Abstract
Chemical genetic analysis of protein kinases involves engineering kinases to be uniquely sensitive to inhibitors and ATP analogs that are not recognized by wild-type kinases. Despite the successful application of this approach to over two dozen kinases, several kinases do not tolerate the necessary modification to the ATP binding pocket, as they lose catalytic activity or cellular function upon mutation of the 'gatekeeper' residue that governs inhibitor and nucleotide substrate specificity. Here we describe the identification of second-site suppressor mutations to rescue the activity of 'intolerant' kinases. A bacterial genetic selection for second-site suppressors using an aminoglycoside kinase APH(3′)-IIIa revealed several suppressor hotspots in the kinase domain. Informed by results from this selection, we focused on the β sheet in the N-terminal subdomain and generated a structure-based sequence alignment of protein kinases in this region. From this alignment, we identified second-site suppressors for several divergent kinases including Cdc5, MEKK1, GRK2 and Pto. The ability to identify second-site suppressors to rescue the activity of intolerant kinases should facilitate chemical genetic analysis of the majority of protein kinases in the genome.
Similar content being viewed by others
Main
The large number of protein kinases in the 'kinome' and their ubiquitous involvement in central signaling pathways in eukaryotic cells has made discovery of inhibitors and substrates of these enzymes an area of intense research. The vast majority of protein kinase inhibitors lack specificity, owing to the highly conserved nature of the ATP binding site within the protein family1,2. Researchers in our laboratory have developed a chemical genetic approach that circumvents the specificity problem associated with conventional small-molecule inhibitors of protein kinases3,4. The approach exploits a conserved, large hydrophobic residue in the kinase active site (termed the gatekeeper5), which makes direct contact with the N6 group of ATP. When this residue is mutated from the naturally occurring bulky residue (methionine, leucine, phenylalanine or threonine, among others) to glycine or alanine, a pocket not found in any wild-type kinase is created within the kinase active site. The engineered kinase, termed an analog-sensitive (as) allele, can be potently and uniquely targeted by inhibitors that contain substituents that occupy this enlarged ATP binding pocket3. Likewise, this enlarged ATP binding pocket allows such mutant kinases to use unnatural N6-modified ATP analogs that are not accepted by any wild-type kinase, which can be used to label and thereby identify direct substrates of the kinase6.
In principle, the predominance of large hydrophobic residues at the gatekeeper position in the protein kinase superfamily should allow application of this chemical genetic approach to any kinase in the kinome for inhibitor and substrate discovery (Fig. 1a). Indeed, we and others have successfully applied this approach to both serine/threonine and tyrosine kinases from diverse families and organisms4,7,8,9,10,11,12,13,14,15,16,17,18,19,20 (Fig. 1b). The acute pharmacological perturbation permits the study of kinases involved in highly dynamic cellular processes, such as actin cytoskeleton reorganization, for which conventional genetic approaches cannot be used7,8. In addition, this chemical genetic approach frequently reveals functions of the kinase not previously exposed by conventional genetic studies4,7,8,9,10,11,12,13,14,15. These studies suggest that the chemical genetic analysis of kinases complements conventional genetics in terms of understanding both the dynamics of kinase signaling and the multiple roles of individual kinases. Thus, it is becoming increasingly important to develop an analog-sensitive allele of each kinase in the genome as a complement to conventional genetic studies of kinase function.
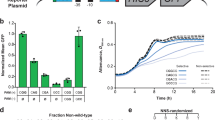
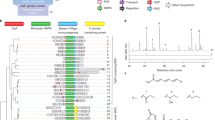
(a) Amino acid distribution at the gatekeeper position in the protein kinase superfamily. Each amino acid is represented by its one letter code on the x-axis. − indicates that a gap is found at this position in the alignment of the sequences of these kinases preventing definite assignment of an amino acid to the gatekeeper position. (b) Classification of kinases according to their tolerance of glycine mutation at the gatekeeper position. Upon mutation of the gatekeeper residue to glycine, kinases that maintain substantial catalytic activity or cellular function are considered tolerant, whereas those that severely lose activity are considered intolerant. * designates kinases discussed in this work.
Despite the broad applicability, a considerable number of kinases, roughly 30% of those tested to date, do not tolerate the mutation of the gatekeeper residue to glycine or alanine and are thus not amenable to study using this approach in its present embodiment (Fig. 1b). These intolerant kinases have a severe loss in catalytic activity and/or cellular function upon introduction of the space-creating gatekeeper mutation. Several such kinases have important roles in diverse cell signaling pathways, such as stress-activated MAPK signaling (MEKK1), mitotic regulation and cytokinesis (Cdc5), G protein–coupled receptor signaling (GRK2) and plant disease resistance (Pto). We describe here a general method to rescue the activity of these intolerant kinases, allowing extension of our chemical genetic methodology to a larger portion of the kinome.
To solve the problem of severe loss in activity caused by introduction of the space-creating gatekeeper mutation in a general manner, we used a stepwise approach to search for second-site suppressor mutations that could rescue the activity of kinases intolerant of a glycine gatekeeper mutation, termed suppressors of glycine gatekeeper (sogg). First, we performed a genetic selection with APH(3′)-IIIa, which does not tolerate mutation of its gatekeeper residue (Met90) to glycine, to identify 'hotspots' of sogg mutations within the kinase domain. This information allowed us to focus on the antiparallel β sheet in the kinase N-terminal subdomain and to generate a structure-based sequence alignment of protein kinases within this region. From this alignment, we identified sogg mutations in four widely divergent intolerant kinases that suppressed the loss of activity caused by the single glycine or alanine gatekeeper mutation. These results provide a basis for the extension of the chemical genetic approach to kinases that do not tolerate the single space-creating gatekeeper mutation.
Results
A genetic selection for sogg in an aminoglycoside kinase
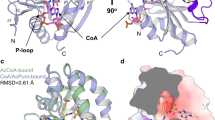
Aminoglycoside kinases confer antibiotic resistance by direct phosphorylation of antibacterial aminoglycosides, thus preventing their binding to the ribosome21. We selected an aminoglycoside kinase, APH(3′)-IIIa22, as the basis for a genetic selection for sogg mutations, because it produces a readily selectable phenotype (cell viability). Despite a low level of amino-acid sequence identity, APH(3′)-IIIa has strong structural homology to protein kinases23. The two families of kinases share similar ATP binding pockets and secondary structure elements (Fig. 2a). Mutation of Met90, the gatekeeper residue in APH(3′)-IIIa, to glycine resulted in an APH(3′)-IIIa allele with greatly reduced resistance to kanamycin when introduced into Escherichia coli (minimum inhibitory concentration (MIC) = 50 μg/ml for the M90G mutant, MIC = 300 μg/ml for the wild type), suggesting that the M90G mutation severely compromised the kinase activity of APH(3′)-IIIa (Fig. 2b). Based on their similar protein structures, enzymatic reactions and defects caused by the homologous gatekeeper mutation, we conclude that APH(3′)-IIIa could serve as a genetically selectable model for intolerant protein kinases.
(a) Ribbon diagram of APH(3′)-IIIa (maroon) in comparison with that of PKA (cyan). The gatekeeper residue, N-terminal lobe and C-terminal lobe in the kinase domain are indicated. (b) Growth curves of E. coli expressing wild-type APH(3′)-IIIa (green) or APH(3′)-IIIaM90G (red) cultured in the presence of different concentrations of kanamycin. OD600 of the cultures was measured after 12 h of incubation at 37 °C. The typical concentration used for kanamycin-resistance selection (100 μg/ml) is indicated with a dotted line in the graph.
We generated a random mutant library of the gene encoding APH(3′)-IIIaM90G and selected for transformants in E. coli with elevated resistance to aminoglycosides. Error-prone PCR was performed to introduce random mutations into the gene encoding APH(3′)-IIIaM90G. The resulting mutant library was first selected for resistant clones to 100 μg/ml of kanamycin, a concentration at which bacteria expressing the parent clone APH(3′)-IIIaM90G (MIC = 50 μg/ml) cannot survive. We next counter-selected the kanamycin resistant colonies against a second, structurally distinct aminoglycoside, neomycin, to eliminate kanamycin-specific mutations from the selection. The clones that survived both the kanamycin and neomycin selections were isolated and sequenced.
Out of a library of approximately 10,000 transformants, 18 unique mutations were identified at 14 positions in APH(3′)-IIIaM90G (Fig. 3a). These mutations can be broadly divided into three categories: mutations at the gatekeeper position, mutations in the C-terminal lobe of APH(3′)-IIIa, and mutations in the N-terminal lobe (Fig. 3a). Identification of rescue mutations at the gatekeeper position (G90V and G90E) validated our selection, as such mutations are expected to increase the activity of the APH gene by reverting Gly90 to larger, more wild type–like residues. The sogg mutations in the C-terminal and N-terminal lobes are of greater interest, because they recover the activity of APH(3′)-IIIa while preserving the space-creating mutation M90G.
(a) Suppressor mutations divided into three categories and listed according to their positions in the primary sequence of APH(3′)-IIIa. (b) Suppressor mutations mapped to the crystal structure of APH(3′)-IIIa. Mutated residues are highlighted and annotated in the APH(3′)-IIIa structure. The gatekeeper is colored red, residues in the N-terminal lobe, blue, and those in the C-terminal lobe, yellow. The structures of the two aminoglycosides used in the selection experiment, kanamycin and neomycin, are shown on the right with their respective primary phosphorylation site marked with a red arrow.
Most sogg mutations in the C-terminal lobe are located near the aminoglycoside-binding pocket (Fig. 3b). Three of these (E101A, E103A and E103V) result in substitution of surface acidic residues with neutral ones. In the crystal structure of APH(3′)-IIIa, a large number of acidic residues were found near the substrate-binding cleft24. These negatively charged residues are in close proximity to one another and may be important for binding positively charged aminoglycosides, yet they may be detrimental to overall protein stability25.
The sogg mutations in the N-terminal lobe are located within the five-stranded antiparallel β sheet. E24K and E24V mutated residues are located near the aminoglycoside binding pocket and may stabilize APH(3′)-IIIaM90G by eliminating surface acidic residues in a manner analogous to the three mutations described above. Another sogg mutation, Y17H, introduces a histidine within range to form a hydrogen bond with the carbonyl oxygen from the preceding α1 helix, based on the crystal structure of APH(3′)-IIIa (Fig. 3b). But this α1 helix is not conserved between aminoglycoside kinases and protein kinases, making it difficult to extend this particular mutation to protein kinases. The sogg mutation nearest to the gatekeeper in both primary sequence and three-dimensional structure is N87T (Fig. 3b). Notably, studies of β sheet–containing proteins indicate that β-branched amino acids (valine, threonine and isoleucine) are generally more favored in β sheets than non–β-branched amino acids26,27. Thus, it seems possible that the functional basis of this sogg mutation is to stabilize the antiparallel β sheet, thereby countering the deleterious effect of the glycine gatekeeper mutation on the same β sheet. Based on this analysis, we consider the β sheet of the N-terminal lobe to be the more favorable region for sogg identification.
Structure-based sequence alignment of kinases
Ideally, identification of sogg mutations should be based primarily on sequence analysis, which is both general and efficient (requiring a minimum number of candidate mutations to be screened). Given that the APH(3′)-IIIa selection revealed that the β sheet of the N-terminal lobe is a hotspot of sogg mutations, we generated a structure-based sequence alignment of both tolerant and intolerant protein kinases within this region. The two edge strands (β1 and β4) are largely exposed and frequently involved in protein-protein interactions, so they were excluded from the alignment. Twelve residues in the remaining three central β strands were included in the alignment (Fig. 4a). We examined these positions for any patterns that could distinguish intolerant kinases from tolerant kinases and tested for rescue of kinase activity (Fig. 4b).
(a) Ribbon diagram of the antiparallel β sheet in the c-Src N-terminal lobe. The residues selected in the alignment are highlighted in different colors according to their side chain orientations. (b) A structure-based sequence alignment of eight substitution-tolerant kinases and five substitution-intolerant kinases at selected positions in the three central β strands (β2, β3 and β5). Color codes for columns follow those in a. Residues that were mutated as potential suppressors in this study are yellow and bold.
Identification of sogg mutations for Cdc5 and MEKK1
Cdc5 is the sole budding yeast member of the polo-like family of kinases (PLK), a family of serine/threonine protein kinases conserved from yeast to human. Cdc5 is essential for cell viability. It functions in mitotic entry, adaptation to the DNA damage checkpoint, spindle checkpoint, mitotic exit and cytokinesis28. Despite extensive evidence underscoring the functional importance of PLK, relatively little is known about the downstream effectors of PLK in their regulation of these diverse cell cycle events. This motivated us to initiate a chemical genetic study of Cdc5.
We assessed the ability of two gatekeeper-modified Cdc5 mutants, cdc5-as1 (L158G) and cdc5-as2 (L158A), to functionally complement the temperature sensitive cdc5-1 allele using a colony-forming efficiency assay. Because cdc5-1 is nonfunctional at restrictive temperatures, viability of the yeast is dependent on the activity of plasmid-borne CDC5 alleles. Unfortunately, cdc5-1 strains transformed with plasmid bearing the genes encoding cdc5-as1 or cdc5-as2 had substantially reduced viability compared with wild-type CDC5 (Fig. 5a). Structure-based sequence alignment revealed that Cdc5 contains cysteine (Cys96) rather than the most commonly occurring valine at the particular position (Fig. 4b), and we introduced the second-site mutation C96V into the kinase and examined its effect on the cellular function of Cdc5 using the colony-forming efficiency assay. Notably, this second-site mutation rescued the in vivo function of both cdc5-as1 or cdc5-as2 close to wild-type levels (Fig. 5a). Moreover, C96V alone did not seem to affect the viability of yeast cells (Supplementary Fig. 1 online), suggesting that the C96V mutation did not cause drastic perturbation to the structure and biological function of Cdc5.
(a) Viability of yeast strains carrying different CDC5 alleles based on colony-forming efficiency. Second-site mutation C96V rescued the cellular function of Cdc5-as1 and Cdc5-as2. (b) Enzymatic activity of MEKK1 proteins revealed in an in vitro kinase assay. A catalytically inactive mutant of SEK (SEK-KR) was used as the substrate in the kinase assay. Second-site mutation C1238V rescued the catalytic activity of MEKK1-as1. (c) Morphine-induced μOR internalization in the presence of different GRK2 alleles (normalized to the wild-type activity). Second-site mutations S268T & S268V increased the cellular function of GRK2-as1 whereas the S268Y mutation had the opposite effect. Data represent the mean ± s.d. of three independent experiments each performed in triplicate. (d) Enzymatic activity of Pto proteins revealed in an in vitro kinase assay. Pti1 was used as the substrate in the kinase assay. One second-site mutation, L68I, substantially rescued the catalytic activity of Pto-as1 whereas the other mutation, L112I, had little effect on activity.
Another intolerant kinase, MEKK1, also contains cysteine (Cys1238) at the homologous position to Cys96 in Cdc5. As we had demonstrated that C96V successfully recovered the cellular function of Cdc5, we went on to test whether the homologous mutation could rescue the activity of MEKK1. As a MAP3K, MEKK1 is involved in many cellular functions including proliferation, differentiation and programmed cell death29. MEKK1 is specifically activated in response to stress signals including TNF-α, microtubule disruption and osmotic shock30,31. In an effort to screen for direct substrates of MEKK1 using N6-substituted ATP analogs, we mutated its gatekeeper residue (Ile1304) to glycine. The resulting MEKK1-as1 had poor enzymatic activity (<4% relative to wild-type MEKK1) in an in vitro kinase assay using catalytically inactive SAPK/ERK kinase (SEK-KR) as substrate (Fig. 5b). C1238V, the homologous mutation to C96V in Cdc5, was introduced to MEKK1-as1 and the catalytic activity of the resulting double mutant was tested in vitro. This second-site mutation increased the kinase activity of MEKK1-as1 by over 15-fold (60% relative to wild-type MEKK1) to nearly wild-type level of activity (Fig. 5b). When introduced into wild-type MEKK1 alone, C1238V caused a slight increase in kinase activity (108% relative to wild-type MEKK1).
The second-site suppressors identified for Cdc5 and MEKK1 involve introduction of a β-branched residue (valine) into the β sheet, which on the structural level is analogous to the β-branched sogg mutation identified for APH(3′)-IIIaM90G (N87T). The recurrence of β-branched sogg mutations led us to propose that introduction of β-branched residues in this β sheet could be a viable route to additional sogg mutations for other intolerant kinases. To test this hypothesis we used two additional intolerant protein kinases.
Identification of sogg mutations for GRK2
G protein–coupled receptor kinases (GRKs) phosphorylate the agonist-activated form of G protein–coupled receptors (GPCRs)32, which stimulates binding of the cytoplasmic accessory protein, arrestin, to the GPCR to initiate internalization and desensitization33,34. We applied the chemical genetic approach to GRK2 with the aim of specifically modulating the functional activity of GRK2 and investigating the role of GRK2 in mediating the morphine-induced internalization of the μ-opioid receptor (μOR).
Mutation of the gatekeeper (Leu271) to glycine in GRK2 afforded GRK2-as1. Functional comparison of GRK2-as1 with wild-type GRK2 was made in human embryonic kidney (HEK) 293 cells stably transfected with μOR. Only upon overexpression of functional GRK2, does morphine become capable of promoting rapid phosphorylation and endocytosis of μOR in HEK 293 cells35,36. This GRK2-dependent μOR internalization provides a quantitative readout of the ability of GRK2 to phosphorylate μOR and consequently induce internalization in a ligand-dependent manner. GRK2-as1 had approximately fivefold lower levels of receptor internalization in response to morphine treatment than wild-type GRK2 in this assay (Fig. 5c).
The structure-based sequence alignment within the β-sheet of the N-terminal sub-domain led us to focus on residue Ser268 of GRK2, which corresponds to Asn87 in APH(3′)-IIIa (Fig. 4b). Inspired by the homologous sogg mutation for APH(3′)-IIIaM90G, N87T, we mutated Ser268 to two different β-branched residues, threonine or valine, as potential suppressors for GRK2-as1. In addition, Ser268 was changed to tyrosine, the most common residue occurring at this position in tolerant kinases (Fig. 4b). Whereas the S268T mutation resulted in partial recovery of cellular function compared to GRK2-as1, the S268V mutation completely restored the activity of GRK2-as1 to wild-type levels as shown in the morphine-induced μOR internalization assay (Fig. 5c). In contrast, the S268Y mutation decreased the activity of GRK2-as1 to a basal level.
Identification of a second-site suppressor for Pto
Pto is a cytoplasmic serine/threonine kinase in tomato plants, which confers resistance to Pseudomonas syringae pv. tomato, the causative agent of bacterial speck disease37. To identify new direct targets of Pto and to dissect phosphorylation cascades originating from the Pto, we mutated the gatekeeper residue (Tyr114) in Pto to glycine or alanine to afford Pto-as1 and Pto-as2 respectively. We expressed glutathione-S-transferase (GST)-Pto fusion proteins in E. coli, and measured the catalytic activity of GST-Pto based on autophosphorylation and transphosphorylation of a substrate (Pti) in a gel-based autoradiography assay. In this assay, both Pto-as1 and Pto-as2 mutants had a severe loss of in vitro phosphorylation activity; the more active Pto-as2 had only 3% activity compared to wild-type Pto (Fig. 5d and data not shown).
We chose to focus on the more active Pto-as2, which contains an alanine gatekeeper. In an effort to identify suppressors of an alanine gatekeeper (soag) for Pto-as2, the sequence alignment within the β sheet of the N-terminal subdomain was again analyzed. Pto contains a leucine at position 68, immediately N-terminally to the catalytically essential lysine residue (Fig. 4b), whereas the β-branched leucine isomer, isoleucine, occurs with the highest frequency at this position among the tolerant kinases. This observation motivated us to test L68I as a suppressor for Pto-as2. In addition to L68I, a mutation at a different position (L112I), which is located two residues N-terminal to the gatekeeper in β5, was also introduced into Pto-as2 as a potential suppressor (Fig. 4b). The two second-site mutations had very different effects on Pto-as2 kinase activity; the L68I mutation increased the enzymatic activity of Pto-as2 by more than tenfold (33% activity, normalized to wild-type Pto),whereas the L112I mutation had little effect (Fig. 5d). Furthermore, the L68I mutation had little effect on the kinase activity when introduced into wild-type Pto alone (Supplementary Fig. 2 online).
Discussion
We undertook a two-step approach combining genetic selection and structure-based sequence analysis to identify second-site suppressors for kinases that do not tolerate the single space-creating gatekeeper mutation. In the first step, an aminoglycoside kinase was used as a model of intolerant protein kinases in a genetic selection for random suppressor mutations, the result of which points to the antiparallel β sheet in the kinase N-terminal lobe as a hotspot for sogg mutations. This five-stranded antiparallel β sheet contains numerous residues important for ATP binding and catalysis (for example, the catalytically essential lysine residue), and thus it is not surprising that a mutation likely destabilizing to the β sheet such as the glycine gatekeeper substitution could diminish the catalytic activity of a kinase. In retrospect, it is rather logical that second-site mutations within the same β sheet can correct the defect caused by the space-creating gatekeeper mutation.
The recognition of this β sheet as a suitable location for sogg mutations allowed us to focus on a subregion of the kinase domain and perform intensive sequence analysis within this small region, which greatly accelerated the identification of sogg mutations for intolerant protein kinases. Aided by a structure-based sequence alignment of kinases within the crucial β sheet, we first succeeded at identifying suppressor mutations for Cdc5 and MEKK1 by substituting an uncommon cysteine with the consensus valine at a particular position. The suppressor mutations of Cdc5 and MEKK1 both introduced β-branched residues like N87T, a sogg identified for APH(3′)-IIIa, consistent with a model that they function by restabilizing the β sheet in the kinase N-terminal lobe. This model is supported by the relative suppressor activity of the three second-site mutations in GRK2, in which case the two β-branched mutations (S268V and S268T) enhanced the cellular function of GRK2-as1 whereas the mutation to the consensus residue (S268Y) abolished GRK2 function. The recently solved crystal structure of GRK2 revealed that the side chain of Phe261 extends from the neighboring β4 strand to contact Ser268 (ref. 38), suggesting that the steric clash introduced by the S268Y mutation with Phe261 may be responsible for the complete disruption of GRK2 function. This result demonstrates that because of the varied local environment, mutation to the 'consensus' residue may not recover kinase activity.
To minimize perturbation to the structures and functions of the kinases, we deliberately designed relatively conservative suppressor mutations (such as C→V, S→T and L→I) at largely buried locations (the three central β strands) in the kinase. Available evidence supports the notion that these sogg mutations do not substantially alter the cellular functions of these kinases. This was highlighted in the case of Cdc5; whereas it was able to rescue the cellular function of the two cdc5-as alleles, the suppressor C96V appears not to appreciably perturb the natural function of Cdc5, indicated by the similar viability of yeast containing cdc5-C96V and CDC5 (Supplementary Fig. 1).
In addition to constituting minimal perturbation to the natural function of the kinase, a second-site suppressor mutation should not negatively affect the ability of the engineered kinase to accept unnatural ATP analogs or inhibitors. As none of the identified suppressor mutations are immediately adjacent to the gatekeeper pocket in space, it seems unlikely these suppressor mutations could decrease the size of the engineered pocket in the studied kinases. In the case of MEKK1, the second-site mutation C1238V actually enhanced the ability of MEKK1-as1 to use an unnatural ATP analog, N6-phenethyl-ATP, suggesting the faithful preservation of the gatekeeper pocket in this double mutant (Supplementary Fig. 3 online). The present study focused on the rescue of the catalytic activity (using ATP) or cellular function of gatekeeper-modified kinases. Characterization of the acceptance of inhibitors or ATP analogs with these doubly engineered kinases are now underway.
Despite the considerably improved kinase activity, the double mutants of MEKK1 and Pto had diminished activity compared to the wild-type kinase. Previous experience with analog-sensitive kinases suggests that cells tolerate moderate loss in activity of engineered kinases. For example, despite a 50-fold decrease in kcat/KM in vitro, CDC28-as1 is capable of replacing endogenous CDC28 in the budding yeast; the cell cycle of the resulting cdc28-as1 strain was not substantially impaired4. Thus it is likely that the activity of the double mutants of MEKK1 and Pto will be sufficient to support the cellular function of these kinases in vivo.
The method that we have developed to discover suppressor mutations for engineering functional analog-sensitive kinases is general and efficient because it is primarily based on kinase sequence information. It should extend our chemical genetic approach to a substantial number of protein kinases that do not tolerate the single gatekeeper mutation and bring us closer to our ultimate goal of performing chemical genetic analysis of every kinase in the kinome.
Methods
Site-directed mutagenesis.
We performed site-directed mutagenesis using the QuikChange system (Stratagene) except in the case of MEKK1, for which overlap extension PCR was used. We confirmed the introduced mutations by sequencing. The information on the mutagenic primers is available in Supplementary Table 1 online.
Random mutagenesis and suppressor selection of APH(3′)-IIIaM90G.
We subcloned APH(3′)-IIIa from the APH(3′)-IIIa/pET22b plasmid22, a gift from G.D. Wright (McMaster University, Ontario, Canada), into the BamHI and EcoRI sites in ProEX-HTb (Gibco BRL). APH(3′)-IIIaM90G was randomly mutagenized using error-prone PCR according to the previously described procedures39 with minor modifications. Mutagenic PCR mixtures were composed of 10 mM Tris-HCl (pH 8.3), 50 mM KCl, 7 mM MgCl2, 0.5 mM MnCl2, 0.2 mM dATP, 0.2 mM dGTP, 1.0 mM dCTP, 1.0 mM dTTP, 50 ng of plasmid DNA, 0.2 μM primer of each primer and 5.0 U of Taq DNA polymerase (Promega) in a final volume of 200 μl. DNA amplification was performed for 30 cycles, each consisting of 30-s denaturation at 95 °C, 1-min primer annealing at 54 °C, and 2-min primer extension at 68 °C. The PCR products were digested with BamHI and EcoRI and ligated into the ProEX-HTb vector. We transformed the ligation products into DH5α E. coli and plated the transformants on LB-agar plates containing 100 μg/ml kanamycin. We then picked the colonies that survived on the kanamycin plate and cultured them. Plasmid DNA was purified from the bacteria and retransformed into DH5α E. coli, fresh transformants were tested for viability on neomycin (100 μg/ml) plates. Plasmid DNA from colonies that consistently survived both kanamycin and neomycin selection was sequenced covering the entire coding region of APH(3′)-IIIa.
Colony-forming efficiency assay of Cdc5.
Yeast strain JC34 bearing the cdc5-1 temperature sensitive allele40 and the plasmid containing wild-type CDC5 was obtained from D.O. Morgan (University of California San Francisco, California, USA). JC34 yeast was transformed with the plasmids containing CDC5 alleles by standard methods and propagated in synthetic dropout medium minus leucine (SD-Leu) to saturation. The cultures were equalized for cell density between strains, tenfold serially diluted and spotted onto a yeast-peptone-dextrose (YPD) plate. The plates were incubated at 37 °C for 2 d, and images was taken on an Alpha Innotech Imager. A second plate was spotted and incubated at 23 °C for 4 d to confirm that identical colony size and number was seen in the absence of selection.
Expression and in vitro kinase assay of MEKK1.
The plasmid (pTM1) containing MEKK1 with an N-terminal EE (MHEEEEYMPMEGPG) epitope tag and a C-terminal chitin binding domain tag was previously described41. The fusion protein of MEKK1 were expressed in CV-1 cells using the vaccinia virus T7 polymerase–driven expression system42. MEKK1 was purified on chitin beads and assayed for activity as previously described41.
Receptor internalization assay of GRK2.
A C-terminally hemagglutinin-tagged version was constructed by subcloning the gene encoding GRK2 into pcDNA3.1 and inserting a hemagglutinin tag (YPYDVPDYA) at its C terminus by standard site-directed mutagenesis. The N-terminally Flag-tagged mouse μOR-1 has been described previously43. HEK 293 cells were transfected with the plasmid containing the gene encoding GRK2 or its mutant alleles along with μOR-1 using Lipofectamine 2000 (Invitrogen) and used for flow cytometry 48 h thereafter. Internalization of μOR-1 in the transfected cells was determined by measuring the relative amount of Flag-tagged receptors present in the plasma membrane after surface labeling with Alexa488-conjugated M1 antibody as described previously44. We performed fluorescence flow cytometry using a FACScan instrument (BD Biosciences). We collected 20,000 cells for each sample, and analyzed triplicate samples in each experiment.
Expression and in vitro kinase assay of Pto.
The pGEX-4T1 plasmid containing the gene encoding Pto fused to GST was described previously45. GST-Pto and its substrate GST-Pti1 (K96N, a catalytically inactive mutant)46 were expressed in DH12S E. coli and affinity-purified using glutathione-agarose beads. We performed kinase assays to test phosphorylation of GST-Pti1(K96N) by GST-Pto as previously described45.
Note: Supplementary information is available on the Nature Methods website.
References
Davies, S.P., Reddy, H., Caivano, M. & Cohen, P. Specificity and mechanism of action of some commonly used protein kinase inhibitors. Biochem. J. 351, 95–105 (2000).
Fabian, M.A. et al. A small molecule-kinase interaction map for clinical kinase inhibitors. Nat. Biotechnol. 23, 329–336 (2005).
Bishop, A.C. et al. Generation of monospecific nanomolar tyrosine kinase inhibitors via a chemical genetic approach. J. Am. Chem. Soc. 121, 627–631 (1999).
Bishop, A.C. et al. A chemical switch for inhibitor-sensitive alleles of any protein kinase. Nature 407, 395–401 (2000).
Adams, J., Huang, P. & Patrick, D. A strategy for the design of multiplex inhibitors for kinase-mediated signalling in angiogenesis. Curr. Opin. Chem. Biol. 6, 486–492 (2002).
Ubersax, J.A. et al. Targets of the cyclin-dependent kinase Cdk1. Nature 425, 859–864 (2003).
Sekiya-Kawasaki, M. et al. Dynamic phosphoregulation of the cortical actin cytoskeleton and endocytic machinery revealed by real-time chemical genetic analysis. J. Cell Biol. 162, 765–772 (2003).
Wang, H. et al. Inducible protein knockout reveals temporal requirement of CaMKII reactivation for memory consolidation in the brain. Proc. Natl. Acad. Sci. USA 100, 4287–4292 (2003).
Weiss, E.L., Bishop, A.C., Shokat, K.M. & Drubin, D.G. Chemical genetic analysis of the budding-yeast p21-activated kinase Cla4p. Nat. Cell Biol. 2, 677–685 (2000).
Weiss, E.L. et al. The Saccharomyces cerevisiae Mob2p-Cbk1p kinase complex promotes polarized growth and acts with the mitotic exit network to facilitate daughter cell–specific localization of Ace2p transcription factor. J. Cell Biol. 158, 885–900 (2002).
Carroll, A.S., Bishop, A.C., DeRisi, J.L., Shokat, K.M. & O'Shea, E.K. Chemical inhibition of the Pho85 cyclin-dependent kinase reveals a role in the environmental stress response. Proc. Natl. Acad. Sci. USA 98, 12578–12583 (2001).
Benjamin, K.R., Zhang, C., Shokat, K.M. & Herskowitz, I. Control of landmark events in meiosis by the CDK Cdc28 and the meiosis-specific kinase Ime2. Genes Dev. 17, 1524–1539 (2003).
Sreenivasan, A., Bishop, A.C., Shokat, K.M. & Kellogg, D.R. Specific inhibition of Elm1 kinase activity reveals functions required for early G1 events. Mol. Cell. Biol. 23, 6327–6337 (2003).
Wan, L., de los Santos, T., Zhang, C., Shokat, K. & Hollingsworth, N.M. Mek1 kinase activity functions downstream of RED1 in the regulation of meiotic double strand break repair in budding yeast. Mol. Biol. Cell 15, 11–23 (2004).
Liu, Y. et al. Two cyclin-dependent kinases promote RNA polymerase II transcription and formation of the scaffold complex. Mol. Cell. Biol. 24, 1721–1735 (2004).
Niswender, C.M. et al. Protein engineering of protein kinase A catalytic subunits results in the acquisition of novel inhibitor sensitivity. J. Biol. Chem. 277, 28916–28922 (2002).
Fan, Q.W., Zhang, C., Shokat, K.M. & Weiss, W.A. Chemical genetic blockade of transformation reveals dependence on aberrant oncogenic signaling. Curr. Biol. 12, 1386–1394 (2002).
Denzel, A. et al. Cutting edge: a chemical genetic system for the analysis of kinases regulating T cell development. J. Immunol. 171, 519–523 (2003).
Habelhah, H. et al. Identification of new JNK substrate using ATP pocket mutant JNK and a corresponding ATP analogue. J. Biol. Chem. 276, 18090–18095 (2001).
Eblen, S.T. et al. Identification of novel ERK2 substrates through use of an engineered kinase and ATP analogs. J. Biol. Chem. 278, 14926–14935 (2003).
Wright, G.D. & Thompson, P.R. Aminoglycoside phosphotransferases: proteins, structure, and mechanism. Front. Biosci. 4, D9–21 (1999).
McKay, G.A., Thompson, P.R. & Wright, G.D. Broad spectrum aminoglycoside phosphotransferase type III from Enterococcus: overexpression, purification, and substrate specificity. Biochemistry 33, 6936–6944 (1994).
Hon, W.C. et al. Structure of an enzyme required for aminoglycoside antibiotic resistance reveals homology to eukaryotic protein kinases. Cell 89, 887–895 (1997).
Fong, D.H. & Berghuis, A.M. Substrate promiscuity of an aminoglycoside antibiotic resistance enzyme via target mimicry. EMBO J. 21, 2323–2331 (2002).
Grimsley, G.R. et al. Increasing protein stability by altering long-range coulombic interactions. Protein Sci. 8, 1843–1849 (1999).
Otzen, D.E. & Fersht, A.R. Side-chain determinants of β-sheet stability. Biochemistry 34, 5718–5724 (1995).
Minor, D.L., Jr. & Kim, P.S. Measurement of the β sheet–forming propensities of amino acids. Nature 367, 660–663 (1994).
Lee, K.S., Park, J.E., Asano, S. & Park, C.J. Yeast polo-like kinases: functionally conserved multitask mitotic regulators. Oncogene 24, 217–229 (2005).
Schlesinger, T.K., Fanger, G.R., Yujiri, T. & Johnson, G.L. The TAO of MEKK. Front. Biosci. 3, D1181–D1186 (1998).
Nemoto, S., DiDonato, J.A. & Lin, A. Coordinate regulation of IκB kinases by mitogen-activated protein kinase kinase kinase 1 and NF-κB–inducing kinase. Mol. Cell. Biol. 18, 7336–7343 (1998).
Xia, Y. et al. MEK kinase 1 is critically required for c-Jun N-terminal kinase activation by proinflammatory stimuli and growth factor–induced cell migration. Proc. Natl. Acad. Sci. USA 97, 5243–5248 (2000).
Zhang, J. et al. Molecular mechanisms of G protein–coupled receptor signaling: role of G protein–coupled receptor kinases and arrestins in receptor desensitization and resensitization. Receptors Channels 5, 193–199 (1997).
Goodman, O.B., Jr. et al. β-arrestin acts as a clathrin adaptor in endocytosis of the β2-adrenergic receptor. Nature 383, 447–450 (1996).
Ferguson, S.S. et al. Role of beta-arrestin in mediating agonist-promoted G protein–coupled receptor internalization. Science 271, 363–366 (1996).
Keith, D.E. et al. Morphine activates opioid receptors without causing their rapid internalization. J. Biol. Chem. 271, 19021–19024 (1996).
Zhang, J. et al. Role for G protein–coupled receptor kinase in agonist-specific regulation of mu-opioid receptor responsiveness. Proc. Natl. Acad. Sci. USA 95, 7157–7162 (1998).
Pedley, K.F. & Martin, G.B. Molecular basis of Pto-mediated resistance to bacterial speck disease in tomato. Annu. Rev. Phytopathol. 41, 215–243 (2003).
Lodowski, D.T., Pitcher, J.A., Capel, W.D., Lefkowitz, R.J. & Tesmer, J.J. Keeping G proteins at bay: a complex between G protein–coupled receptor kinase 2 and Gβγ. Science 300, 1256–1262 (2003).
Miyazaki, C. et al. Changes in the specificity of antibodies by site-specific mutagenesis followed by random mutagenesis. Protein Eng. 12, 407–415 (1999).
Jaspersen, S.L., Charles, J.F., Tinker-Kulberg, R.L. & Morgan, D.O. A late mitotic regulatory network controlling cyclin destruction in Saccharomyces cerevisiae. Mol. Biol. Cell 9, 2803–2817 (1998).
Cross, J.V. & Templeton, D.J. Oxidative stress inhibits MEKK1 by site-specific glutathionylation in the ATP-binding domain. Biochem. J. 381, 675–683 (2004).
Fuerst, T.R., Niles, E.G., Studier, F.W. & Moss, B. Eukaryotic transient-expression system based on recombinant vaccinia virus that synthesizes bacteriophage T7 RNA polymerase. Proc. Natl. Acad. Sci. USA 83, 8122–8126 (1986).
Tsao, P.I. & von Zastrow, M. Type-specific sorting of G protein–coupled receptors after endocytosis. J. Biol. Chem. 275, 11130–11140 (2000).
Gage, R.M., Kim, K.A., Cao, T.T. & von Zastrow, M. A transplantable sorting signal that is sufficient to mediate rapid recycling of G protein–coupled receptors. J. Biol. Chem. 276, 44712–44720 (2001).
Sessa, G., D'Ascenzo, M., Loh, Y.T. & Martin, G.B. Biochemical properties of two protein kinases involved in disease resistance signaling in tomato. J. Biol. Chem. 273, 15860–15865 (1998).
Zhou, J., Loh, Y.T., Bressan, R.A. & Martin, G.B. The tomato gene Pti1 encodes a serine/threonine kinase that is phosphorylated by Pto and is involved in the hypersensitive response. Cell 83, 925–935 (1995).
Acknowledgements
We thank G.D. Wright for providing the APH(3′)-IIIa expression plasmid. We also thank M. Simon, Z. Knight and S. DiMagno for critical reading of the manuscript. This work was supported by the National Institutes of Health (R01EB001987 & AI44009 to K.M.S.) and Binational Science Foundation (Grant 2001124 to G.S.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Viability of yeast strains carrying different Cdc5 alleles based on colony-forming efficiency. (PDF 248 kb)
Supplementary Fig. 2
Enzymatic activity of Pto proteins revealed in an in vitro autophosphorylation kinase assay. (PDF 127 kb)
Supplementary Fig. 3
Catalytic activity of MEKK1 proteins using N6-phenethyl-ATP revealed in an in vitro kinase assay. (PDF 423 kb)
Supplementary Table 1
Mutagenic primers used in the paper. (PDF 85 kb)
Rights and permissions
About this article
Cite this article
Zhang, C., Kenski, D., Paulson, J. et al. A second-site suppressor strategy for chemical genetic analysis of diverse protein kinases. Nat Methods 2, 435–441 (2005). https://doi.org/10.1038/nmeth764
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth764
This article is cited by
-
Crystal structure of the human PRPK–TPRKB complex
Communications Biology (2021)
-
ASKA technology-based pull-down method reveals a suppressive effect of ASK1 on the inflammatory NOD-RIPK2 pathway in brown adipocytes
Scientific Reports (2021)
-
A chemical genetics approach to examine the functions of AAA proteins
Nature Structural & Molecular Biology (2021)
-
Chemical genetics strategy to profile kinase target engagement reveals role of FES in neutrophil phagocytosis
Nature Communications (2020)
-
Cdk9 regulates a promoter-proximal checkpoint to modulate RNA polymerase II elongation rate in fission yeast
Nature Communications (2018)