Abstract
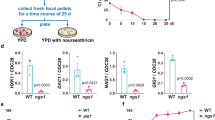
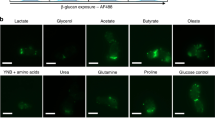
As they proliferate, fungi expose antigens at their cell surface that are potent stimulators of the innate immune response, and yet the commensal fungus Candida albicans is able to colonize immuno competent individuals. We show that C. albicans may evade immune detection by presenting a moving immunological target. We report that the exposure of β-glucan, a key pathogen-associated molecular pattern (PAMP) located at the cell surface of C. albicans and other pathogenic Candida species, is modulated in response to changes in the carbon source. Exposure to lactate induces β-glucan masking in C. albicans via a signalling pathway that has recruited an evolutionarily conserved receptor (Gpr1) and transcriptional factor (Crz1) from other well-characterized pathways. In response to lactate, these regulators control the expression of cell-wall-related genes that contribute to β-glucan masking. This represents the first description of active PAMP masking by a Candida species, a process that reduces the visibility of the fungus to the immune system.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
14 July 2017
In the PDF version of this article previously published, the year of publication provided in the footer of each page and in the 'How to cite' section was erroneously given as 2017, it should have been 2016. This error has now been corrected. The HTML version of the article was not affected.
References
Brown, G. D. & Gordon, S. Immune recognition. A new receptor for β-glucans. Nature 413, 36–37 (2001).
Hardison, S. E. & Brown, G. D. C-type lectin receptors orchestrate antifungal immunity. Nat. Immunol. 13, 817–822 (2012).
O'Meara, T. R. & Alspaugh, J. A. The Cryptococcus neoformans capsule: a sword and a shield. Clin. Microbiol. Rev. 25, 387–408 (2012).
Carrion, S. d. J. et al. The RodA hydrophobin on Aspergillus fumigatus spores masks dectin-1- and dectin-2-dependent responses and enhances fungal survival in vivo. J. Immunol. 191, 2581–2588 (2013).
Gow, N. A. R. & Hube, B. Importance of the Candida albicans cell wall during commensalism and infection. Curr. Opin. Microbiol. 15, 406–412 (2012).
Wheeler, R. T. & Fink, G. R. A drug-sensitive genetic network masks fungi from the immune system. PLoS Pathog. 2, e35 (2006).
Wheeler, R. T., Kombe, D., Agarwala, S. D. & Fink, G. R. Dynamic, morphotype-specific Candida albicans beta-glucan exposure during infection and drug treatment. PLoS Pathog. 4, e1000227 (2008).
Nett, J. E., Sanchez, H., Cain, M. T. & Andes, D. R. Genetic basis of Candida biofilm resistance due to drug-sequestering matrix glucan. J. Infect. Dis. 202, 171–175 (2010).
Gantner, B. N., Simmons, R. M. & Underhill, D. M. Dectin-1 mediates macrophage recognition of Candida albicans yeast but not filaments. EMBO J. 24, 1277–1286 (2005).
Hopke, A. et al. Neutrophil attack triggers extracellular trap-dependent Candida cell wall remodeling and altered immune recognition. PLoS Pathog. 12, e1005644 (2016).
Sosinska, G. J. et al. Hypoxic conditions and iron restriction affect the cell-wall proteome of Candida albicans grown under vagina-simulative conditions. Microbiology 154, 510–520 (2008).
Ene, I. V. et al. Host carbon sources modulate cell wall architecture, drug resistance and virulence in a fungal pathogen. Cell. Microbiol. 14, 1319–1335 (2012).
Munro, C. A. et al. The PKC, HOG and Ca2+ signalling pathways co-ordinately regulate chitin synthesis in Candida albicans. Mol. Microbiol. 63, 1399–1413 (2007).
Heilmann, C. J. et al. Surface stress induces a conserved cell wall stress response in the pathogenic fungus Candida albicans. Eukaryot. Cell 12, 254–264 (2013).
Galán-Díez, M. et al. Candida albicans beta-glucan exposure is controlled by the fungal CEK1-mediated mitogen-activated protein kinase pathway that modulates immune responses triggered through dectin-1. Infect. Immun. 78, 1426–1436 (2010).
Ene, I. V., Cheng, S.-C., Netea, M. G. & Brown, A. J. P. Growth of Candida albicans cells on the physiologically relevant carbon source lactate affects their recognition and phagocytosis by immune cells. Infect. Immun. 81, 238–248 (2013).
Barelle, C. J. et al. Niche-specific regulation of central metabolic pathways in a fungal pathogen. Cell. Microbiol. 8, 961–971 (2006).
Owen, D. H. & Katz, D. F. A vaginal fluid simulant. Contraception 59, 91–95 (1999).
Ene, I. V. et al. Cell wall remodeling enzymes modulate fungal cell wall elasticity and osmotic stress resistance. mBio 6, e00986 (2015).
Marakalala, M. J. et al. Differential adaptation of Candida albicans in vivo modulates immune recognition by dectin-1. PLoS Pathog. 9, e1003315 (2013).
Vieira, N. et al. Functional specialization and differential regulation of short-chain carboxylic acid transporters in the pathogen Candida albicans. Mol. Microbiol. 75, 1337–1354 (2010).
Kuei, C. et al. Study of GPR81, the lactate receptor, from distant species identifies residues and motifs critical for GPR81 functions. Mol. Microbiol. 80, 848–858 (2011).
Maidan, M. M. et al. The G protein-coupled receptor Gpr1 and the Gα protein Gpa2 act through the cAMP-protein kinase A pathway to induce morphogenesis in Candida albicans. Mol. Biol. Cell 16, 1971–1986 (2005).
Lemaire, K., Van de Velde, S., Van Dijck, P. & Thevelein, J. M. Glucose and sucrose act as agonist and mannose as antagonist ligands of the G protein-coupled receptor Gpr1 in the yeast Saccharomyces cerevisiae. Mol. Cell 16, 293–299 (2004).
Johnston, C. A. & Siderovski, D. P. Receptor-mediated activation of heterotrimeric G-proteins: current structural insights. Mol. Pharmacol. 72, 219–230. (2007).
Leberer, E. et al. Virulence and hyphal formation of Candida albicans require the Ste20p-like protein kinase CaCla4p. Curr. Biol. 7, 539–546 (1997).
Santos, M. & de Larrinoa, I. F. Functional characterization of the Candida albicans CRZ1 gene encoding a calcineurin-regulated transcription factor. Curr. Genet. 48, 88–100 (2005).
Karababa, M. et al. CRZ1, a target of the calcineurin pathway in Candida albicans. Mol. Microbiol. 59, 1429–1451 (2006).
Kelly, M. T. et al. The Candida albicans CaACE2 gene affects morphogenesis, adherence and virulence. Mol. Microbiol. 53, 969–983 (2004).
Desai, P. R., van Wijlick, L., Kurtz, D., Juchimiuk, M. & Ernst, J. F. Hypoxia and temperature regulated morphogenesis in Candida albicans. PLoS Genet. 11, e1005447 (2015).
Hall, R. A. et al. The Mnn2 mannosyltransferase family modulates mannoprotein fibril length, immune recognition and virulence of Candida albicans. PLoS Pathog. 9, e1003276 (2013).
Bain, J. M. et al. Candida albicans hypha formation and mannan masking of β-glucan inhibit macrophage phagosome maturation. mBio 5, e01874-14 (2014).
Flint, H. J., Scott, K. P., Louis, P. & Duncan, S. H. The role of the gut microbiota in nutrition and health. Nat. Rev. Gastroenterol. Hepatol. 9, 577–589 (2012).
Jacobs, I. Blood lactate. Implications for training and sports performance. Sports Med. 3, 10–25 (1986).
Netea, M. G. et al. Immune sensing of Candida albicans requires cooperative recognition of mannans and glucans by lectin and Toll-like receptors. J. Clin. Invest. 116, 1642–1650 (2006).
Ferwerda, B. et al. Human dectin-1 deficiency and mucocutaneous fungal infections. N. Engl. J. Med. 361, 1760–1767 (2009).
Iliev, I. D. et al. Interactions between commensal fungi and the C-type lectin receptor dectin-1 influence colitis. Science 336, 1314–1317 (2012).
Wheeler, R. T., Kupiec, M., Magnelli, P., Abeijon, C. & Fink, G. R. A Saccharomyces cerevisiae mutant with increased virulence. Proc. Natl Acad. Sci. USA 100, 2766–2770 (2003).
Chen, Y. L. et al. Convergent evolution of calcineurin pathway roles in thermotolerance and virulence in Candida glabrata. G3 2, 675–691 (2012).
Lev, S., Desmarini, D., Chayakulkeeree, M., Sorrell, T. C. & Djordjevic, J. T. The Crz1/Sp1 transcription factor of Cryptococcus neoformans is activated by calcineurin and regulates cell wall integrity. PLoS ONE 7, e51403 (2012).
Ene, I. V. et al. Carbon source-induced reprogramming of the cell wall proteome and secretome modulates the adherence and drug resistance of the fungal pathogen Candida albicans. Proteomics 12, 3164–3179 (2012).
Gravelat, F. N. et al. Aspergillus galactosaminogalactan mediates adherence to host constituents and conceals hyphal β-glucan from the immune system. PLoS Pathog. 9, e1003575 (2013).
Rappleye, C. A. & Goldman, W. E. Fungal stealth technology. Trends Immunol. 29, 18–24 (2008).
Noble, S. M. & Johnson, A. D. Strains and strategies for large-scale gene deletion studies of the diploid human fungal pathogen Candida albicans. Eukaryot. Cell 4, 298–309 (2005).
Lewis, L. E. et al. Stage specific assessment of Candida albicans phagocytosis by macrophages identifies cell wall composition and morphogenesis as key determinants. PLoS Pathog. 8, e1002578 (2012).
Netea, M. G. et al. Increased susceptibility of TNF-α lymphotoxin-α double knockout mice to systemic candidiasis through impaired recruitment of neutrophils and phagocytosis of Candida albicans. J. Immunol. 163, 1498–1505 (1999).
Graham, L. M. et al. Soluble dectin-1 as a tool to detect β-glucans. J. Immunol. Methods 314, 164–169 (2006).
Shahana, S. et al. New Clox Systems for rapid and efficient gene disruption in Candida albicans. PLoS ONE 9, e100390 (2014).
Inglis, D. O . et al. The Candida Genome Database incorporates multiple Candida species: multispecies search and analysis tools with curated gene and protein information for Candida albicans and Candida glabrata. Nucleic Acids Res. 40, D667–D674 (2012).
Bindea, G. et al. ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25, 1091–1093 (2009).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Acknowledgements
The authors apologize to colleagues whose work we have been unable to cite because of space constraints. The authors thank F. Odds for access to his collection of clinical isolates. The authors thank R. Yuecel and L. Duncan in the Iain Fraser Cytometry Centre (Aberdeen University), and L. Wight and K. MacKenzie in the Microscopy and Histology Core Facility (Aberdeen University) for their expert help with the cytometry and microscopy experiments. The authors acknowledge the assistance of staff at the University of Aberdeen Medical Research Facility, and B. Martin in the Centre for Genome Enabled Biology and Medicine (Aberdeen University) for help with the RNA sequencing. A.J.P.B. was supported by the European Research Council (STRIFE, ERC-2009-AdG-249793), the UK Medical Research Council (MR/M026663/1), the UK Biotechnology and Biological Research Council (BB/K017365/1) and the Wellcome Trust (080088; 097377). E.R.B. was supported by the UK Biotechnology and Biological Research Council (BB/M014525/1), G.M.A. by CNPq-Brazil (Science without Borders fellowship 202976/2014–9), G.D.B. by the Wellcome Trust (102705), C.A.M. by the UK Medical Research Council (G0400284), D.M.M. by the UK National Centre for the Replacement, Refinement and Reduction of Animals in Research (NC/K000306/1), N.A.R.G. and J.W. by the Wellcome Trust (086827, 075470 and 101873) and Wellcome Trust Strategic Award in Medical Mycology and Fungal Immunology (097377). This work was also supported by the MRC Centre for Medical Mycology and the University of Aberdeen (MR/N006364/1). All animal experiments were performed under UK Home Office project licence PPL 60/4135 (granted to D.M.M.) in accordance with Home Office ethical guidelines.
Author information
Authors and Affiliations
Contributions
E.R.B., D.S.C., G.M.A. and A.J.P.B. planned the experiments. E.R.B., G.M.A., D.S.C., J.M., J.M.B., J.W., M.D.P. and D.M.M. performed the experiments. Animal experiments were performed by D.M.M., E.R.B. and S.E.H. S.L.K. and E.R.B performed bioinformatic analyses. S.E.H., L.A.W., L.P.E., C.A.M., N.A.R.G. and G.D.B. provided materials and insight during this study. E.R.B. and A.J.P.B. drafted the manuscript with contributions from J.M., G.M.A., D.S.C., J.M.B., C.A.M., N.A.R.G., G.D.B. and D.M.M.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary Figures 1–9 and Supplementary References (PDF 18644 kb)
Supplementary Tables
Supplementary Tables 1–4 (XLSX 74 kb)
Rights and permissions
About this article
Cite this article
Ballou, E., Avelar, G., Childers, D. et al. Lactate signalling regulates fungal β-glucan masking and immune evasion. Nat Microbiol 2, 16238 (2017). https://doi.org/10.1038/nmicrobiol.2016.238
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2016.238
This article is cited by
-
Immunogenicity and efficacy of CNA25 as a potential whole-cell vaccine against systemic candidiasis
EMBO Molecular Medicine (2024)
-
The nematode-trapping fungus Arthrobotrys oligospora detects prey pheromones via G protein-coupled receptors
Nature Microbiology (2024)
-
Reappraisal of intra-abdominal candidiasis: insights from peritoneal fluid analysis
Intensive Care Medicine Experimental (2023)
-
Architecture of the dynamic fungal cell wall
Nature Reviews Microbiology (2023)
-
Microglia are not protective against cryptococcal meningitis
Nature Communications (2023)