Abstract
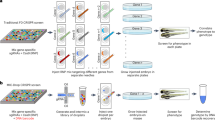
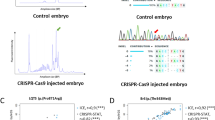
Identifying genes involved in biological processes is critical for understanding the molecular building blocks of life. We used engineered CRISPR (clustered regularly interspaced short palindromic repeats) to efficiently mutate specific loci in zebrafish (Danio rerio) and screen for genes involved in vertebrate biological processes. We found that increasing CRISPR efficiency by injecting optimized amounts of Cas9-encoding mRNA and multiplexing single guide RNAs (sgRNAs) allowed for phenocopy of known mutants across many phenotypes in embryos. We performed a proof-of-concept screen in which we used intersecting, multiplexed pool injections to examine 48 loci and identified two new genes involved in electrical-synapse formation. By deep sequencing target loci, we found that 90% of the genes were effectively screened. We conclude that CRISPR can be used as a powerful reverse genetic screening strategy in vivo in a vertebrate system.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
References
Chang, N. et al. Genome editing with RNA-guided Cas9 nuclease in zebrafish embryos. Cell Res. 23, 465–472 (2013).
Hwang, W.Y. et al. Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat. Biotechnol. 31, 227–229 (2013).
Hsu, P.D., Lander, E.S. & Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Peng, Y. et al. Making designer mutants in model organisms. Development 141, 4042–4054 (2014).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Jao, L.-E., Wente, S.R. & Chen, W. Efficient multiplex biallelic zebrafish genome editing using a CRISPR nuclease system. Proc. Natl. Acad. Sci. USA 110, 13904–13909 (2013).
Sung, Y.H. et al. Highly efficient gene knockout in mice and zebrafish with RNA-guided endonucleases. Genome Res. 24, 125–131 (2014).
Rohrschneider, M.R., Elsen, G.E. & Prince, V.E. Zebrafish Hoxb1a regulates multiple downstream genes including prickle1b. Dev. Biol. 309, 358–372 (2007).
Jessen, J.R. et al. Zebrafish trilobite identifies new roles for Strabismus in gastrulation and neuronal movements. Nat. Cell Biol. 4, 610–615 (2002).
Satou, C. et al. Functional role of a specialized class of spinal commissural inhibitory neurons during fast escapes in zebrafish. J. Neurosci. 29, 6780–6793 (2009).
Pereda, A.E. Electrical synapses and their functional interactions with chemical synapses. Nat. Rev. Neurosci. 15, 250–263 (2014).
Rash, J.E. et al. Molecular and functional asymmetry at a vertebrate electrical synapse. Neuron 79, 957–969 (2013).
Wang, T., Wei, J.J., Sabatini, D.M. & Lander, E.S. Genetic screens in human cells using the CRISPR-Cas9 system. Science 343, 80–84 (2014).
Craig, A.M. & Kang, Y. Neurexin-neuroligin signaling in synapse development. Curr. Opin. Neurobiol. 17, 43–52 (2007).
Xiao, A. et al. Chromosomal deletions and inversions mediated by TALENs and CRISPR/Cas in zebrafish. Nucleic Acids Res. 41, e141 (2013).
Hsu, P.D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Gagnon, J.A. et al. Efficient mutagenesis by Cas9 protein-mediated oligonucleotide insertion and large-scale assessment of single-guide RNAs. PLoS One 9, e98186 (2014).
Howe, K. et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature 496, 498–503 (2013).
Kelly, A. & Hurlstone, A.F. The use of RNAi technologies for gene knockdown in zebrafish. Brief Funct. Genomics 10, 189–196 (2011).
Schulte-Merker, S. & Stainier, D.Y.R. Out with the old, in with the new: reassessing morpholino knockdowns in light of genome editing technology. Development 141, 3103–3104 (2014).
Hruscha, A. et al. Efficient CRISPR/Cas9 genome editing with low off-target effects in zebrafish. Development 140, 4982–4987 (2013).
Twigg, S.R.F. et al. Cellular interference in craniofrontonasal syndrome: males mosaic for mutations in the X-linked EFNB1 gene are more severely affected than true hemizygotes. Hum. Mol. Genet. 22, 1654–1662 (2013).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Bassett, A.R., Tibbit, C., Ponting, C.P. & Liu, J.-L. Highly efficient targeted mutagenesis of Drosophila with the CRISPR/Cas9 system. Cell Rep. 4, 220–228 (2013).
Lo, T.-W. et al. Precise and heritable genome editing in evolutionarily diverse nematodes using TALENs and CRISPR/Cas9 to engineer insertions and deletions. Genetics 195, 331–348 (2013).
Nakayama, T. et al. Cas9-based genome editing in Xenopus tropicalis. Methods Enzymol. 546, 355–375 (2014).
Yen, S.-T. et al. Somatic mosaicism and allele complexity induced by CRISPR/Cas9 RNA injections in mouse zygotes. Dev. Biol. 393, 3–9 (2014).
Yu, C., Zhang, Y., Yao, S. & Wei, Y. A PCR based protocol for detecting indel mutations induced by TALENs and CRISPR/Cas9 in zebrafish. PLoS One 9, e98282 (2014).
Pan, L. et al. Rapid identification and recovery of ENU-induced mutations with next-generation sequencing and paired-end low-error analysis. BMC Genomics 16, 1263 (2015).
Breese, M.R. & Liu, Y. NGSUtils: a software suite for analyzing and manipulating next-generation sequencing datasets. Bioinformatics 29, 494–496 (2013).
Güell, M., Yang, L. & Church, G.M. Genome editing assessment using CRISPR Genome Analyzer (CRISPR-GA). Bioinformatics 30, 2968–2970 (2014).
Kimmel, C.B., Ballard, W.W., Kimmel, S.R., Ullmann, B. & Schilling, T.F. Stages of embryonic development of the zebrafish. Dev. Dyn. 203, 253–310 (1995).
Higashijima, S., Hotta, Y. & Okamoto, H. Visualization of cranial motor neurons in live transgenic zebrafish expressing green fluorescent protein under the control of the islet-1 promoter/enhancer. J. Neurosci. 20, 206–218 (2000).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Acknowledgements
We thank R. Garcia for superb animal care, the Moens lab for discussion and editing, J. Spiewak and the Parichy lab at the University of Washington for slc24a5 mutant embryos, S. Peterson and the Postlethwait lab at the University of Oregon for advice on sgRNA cloning, and I. Phelps and D. Doherty for help and access to the MiSeq machine. Funding was provided by the National Institutes of Health (R01HD076585 and R21NS076950 to C.B.M. and K99NS085035 to A.C.M.).
Author information
Authors and Affiliations
Contributions
A.N.S., C.F.D., A.C.W. and A.C.M. performed experiments, acquired and quantified data, and generated images for publication. A.C.M., A.N.S. and C.B.M. wrote the manuscript. All authors edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Optimizing Cas9 and sgRNA concentrations for increasing mutation efficiency.
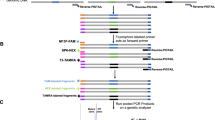
(a) Wild-type, 2-dpf slc24a5b1 embryos and a series of slc24a5 CRISPR-injected embryos; anterior is to the left. Wild-type pigmentation is lost in slc24a5 (‘golden’) mutants. Injection of Cas9-encoding mRNA and an sgRNA targeting slc24a5 resulted in embryos with mosaic loss of eye pigmentation (a ʹ). (b) Model of the slc24a5 CRISPR target site with arrows denoting PCR primers used for sequence analysis of the locus. (c) Sequencing from embryos injected with 100/1,200 pg of slc24a5 sgRNA/Cas9-encoding mRNA. Deletions are denoted by dashes (-), and insertions are in bold. The underlined sequence denotes the NGG motif used by Cas9. (d) Quantitation of eye pigmentation as assessed by mean pixel gray value of the eye with varying amounts of Cas9-encoding mRNA and a constant 100 pg of slc24a5 sgRNA. Each open triangle represents an individual animal. (e,f) Quantitation of toxicity seen in injected embryos for varying amounts of Cas9-encoding mRNA and a constant 100 pg of slc24a5 sgRNA (e) or a constant 1,200 pg of Cas9 with varying slc24a5 sgRNA (f). Toxicity encompassed embryo death, edema, localized cell death and general developmental defects. In d, N = 24 for each condition; error bars denote s.e.m. In e and f, N > 77 embryos for each point.
Supplementary Figure 2 Optimized CRISPR can recapitulate known mutant defects in injected embryos.
(a) Model of pk1b and vangl2 sgRNA target sites. Underlined sequence denotes the NGG motif used by Cas9. (b) Images are 20-µm dorsal-view projections of the hindbrain at 2 dpf. Anterior is up. Scale bar, 20 µm. Larvae are transgenic for Tg(isl1:GFP)rw0, which marks the branchiomotor neurons. A subset of these, the facial branchiomotor neurons (shown), undergo a stereotypical migration from rhombomere 4 (r4) to r6. Mutants in the planar cell polarity pathway (pk1b and vangl2) cause a failure in this migration. These phenotypes can be recapitulated in injected CRISPR F0 embryos. (c) Quantitation of migration defects. (d) Lateral views of 1-dpf embryos. Scale bar, 1 mm. vangl2 mutants have defects in anterior-posterior axis elongation due to defects in mesodermal convergence and extension. This can be recapitulated in injected CRISPR F0 embryos. (e) Quantitation of convergent extension defect. In c and e, N > 24 embryos for each bar.
Supplementary Figure 3 Optimizing Cas9 and sgRNA concentrations for electrical-synapse phenotypes.
(a) Quantitation of mosaic electrical-synapse loss in injected embryos for varying amounts of gjd1a sgRNA injected with 1,200 pg of Cas9-encoding mRNA. (b) Quantitation of toxicity seen in injected embryos for varying amounts of gjd1a sgRNA injected with 1,200 pg of Cas9-encoding mRNA. In a, N = 24 embryos for each bar. In b, N > 86 embryos for each point.
Supplementary Figure 4 Analysis of sgRNAs and genomic alterations in CRISPR injected embryos revealed by targeted deep sequencing of screen targets.
(a) Comparison of GC content versus indel frequency. Individual targets are denoted by open squares. The solid line is a linear trend line of the data (R2 = 0.0233). (b) Analysis of sgRNA indel efficiency based on binding the non-coding (–, N = 10) or coding (+, N = 32) strand. Box-and-whisker plot denotes median, quartiles, and extremes of the data. (c) Heat map for positions 2 to N of the sgRNA showing the relative increase in indel efficiency for each specific position. N is the first position of the PAM sequence. Color scale represents increased (orange) or decreased (cyan) indel efficiency relative to average. (d) Percentage of indels that caused out-of-frame changes to the target sequence. Individual targets are denoted by open squares. (e) Histogram of deletion positions relative to the NGG site bound by Cas9. Data are included from all targets aligned to the NGG. (f) Distribution of deletion sizes. Data are combined from all targets. The x-axes of e and f are log scales.
Supplementary Figure 5 Direct genotype-phenotype association for tjp1b and electrical-synapse loss.
(a) Each triangle represents an individual animal injected with 16.5/1,200 pg tjp1b_L sgRNA/Cas9-encoding mRNA. Spinal cords were analyzed for the loss of synapses and heads were assayed for mutational efficiency on the basis of qPCR analysis of the tjp1b locus. We found that heads and spinal cords had indistinguishable mutational efficiencies (heads, 0.59 ± 0.04; tail, 0.61 ± 0.05; P = 0.84; values are average ± s.e.m. compared by Student’s t-test). The square of the mutational efficiency is plotted on the x-axis on the basis of the expectation that phenotypes are expected to be due to biallelic loss of a gene. There is a strong correlation of genotype to phenotype (R2 = 0.98). The lines represent the expected genotype-phenotype association, assuming random mutagenesis and that deleterious alleles occur only with out-of-frame mutations (dotted line, y = (2/3)2x) or both in-frame and out-of-frame mutations (solid line, y = (3/3)2x). The fact that the phenotypic rate approaches 100% suggests either that in-frame deletions are deleterious or that heterozygous loss of tjp1b effects synapse formation. Note that we found that the qPCR genotypic analysis underestimated the mutational efficiency by 6%–15% (see Online Methods). This underestimation would result in an increased slope of genotype-phenotype association plotted here. (b–d) Images are 15-µm dorsal-view projections of two spinal cord segments at 5 dpf. Anterior is to the left. Scale bar, 10 µm. Larvae were stained for Connexin36 (Cx36, white) to mark the electrical synapses and for neurofilaments (RMO44, red) to mark neuronal processes, including M and CoLo. Individual Cx36 channel is shown in neighboring panel. Heterozygous loss of tjp1b has no effect on M synapse formation, whether the germline-transmitted mutation is out-of-frame (b, tjp1bfh449/+) or in-frame (c, tjp1bfh451/+). However, trans-heterozygous tjp1b mutants carrying an out-of-frame and an in-frame mutation (tjp1bfh449/fh451) lose electrical synapses, confirming that in-frame mutations are deleterious at this locus (d). In a, N = 32 embryos.
Supplementary Figure 6 Retesting sgRNAs with low indel rates from the screen revealed that most were due to degradation.
(a) qPCR quantitation of mutational efficiency in injected embryos for each of the failed target sgRNAs from the screen and a newly synthesized version. Each was individually injected at 100/1,200 pg sgRNA/Cas9-encoding mRNA. Mutation efficiency was assessed by qPCR and was done in triplicate; error bars denote s.e.m. (b) nrxn1b_S contained two polymorphisms in the designed sgRNA as compared to the genomic sequence of the fish into which it was injected. These differences likely reflect its failure to induce indels. In a, each bar represents five embryos pooled in three replicates; error bars denote s.e.m.
Supplementary Figure 7 Visual method for design, synthesis, quality control and injection of sgRNAs and Cas9-encoding mRNA into zebrafish embryos.
See the Online Methods for details on each step.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Table 1 (PDF 5579 kb)
Rights and permissions
About this article
Cite this article
Shah, A., Davey, C., Whitebirch, A. et al. Rapid reverse genetic screening using CRISPR in zebrafish. Nat Methods 12, 535–540 (2015). https://doi.org/10.1038/nmeth.3360
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3360
This article is cited by
-
Exploring hematopoiesis in zebrafish using forward genetic screening
Experimental & Molecular Medicine (2024)
-
Directionality of developing skeletal muscles is set by mechanical forces
Nature Communications (2023)
-
A CRISPR-Cas9-mediated F0 screen to identify pro-regenerative genes in the zebrafish retinal pigment epithelium
Scientific Reports (2023)
-
Large-scale F0 CRISPR screens in vivo using MIC-Drop
Nature Protocols (2023)
-
Generation of Zebrafish Models of Human Retinitis Pigmentosa Diseases Using CRISPR/Cas9-Mediated Gene Editing System
Molecular Biotechnology (2023)