Abstract
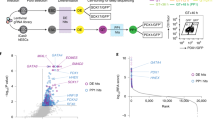
The genomic regulatory programmes that underlie human organogenesis are poorly understood. Pancreas development, in particular, has pivotal implications for pancreatic regeneration, cancer and diabetes. We have now characterized the regulatory landscape of embryonic multipotent progenitor cells that give rise to all pancreatic epithelial lineages. Using human embryonic pancreas and embryonic-stem-cell-derived progenitors we identify stage-specific transcripts and associated enhancers, many of which are co-occupied by transcription factors that are essential for pancreas development. We further show that TEAD1, a Hippo signalling effector, is an integral component of the transcription factor combinatorial code of pancreatic progenitor enhancers. TEAD and its coactivator YAP activate key pancreatic signalling mediators and transcription factors, and regulate the expansion of pancreatic progenitors. This work therefore uncovers a central role for TEAD and YAP as signal-responsive regulators of multipotent pancreatic progenitors, and provides a resource for the study of embryonic development of the human pancreas.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Accession codes
References
Fang, H. et al. An organogenesis network-based comparative transcriptome analysis for understanding early human development in vivo and in vitro. BMC Syst. Biol. 5, 108 (2011).
Fang, H. et al. Transcriptome analysis of early organogenesis in human embryos. Dev. Cell 19, 174–184 (2010).
Pan, F. C. & Wright, C. Pancreas organogenesis: from bud to plexus to gland. Dev. Dynam. 240, 530–565 (2011).
Zaret, K. S. & Grompe, M. Generation and regeneration of cells of the liver and pancreas. Science 322, 1490–1494 (2008)10.1126/science.1161431
Lango Allen, H. et al. GATA6 haploinsufficiency causes pancreatic agenesis in humans. Nat. Genet. 44, 20–22 (2012).
Xuan, S. et al. Pancreas-specific deletion of mouse Gata4 and Gata6 causes pancreatic agenesis. J. Clin. Invest. 122, 3516–3528 (2012).
Carrasco, M., Delgado, I., Soria, B., Martín, F. & Rojas, A. GATA4 and GATA6 control mouse pancreas organogenesis. J. Clin. Invest. 122, 3504–3515 (2012).
Offield, M. F. et al. PDX-1 is required for pancreatic outgrowth and differentiation of the rostral duodenum. Development 122, 983–995 (1996).
Stoffers, D. A., Zinkin, N. T., Stanojevic, V., Clarke, W. L. & Habener, J. F. Pancreatic agenesis attributable to a single nucleotide deletion in the human IPF1 gene coding sequence. Nat. Genet. 15, 106–110 (1997).
Haumaitre, C. et al. Lack of TCF2/vHNF1 in mice leads to pancreas agenesis. Proc. Natl Acad. Sci. USA 102, 1490–1495 (2005).
Jacquemin, P. et al. Transcription factor hepatocyte nuclear factor 6 regulates pancreatic endocrine cell differentiation and controls expression of the proendocrine gene ngn3. Mol. Cell. Biol. 20, 4445–4454 (2000).
Gao, N. et al. Dynamic regulation of Pdx1 enhancers by Foxa1 and Foxa2 is essential for pancreas development. Genes Dev. 22, 3435–3448 (2008).
Piper, K. et al. Novel SOX9 expression during human pancreas development correlates to abnormalities in Campomelic dysplasia. Mech. Dev. 116, 223–226 (2002).
Seymour, P. A. et al. SOX9 is required for maintenance of the pancreatic progenitor cell pool. Proc. Natl Acad. Sci. USA 104, 1865–1870 (2007).
Krapp, A. et al. The p48 DNA-binding subunit of transcription factor PTF1 is a new exocrine pancreas-specific basic helix-loop-helix protein. EMBO J. 15, 4317–4329 (1996).
Jennings, R. E. et al. Development of the human pancreas from foregut to endocrine commitment. Diabetes 62, 3514–3522 (2013).
Cho, C. H. H. et al. Inhibition of activin/nodal signalling is necessary for pancreatic differentiation of human pluripotent stem cells. Diabetologia 55, 3284–3295 (2012).
Xie, R. et al. Dynamic chromatin remodeling mediated by polycomb proteins orchestrates pancreatic differentiation of human embryonic stem cells. Cell Stem Cell 12, 224–237 (2013)
Kroon, E., Martinson, L. A., Kadoya, K. & Bang, A. G. Pancreatic endoderm derived from human embryonic stem cells generates glucose-responsive insulin-secreting cells in vivo. Nature 26, 443–452 (2008)
Borowiak, M., Maehr, R., Chen, S., Chen, A. E. & Tang, W. Small molecules efficiently direct endodermal differentiation of mouse and human embryonic stem cells. Cell Stem Cell 4, 348–358 (2009)
Rodríguez-Seguel, E. et al. Mutually exclusive signaling signatures define the hepatic and pancreatic progenitor cell lineage divergence. Genes Dev. 27, 1932–1946 (2013).
Cortijo, C., Gouzi, M., Tissir, F. & Grapin-Botton, A. Planar cell polarity controls pancreatic β cell differentiation and glucose homeostasis. Cell Rep. 2, 1593–1606 (2012).
Rada-Iglesias, A., Bajpai, R., Swigut, T. & Brugmann, S. A. A unique chromatin signature uncovers early developmental enhancers in humans. Nature 470, 279–283 (2011)
Creyghton, M. P. et al. Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc. Natl Acad. Sci. USA 107, 21931–21936 (2010).
Pasquali, L. et al. Pancreatic islet enhancer clusters enriched in type 2 diabetes risk-associated variants. Nat. Genet. 46, 136–143 (2014).
Oliver-Krasinski, J. M. & Stoffers, D. A. On the origin of the β cell. Genes Dev. 22, 1998–2021 (2008).
Zhao, B. et al. TEAD mediates YAP-dependent gene induction and growth control. Genes Dev. 22, 1962–1971 (2008).
George, N. M., Day, C. E., Boerner, B. P., Johnson, R. L. & Sarvetnick, N. E. Hippo Signaling Regulates Pancreas Development through Inactivation oelf Yap. Mol. Cell. Biol. 32, 5116–5128 (2012).
Liu-Chittenden, Y. et al. Genetic and pharmacological disruption of the TEAD-YAP complex suppresses the oncogenic activity of YAP. Genes Dev. 26, 1300–1305 (2012).
Elghazi, L., Cras-Méneur, C., Czernichow, P. & Scharfmann, R. Role for FGFR2IIIb-mediated signals in controlling pancreatic endocrine progenitor cell proliferation. Proc. Natl Acad. Sci. USA 99, 3884–3889 (2002).
Lynn, F. C. et al. Sox9 coordinates a transcriptional network in pancreatic progenitor cells. Proc. Natl Acad. Sci. USA 104, 10500–10505 (2007).
Sawada, A. et al. Tead proteins activate the Foxa2 enhancer in the node in cooperation with a second factor. Development 132, 4719–4729 (2005).
Whyte, W. A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Weedon, M. N. et al. Recessive mutations in a distal PTF1A enhancer cause isolated pancreatic agenesis. Nat. Genet. 46, 61–64 (2014).
Hezel, A. F., Kimmelman, A. C., Stanger, B. Z., Bardeesy, N. & DePinho, R. A. Genetics and biology of pancreatic ductal adenocarcinoma. Genes Dev. 20, 1218–1249 (2006).
Rooman, I. & Real, F. X. Pancreatic ductal adenocarcinoma and acinar cells: a matter of differentiation and development? Gut 61, 449–458 (2012).
Kapoor, A. et al. Yap1 activation enables bypass of oncogenic Kras addiction in pancreatic cancer. Cell 158, 185–197 (2014).
Zhang, W. et al. Downstream of mutant KRAS, the transcription regulator YAP is essential for neoplastic progression to pancreatic ductal adenocarcinoma. Sci. Signal. 7, ra42 (2014).
Zhao, B., Tumaneng, K. & Guan, K. L. The Hippo pathway in organ size control, tissue regeneration and stem cell self-renewal. Nat. Cell Biol. 13, 877–883 (2011)
Bardeesy, N. & Stanger, B. Z. Hippo signaling regulates differentiation and maintenance in the exocrine pancreas. Gastroenterology 144, 1543–53–1553.e1 (2013).
Fujitani, Y. et al. Targeted deletion of a cis-regulatory region reveals differential gene dosage requirements for Pdx1 in foregut organ differentiation and pancreas formation. Genes Dev. 20, 253–266 (2006).
Gannon, M., Gamer, L. W. & Wright, C. V. Regulatory regions driving developmental and tissue-specific expression of the essential pancreatic gene pdx1. Dev. Biol. 238, 185–201 (2001).
O’Rahilly, R., Müller, F., Hutchins, G. M. & Moore, G. W. Computer ranking of the sequence of appearance of 73 features of the brain and related structures in staged human embryos during the sixth week of development. Am. J. Anat. 180, 69–86 (1987).
Maestro, M. A. et al. Hnf6 and Tcf2 (MODY5) are linked in a gene network operating in a precursor cell domain of the embryonic pancreas. Hum. Mol. Genet. 12, 3307–3314 (2003).
Piper, K. et al. β cell differentiation during early human pancreas development. J. Endocrinol. 181, 11–23 (2004).
Petzold, K. M. & Spagnoli, F. M. A system for ex vivo culturing of embryonic pancreas. J. Vis. Exp. 66, e3979 (2012).
Gaulton, K. J. et al. A map of open chromatin in human pancreatic islets. Nat. Genet. 42, 255–259 (2010).
Luco, R. F., Maestro, M. A., Sadoni, N., Zink, D. & Ferrer, J. Targeted deficiency of the transcriptional activator Hnf1α alters subnuclear positioning of its genomic targets. PLoS Genet. 4, e1000079 (2008).
van Arensbergen, J. et al. Derepression of Polycomb targets during pancreatic organogenesis allows insulin-producing β-cells to adopt a neural gene activity program. Genome Res. 20, 722–732 (2010).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Morán, I. et al. Human β cell transcriptome analysis uncovers lncRNAs that are tissue-specific, dynamically regulated, and abnormally expressed in type 2 diabetes. Cell Metab. 16, 435–448 (2012).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
McLean, C. Y., Bristor, D., Hiller, M. & Clarke, S. L. GREAT improves functional interpretation of cis-regulatory regions. Nature 28, 495–501 (2010).
Supek, F., Bošnjak, M., Škunca, N. & Šmuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS ONE 6, e21800 (2011).
de Hoon, M. J. L., Imoto, S., Nolan, J. & Miyano, S. Open source clustering software. Bioinformatics 20, 1453–1454 (2004).
Saldanha, A. J. Java Treeview—extensible visualization of microarray data. Bioinformatics 20, 3246–3248 (2004).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Gupta, S., Stamatoyannopoulos, J. A., Bailey, T. L. & Noble, W. S. Quantifying similarity between motifs. Genome Biol. 8, R24 (2007).
Whitlock, M. C. Combining probability from independent tests: the weighted Z-method is superior to Fisher’s approach. J. Evol. Biol. 18, 1368–1373 (2005).
Derrien, T. et al. Fast computation and applications of genome mappability. PLoS ONE 7, e30377 (2012).
Esni, F., Miyamoto, Y., Leach, S. D. & Ghosh, B. Primary explant cultures of adult and embryonic pancreas. Methods Mol. Med. 103, 259–271 (2005).
Skouloudaki, K. et al. Scribble participates in Hippo signaling and is required for normal zebrafish pronephros development. Proc. Natl Acad. Sci. USA 106, 8579–8584 (2009).
Chiang, E. F. et al. Two sox9 genes on duplicated zebrafish chromosomes: expression of similar transcription activators in distinct sites. Dev. Biol. 231, 149–163 (2001).
Argenton, F., Zecchin, E. & Bortolussi, M. Early appearance of pancreatic hormone-expressing cells in the zebrafish embryo. Mech. Dev. 87, 217–221 (1999).
Jowett, T. & Lettice, L. Whole-mount in situ hybridizations on zebrafish embryos using a mixture of digoxigenin- and fluorescein-labelled probes. Trends Genet. 10, 73–74 (1994).
Bessa, J. et al. Zebrafish enhancer detection (ZED) vector: A new tool to facilitate transgenesis and the functional analysis of cis-regulatory regions in zebrafish. Dev. Dynam. 238, 2409–2417 (2009).
Kawakami, K., Shima, A. & Kawakami, N. Identification of a functional transposase of the Tol2 element, an Ac-like element from the Japanese medaka fish, and its transposition in the zebrafish germ lineage. Proc. Natl Acad. Sci. USA 97, 11403–11408 (2000).
Binot, A-C. et al. Nkx6.1 and nkx6.2 regulate α- and β-cell formation in zebrafish by acting on pancreatic endocrine progenitor cells. Dev. Biol. 340, 397–407 (2010).
Acknowledgements
The research was supported by the National Institute for Health Research (NIHR) Imperial Biomedical Research Centre. Work was funded by grants from the Ministerio de Economía y Competitividad (CB07/08/0021, SAF2011-27086, PLE2009-0162 to J.F., BFU2013-41322-P to J.L.G-S.), the Andalusian Government (BIO-396 to J.L.G-S.), the Wellcome Trust (WT088566 and WT097820 to N.A.H., WT101033 to J.F.), the Manchester Biomedical Research Centre, ERC advanced starting grant IMDs (C.H-H.C. and L.V.) and the Cambridge Hospitals National Institute for Health Research Biomedical Research Centre (L.V.). R.E.J. is a Medical Research Council clinical training fellow. The authors are grateful to C. Wright (Vanderbilt University) for zebrafish Pdx1 antiserum, J. Postlethwait (Purdue University) for a Sox9b clone, H. Sasaki (Kumamoto University) for a TEAD–EnR clone, C. Vinod and L. Abi for research nurse assistance, and clinical colleagues at Central Manchester University Hospitals NHS Foundation Trust. The authors thank J. Garcia-Hurtado for technical assistance (IDIBAPS).
Author information
Authors and Affiliations
Contributions
J.F. coordinated the overall project and supervised epigenomic analysis and mouse studies, N.A.H. supervised human embryo characterization, L.V. supervised hESC differentiation studies and J.L.G-S. supervised zebrafish studies. I.C., S.A.R-S., C.H-H.C., J.B., M.R., M.L., M.C., A.B., M.A.M. and R.E.J. designed, carried out and analysed experiments. N.C. carried out experiments. I.C., S.A.R-S., J.P-C., L.P. and I.M. carried out computational analysis. I.C., S.A.R-S. and J.F. wrote the manuscript with contributions from C.H-H.C., J.B., M.R., M.L., J.P-C., N.A.H., J.L.G-S. and L.V.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Human in vitro MPCs recapitulate key features of in vivo MPCs.
(a) Immunohistochemistry analysis of in vivo MPCs from Carnegie stage 16–18 human embryonic pancreas, and immunofluorescence analysis of in vitro MPCs show expression of stage-delimiting MPC TFs in both sources of MPCs. (b) Heatmap showing RNA-seq FPKM signal in MPCs and 23 control tissues for TFs that are enriched in pancreatic MPCs and for a similar number of known lineage-specific non-pancreatic TFs. (c) Expression correlation matrix showing Spearman coefficient values for transcript levels from in vivo and in vitro MPCs vs. 23 control tissues. (d) Z score correlation density plots. Comparisons of in vivo MPCs with an unrelated tissue (fetal heart, left panel), or between tissues from the same lineage, but different stages (adult and fetal heart, right panel), do not show high correlation, in contrast with data presented in Fig. 1c that shows highly correlated Z-scores for in vivo and in vitro MPCs. Spearman coefficient values are shown for each comparison. Color scale depicts number of transcripts. (e) Motif discovery in different FOXA2 ChIP-seq datasets, shows a similar binding motif for this TF in all samples. P values and percentages of bound versus background regions are indicated below each motif logo. (f) Regions enriched in FOXA2 and H3K4me1 in chromatin from in vivo MPCs also show H3K4me1-enrichment in in vitro MPCs, but not in control samples (mammary epithelial cells, myotubes, CD133 + umbilical cord blood and hESCs). The heatmap shows FOXA2 and H3K4me1 signal centered on these regions (see Methods for details). Note that even though the regions were pre-selected from in vivo MPC data, H3K4me1-enrichment is stronger in chromatin from in vitro MPCs, reflecting the larger number of cells used for ChIP-seq. (g) H3K4me1 and FOXA2 signals in the whole genome were binned in 5 Kb for H3K4me1 and 1 Kb for FOXA2. These signals were highly correlated in biological replicates (R > 0.8).
Supplementary Figure 2 Human pancreatic MPC enhancers.
(a) Examples showing how in vitro MPCs recapitulate the epigenomic landscape of in vivo MPCs. HNF1B encodes a TF that important for pancreas development, FZD2 is a non-canonical WNT signaling component, and HES1 is a transcriptional repressor that controls growth and differentiation of pancreatic MPCs. (b) Enhancers were defined as H3K27ac islands in the in vitro MPCs that overlapped H3K4me1 islands in both in vitro and in vivo MPCs. We discarded regions overlapping promoters (1 Kb upstream and 2 Kb downstream of RefSeq TSS) and any regions smaller than 50 bp. This revealed 9,669 MPC enhancers. (c) MPC enhancers are tissue- and stage-selective. Enhancers were defined for 8 tissues in a similar manner to MPCs (Supplementary Table 8). Each pie chart shows in red the proportion of MPC enhancers that are inactive in each tissue. We defined MPC-selective enhancers as those that were inactive in at least 6 out of 7 non-pancreatic tissues. (d) Enriched annotated functions among genes that are associated with three or more MPC-selective enhancers. The graph shows fold enrichment values and P values calculated with GREAT53.
Supplementary Figure 3 MPC enhancers are enriched in TEAD motifs.
(a) De novo motif search in MPC-selective enhancers revealed strong enrichment for TEAD recognition sequences, similarly to what we observed for the whole set of MPC enhancers. Other enriched matrices match binding sites of known pancreatic regulators. (b) TEAD motifs are highly enriched in enhancers bound by FOXA2 in both in vivo and in vitro MPCs, but not in enhancers bound by FOXA2 in adult pancreatic islets.
Supplementary Figure 4 TEAD1 is a core component of the combination of TFs that bind to MPC enhancers.
(a) TEAD1 is expressed in PDX1+ in vivo MPCs from human pancreas of Carnegie stage 19. (b) De novo analysis of over-represented sequence motifs for regions bound by each of the TFs examined in this study. As expected, each dataset showed a top scoring motif that coincided with the immunoprecipitated TF. A marked co-enrichment of many known pancreatic TF and TEAD1 motifs was observed. P values and percentages of bound vs. background regions are indicated below each motif logo. (c) Examples showing CRMs bound by multiple TFs (regions highlighted in yellow). (d) TF binding and co-binding preferentially occurs at MPC enhancers. Note that TEAD1 binding and co-binding enrichment is comparable to the enrichments found for other TFs. Binding fold enrichment was calculated over 1,000 permutations of enhancer or promoter genomic positions. (e) MPC enhancers bound by any of the pancreatic TFs or TEAD1 show a high degree of co-binding with other TFs. Total number of peaks for each TF is shown below the corresponding column. (f) Representative examples of known Hippo pathway targets showing TEAD1 binding at their promoter regions.
Supplementary Figure 5 Functional validation of CRMs as transcriptional enhancers in pancreatic MPCs.
(a) Functional validation of CRMs as transcriptional enhancers in human progenitors. Thirty-two CRMs and 8 negative control regions were cloned into the pGL4.23 vector and tested in reporter assays. Reporter activity was compared to empty pGL4.23. ∗ Two-tailed Student’s t test P<0.05 (P values fully listed in Supplementary Table 22). n = 3-4 independent transfections per enhancer, 8 of 32 constructs were tested in an independent experiment that yielded comparable results. (b) Functional validation of unannotated CRMs identified in the vicinity of MPC-enriched genes in 24 hpf zebrafish embryos. Eight out of 10 TEAD1-bound CRMs yielded activation of a minimal promoter driving GFP (see also Fig. 5c–e and Supplementary Table 21). Pancreatic progenitors were identified by co-staining Nkx6.1 and either Pdx1 or insulin. Note that in zebrafish Nkx6.1 is expressed in early pancreatic progenitors but not in endocrine cells, unlike in mammalian embryos, which show Nkx6.1 expression in both cellular compartments68. The percentage of transgenics showing activation of GFP in the pancreatic domain for each CRM and in control injections is presented in the bar plot in Fig. 5e. Dashed lines demarcate the pancreatic progenitor domain (Nkx6.1+ cells). y: yolk autofluorescence, s: somites showing crossreactivity with anti-Pdx1 serum.
Supplementary Figure 6 Developmental expression of YAP.
(a) Immunofluorescence images of hESCs and different stages of differentiation show that YAP is strongly expressed in the nuclei of hESCs (white arrows), whereas a marked decrease in YAP immunoreactivity is observed in definitive endoderm and dorsal foregut stages (days 3 and 6)(white arrowheads). In days 3 and 6, YAP is not detected in a subset of SOX17+ and FOXA2+ cells, respectively (hollow arrowheads). (b) YAP is present in the nuclei of dorsal foregut endoderm cells of human Carnegie Stage 10 embryos. AIP: anterior intestinal portal, fg: foregut, lm: lateral mesoderm, nc: notochord, nt: neural tube. (c) Immunofluorescence images of mouse E10.5 and E12.5 embryonic pancreas show Tead1 and Yap expression in most nuclei of Pdx1 + MPCs (white arrows) and in the surrounding mesenchyme. Yap expression is absent in glucagon-expressing endocrine cells (hollow arrowheads). The squares in the leftmost panels depict areas shown at higher power in other panels. du: duodenum, dp: dorsal pancreas. (d,e) Yap is broadly expressed in the nuclei of pancreatic mesenchyme and epithelium from E12.5 and E14.5 mouse embryonic pancreas, yet shows cytoplasmic localization in Cpa1 + progenitor cells (white arrowheads) and is undetectable in early Pax6 + endocrine cells (hollow arrowheads). (f) In the adult mouse pancreas Yap is present in nuclei from ducts (white arrows), and not in endocrine (hollow arrowheads) or acinar cells (white arrowheads). (g) YAP is expressed in nuclei of SOX9 + epithelial cells but absent in insulin-expressing endocrine cells from 14 weeks post-conception (WPC) human pancreas. (h) PDX1 co-stains with YAP and TEAD1 in the nucleus of in vitro MPCs. (i) Nuclear YAP is not detected in differentiated insulin-expressing cells derived from hESCs (hollow arrowheads).
Supplementary Figure 7 Knockdown of Yap1 or dominant inhibition of Tead reduces pancreas size in zebrafish.
(a) Injection of a morpholino targeting yap1 (Mo-yap1) or mRNA encoding a TEAD protein fused to the transcriptional repressor domain of Engrailed (TEAD-EnR) decreased the number of insulin expressing cells detected by in situ hybridization. This phenotype was rescued by co-injection of Mo-yap1 with an in vitro synthesized yap1 mRNA that is not sensitive to morpholino inhibition. The percentage of embryos from each condition showing reduced insulin-positive cells was quantified as an indication of pancreatic hypoplasia, and displayed in the graph shown on the side (n = 46-71 embryos per condition). Scale bar = 0.25 mm. (b) Mo-yap1 increased ectopic expression of pancreatic markers. The panels show insulin in situ hybridization in control and Morpholino-treated 24 hpf zebrafish embryos. Scale bar = 0.25 mm.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1314 kb)
Supplementary Tables 1–23
Supplementary Information (XLSX 6232 kb)
Rights and permissions
About this article
Cite this article
Cebola, I., Rodríguez-Seguí, S., Cho, CH. et al. TEAD and YAP regulate the enhancer network of human embryonic pancreatic progenitors. Nat Cell Biol 17, 615–626 (2015). https://doi.org/10.1038/ncb3160
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3160
This article is cited by
-
Inhibition of the YAP-MMB interaction and targeting NEK2 as potential therapeutic strategies for YAP-driven cancers
Oncogene (2024)
-
A statistical method for quantifying progenitor cells reveals incipient cell fate commitments
Nature Methods (2024)
-
Expansion of ventral foregut is linked to changes in the enhancer landscape for organ-specific differentiation
Nature Cell Biology (2023)
-
Context-dependent transcriptional regulations of YAP/TAZ in stem cell and differentiation
Stem Cell Research & Therapy (2022)
-
Stepwise differentiation of functional pancreatic β cells from human pluripotent stem cells
Cell Regeneration (2022)