiments. C.T. carried out the modelling analysis. C.T., S.S., M.B. and P.L. provided materials and reagents. C.T., R.S. and P.L. interpreted the results and wrote the manuscript, with critical input from S.S. and M.B. All authors approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 683 kb)
Supplementary movie S1
Supplementary movie S1 (AVI 1107 kb)
Supplementary movie S2
Supplementary movie S2 (AVI 413 kb)
Rights and permissions
About this article
Cite this article
Tan, C., Saurabh, S., Bruchez, M. et al. Molecular crowding shapes gene expression in synthetic cellular nanosystems. Nature Nanotech 8, 602–608 (2013). https://doi.org/10.1038/nnano.2013.132
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2013.132
This article is cited by
-
Negative autoregulation controls size scaling in confined gene expression reactions
Scientific Reports (2022)
-
pSpatiocyte: a high-performance simulator for intracellular reaction-diffusion systems
BMC Bioinformatics (2020)
-
Instantaneous fibrillation of egg white proteome with ionic liquid and macromolecular crowding
Communications Materials (2020)
-
New opportunities for creating man-made bioarchitectures utilizing microfluidics
Biomedical Microdevices (2019)
-
Spatial Stochastic Intracellular Kinetics: A Review of Modelling Approaches
Bulletin of Mathematical Biology (2019)