Abstract
Dendritic cells serve a key function in host defence, linking innate detection of microbes to activation of pathogen-specific adaptive immune responses1,2. Whether there is cell-intrinsic recognition of human immunodeficiency virus (HIV) by host innate pattern-recognition receptors and subsequent coupling to antiviral T-cell responses is not yet known3. Dendritic cells are largely resistant to infection with HIV-14, but facilitate infection of co-cultured T-helper cells through a process of trans-enhancement5,6. Here we show that, when dendritic cell resistance to infection is circumvented7,8, HIV-1 induces dendritic cell maturation, an antiviral type I interferon response and activation of T cells. This innate response is dependent on the interaction of newly synthesized HIV-1 capsid with cellular cyclophilin A (CYPA) and the subsequent activation of the transcription factor IRF3. Because the peptidylprolyl isomerase CYPA also interacts with HIV-1 capsid to promote infectivity, our results indicate that capsid conformation has evolved under opposing selective pressures for infectivity versus furtiveness. Thus, a cell-intrinsic sensor for HIV-1 exists in dendritic cells and mediates an antiviral immune response, but it is not typically engaged owing to the absence of dendritic cell infection. The virulence of HIV-1 may be related to evasion of this response, the manipulation of which may be necessary to generate an effective HIV-1 vaccine.
Similar content being viewed by others
Main
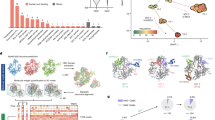
The exposure of monocyte-derived dendritic cells (MDDCs) to GFP-encoding HIV-1 pseudotyped with vesicular stomatitis virus protein G (VSV-G) (hereafter referred to as HIV-GFP(G); multiplicity of infection (MOI) 1–2, see Supplementary Table 1) resulted in little infection and the absence of cell activation, as monitored by the expression of CD86, CD80, CD38 and CD83 (Fig. 1a and Supplementary Fig. 2a). Likewise, VSV-G-pseudotyped SIVmac239 virus-like particles (SIV-VLP(G)) had no effect on dendritic cell activation. In contrast, co-infection of MDDCs with HIV-GFP(G) and SIV-VLP(G), which provides Vpx-mediated relief of restriction to HIV-1 replication8, resulted in GFP expression in more than 85% of the cells as well as upregulation of CD86 and other activation markers after 48 h (Fig. 1a and Supplementary Fig. 2a). Entry of both virions into the cytoplasm was required (Supplementary Fig. 2b), and activation occurred only beyond a variable threshold of infection (Supplementary Figs 2c and 3a). A virus that expressed all accessory proteins and a CCR5-tropic replication-competent virus also infected MDDCs in the presence of SIV-VLP(G) and induced expression of CD86 (Supplementary Figs 3b and 4a). Expression of a Vpx–Vpr fusion protein in packaging cells rescued the ability of HIV-GFP(G) to productively infect MDDCs and to induce CD86 upregulation, indicating that Vpx is the only SIV-VLP component required for infection with HIV-1 (Supplementary Fig. 4b). Thus, MDDCs have an intact mechanism of activation after HIV-1 infection.
a, GFP expression and CD86 surface expression in MDDCs at 48 h after infection with HIV-GFP(G) and SIV-VLP(G), alone or in combination. b, Immunoblot of phospho-STAT1 and total actin expression over time in MDDCs infected with HIV-GFP(G) and SIV-VLP(G) or treated with LPS. c, Type I IFN activity in the supernatant of MDDCs or activated CD4+ T cells infected with dilutions of HIV-GFP(G) with SIV-VLP(G) or with Sendai virus (SeV). d, Type I IFN activity in supernatants of MDDCs, 293FT and THP-1 cells infected with dilutions of HIV-GFP(G) with or without SIV-VLP(G) or transfected with poly(I:C) or RNA from Newcastle disease virus (NDV)-infected cells using lipofectamine (lipo).
As observed with MDDCs, infection of primary peripheral blood CD11c+ dendritic cells with HIV-GFP(G) did not result in detectable expression of GFP (Supplementary Fig. 4c). However, CD11c+ dendritic cells were infected with HIV-GFP(G) in the presence of SIV-VLP(G), resulting in upregulation of CD86 in a proportion of cells similar to that observed after incubation with polyriboinosinic:polyribocytidylic acid (poly(I:C)).
Genome-wide expression profiling demonstrated induction of a type I interferon (IFN) response following co-infection, but not following infection with either HIV-1 or SIV particles (Supplementary Figs 5a–c). After infection, expression of IFN-regulated genes was delayed as compared to the response to lipopolysaccharide (LPS) (Supplementary Fig. 5d). Accordingly, STAT1 phosphorylation was present at 2 h after LPS treatment, but only at 22 h after infection (Fig. 1b). Type I IFN was produced by infected MDDCs over the course of 48 h (Supplementary Fig. 6a), but was not detected after infection of CD4+ T cells (Fig. 1c), 293T cells and THP-1 cells (Fig. 1d), despite the ability of these cells to produce type I IFN after other viral innate stimuli. Blocking antibodies against IFN-β, but not IFN-α, reduced expression of the activation markers on MDDCs (Supplementary Fig. 6b, c). Further neutralization of type I and type III IFN did not improve the inhibition (data not shown). Together, these results indicate that CD86 induction is mainly due to the production of soluble type I IFN-β, in accordance with observations in murine dendritic cells9.
Next we sought to determine which step of the viral replication cycle is required for MDDC activation. Inhibitors of HIV-1 reverse transcriptase (zidovudine (AZT)) and integrase (raltegravir) inhibited transduction efficiency and MDDC activation only when added during the first 24 h (Supplementary Fig. 7a) and had no effect on LPS- or poly(I:C)-induced CD86 upregulation (data not shown). These results indicated that dendritic cell activation is induced after integration. There was no activation of MDDCs after infection with an HIV-1-based vector devoid of viral-protein-coding sequences (LKO1gfp) (Supplementary Fig. 7b). Therefore, we introduced mutations in the packaged HIV-GFP genome and evaluated activation of infected MDDCs. Inactivation of Rev, which is required for nuclear export of unspliced viral RNA10, and abrogation of Gag expression prevented MDDC activation (Fig. 2a); however, mutation of the PTAP sequence in p6, required for viral budding11, and treatment with HIV-1 protease inhibitors (Supplementary Fig. 7c) had no effect. In the absence of SIV-VLP(G), intracellular HIV-1 capsid from incoming viral particles failed to induce CD86 expression (Supplementary Fig. 8). These results indicated that newly synthesized GagPol is required for dendritic cell activation, which is consistent with the delayed induction of the type I IFN response. Next we tested a panel of viruses with HIV-1-capsid mutations12 for the ability to induce the innate response in MDDCs. These mutants were defective for dendritic cell infection (Supplementary Fig. 9) and were thus partially rescued by co-transfection of wild-type viral proteins in the packaging cells. The T54A/N57A and Q63A/Q67A mutants induced CD86 expression despite reduced infectivity compared to wild-type virus (Fig. 2b and Supplementary Fig. 10). In contrast, infection with the G89V mutant, which is compromised for HIV-1-capsid binding to CYPA, a peptidylprolyl isomerase required for optimal HIV-1 infectivity13, resulted in substantially reduced CD86 expression at similar levels of infection.
a, GFP and CD86 expression in MDDCs infected with HIV-GFP(G) or its mutants, ΔRev, ΔGag or PTAP− in the presence of SIV-VLP(G). HIV-GFP was rescued in all cases by co-expression of wild-type proteins in packaging cells. b, Effect of HIV-1 capsid mutations on the proportion of GFP+-infected MDDCs that express CD86. MDDCs were infected with serially diluted wild-type (WT) (pLaiΔEnv-GFP3(G)) or HIV-1 capsid mutants (T54A/N57A, Q63A/Q67A or G89V), in the presence of SIV-VLP(G). HIV-1 capsid mutant infectivity was rescued by co-expression of wild-type proteins in packaging cells. *P < 0.026 (n = 9). c, Effect of cyclosporin A and FK506 on expression of CD86 in MDDCs infected with HIV-GFP(G) and SIV-VLP(G) or after treatment with LPS. Cyclosporin A and FK506 target the calcineurin pathway but FK506 does not bind to CYPA. d, Expression of GFP, RFP and CD86 in HIV-infected cells after CYPA knockdown by RNAi. MDDCs were transduced with GFP-encoding control shRNA vector or a shRNA vector targeting CYPA, in the presence of SIV-VLP(G), and subsequently challenged with HDV-IRES-RFP(G) or treated with LPS. Right, cells are gated on GFP+ populations shown on the left. Experiments were performed on a total of at least six donors, except c, which was performed on four donors.
Treatment with cyclosporin A, which disrupts the interaction between CYPA and the HIV-1 capsid14, prevented MDDC activation after infection with HIV-GFP(G) and SIV-VLP(G), but not after treatment with LPS (Fig. 2c). Because cyclosporin A also inhibits infection with HIV-1, we assessed its effect when administered at different times after infection of MDDCs. When cyclosporin A was added as late as 12 h after infection, it prevented upregulation of CD86 despite highly efficient infection and expression of HIV-1 capsid (Supplementary Fig. 11).
To study the role of CYPA and other host genes in innate immune signalling following productive infection of MDDCs with HIV-1, we used an RNA interference approach using short hairpin RNA lentiviral vectors (that also express GFP)15 along with SIV-VLP(G). Knockdown of CYPA (also known as PPIA) markedly reduced expression of its product and prevented CD86 upregulation following infection with HIV-1 (HDV-IRES-RFP(G), a VSV-G-pseudotyped HIV-derived vector encoding an IRES-regulated RFP reporter, see Methods), but not after treatment with LPS (Fig. 2d and Supplementary Fig. 12a). The interaction between CYPA and newly synthesized HIV-1 capsid is therefore essential for the innate response of MDDCs to HIV-1.
Type I IFN responses following infection with several viruses require the phosphorylation, dimerization and nuclear translocation of IRF316. Productive infection of MDDCs with HIV-1 resulted in cyclosporin-A-sensitive nuclear accumulation of phosphorylated IRF3 (Fig. 3a). Knockdown of IRF3 in MDDCs abrogated the induction of CD86 after infection with HIV-1 and, as expected, after treatment with LPS or poly(I:C) but not after treatment with curdlan (β-1,3-glucan), indicating that IRF3 knockdown did not lead to an intrinsic defect in CD86 expression (Fig. 3b and Supplementary Fig. 12b). IRF3 knockdown, as well as CYPA knockdown, also increased the threshold at which virus induced CD86 and CD38 (Supplementary Fig. 12c, d).
a, Tubulin, histone H3, IRF3 and phospho-Ser396-IRF3 expression in cytoplasmic and nuclear fractions of MDDCs infected with SIV-VLP(G) and dilutions of HIV-GFP(G) in the presence or the absence of cyclosporin A. Cells were harvested 8 h after infection or after control treatment with poly(I:C). b, GFP, RFP and CD86 expression in MDDCs initially transduced with GFP-encoding-control shRNA vector or a shRNA vector targeting IRF3, and subsequently challenged with HDV-IRES-RFP(G) or treated with poly(I:C) or curdlan. Right, cells are gated on GFP+ transduced populations.
To determine if productive infection and subsequent activation of MDDCs influence antiviral adaptive immunity, first we examined whether HIV-infected dendritic cells could activate HIV-1 Gag-specific CD4+ and CD8+-T-cell clones. In the presence of MDDCs incubated with HIV-1 alone, low levels of IFNγ were detected17. In contrast, MDDCs infected with HIV-GFP(G) and SIV-VLP(G) stimulated a high proportion of major histocompatibility complex (MHC) class I and class II restricted T-cell clones to produce IFN-γ (Supplementary Fig. 13a). Maturation induced by unrelated Toll-like receptor (TLR) ligands coupled with abortive HIV infection was not sufficient for MDDCs to potently stimulate HIV-antigen-specific T cells (Supplementary Fig. 13b).
To measure directly the contribution of co-stimulation to T-cell activation, we examined the polyclonal proliferation of naive CD4+ T cells in response to infected dendritic cells in the presence of suboptimal concentrations of anti-CD3 antibody18,19,20. Under these conditions, T cells that were co-cultured with productively infected and activated MDDCs proliferated through several cell cycles whereas T cells cultured with the abortively infected or uninfected MDDCs showed little proliferation (Fig. 4a and Supplementary Fig. 14). Next we examined the effect of SCY, a non-immunosuppressive cyclosporin A analogue21,22 that, unlike cyclosporin A, does not have any direct effect on the activation or proliferation of T cells21 (Supplementary Fig. 15a). SCY inhibited dendritic cell activation induced by HIV-GFP(G) similarly to cyclosporin A at similar levels of infection. Dendritic cells treated with SCY or the reverse transcriptase inhibitor AZT at the time of HIV-GFP(G) and SIV-VLP(G) infection showed a reduced ability to induce proliferation (Fig. 4b and Supplementary Figs 15b and 16), as did dendritic cells infected with the G89V HIV-1 capsid mutant (Supplementary Fig. 17). These results are consistent with a requirement for interaction of newly synthesized HIV-1 capsid with CYPA in the induction of dendritic cell co-stimulatory activity.
a, GFP and CD86 expression in control and HIV-1-infected dendritic cells (top) and carboxyfluorescein succinimidyl ester (CFSE) dilution (bottom) in CFSE-labelled naive CD4+ T cells cultured with the dendritic cells for 4 days in the presence of anti-CD3 antibody. b, GFP and CD86 expression in dendritic cells (top) and CFSE dilution (bottom) in naive CD4+ T cells cultured for 4 days with untreated dendritic cells or dendritic cells treated with 25 µM AZT or 1 µM SCY after infection. c, Induction of a type-I-IFN-dependent antiviral state inhibits MDDC-dependent trans-enhancement. MDDCs were infected with dilutions of HDV-IRES-RFP(G) with or without SIV-VLP(G) in the presence or absence of type-I-IFN-neutralizing reagents. Activated CD4+ T cells and a CCR5-tropic HIV-1-GFP (R5–GFP) were added 2 days later. RFP and CD86 expression and GFP expression were measured in dendritic cells and CD4+ T cells, respectively. Trans-enhancement is indicated by the increase in GFP+ T cells in the presence or absence of MDDCs in the top panel (error bars indicate standard error of the mean for three independent donors).
Trans-enhancement by MDDCs of CD4+ T-cell infection with a CCR5-tropic virus encoding GFP was inhibited if the dendritic cells were previously infected with HIV-1. The inhibition was relieved by neutralizing antibody against IFN-β, indicating that the innate response to HIV-1 in dendritic cells restricts infection of surrounding T cells (Fig. 4c) and suggesting that activation of such a response may also limit infection in vivo.
Our results show that, in contrast to CD4+ T cells, human dendritic cells have intrinsic machinery for responding to infection with HIV-1 and for activating antiviral defences and adaptive immunity (Supplementary Fig. 1). However, they are unlikely to do so effectively in infected individuals because HIV-1 fails to replicate in dendritic cells. HIV-2, which is not pandemic23, encodes Vpx and has the potential to infect and activate MDDCs in a CYPA-dependent manner (Supplementary Fig. 18), which is consistent with the reported ability of HIV-2 capsid to bind human CYPA24,25. The finding that newly synthesized HIV-1 capsid is required to induce dendritic cell activation through a pathway involving CYPA and IRF3 implicates an intracellular viral protein—in addition to the already known nucleic acids of numerous other viruses—among the type I IFN-inducing pathogen-associated molecular patterns26 and constitutes the first description of a cell-intrinsic recognition mechanism of retroviruses26. It will be important to determine whether the mechanism described herein contributes to the control of viral load in individuals infected with HIV-2, as well as in HIV-1-infected long-term non-progressors or ‘elite controllers’27. A better mechanistic understanding of this dendritic-cell-intrinsic signalling pathway may also inform HIV vaccine development.
Methods Summary
Monocytes were isolated and incubated with granulocyte–macrophage colony-stimulating factor (GM-CSF) and IL-4 to induce dendritic cell differentiation. Pseudotyped viruses and virus-like particles were produced by transient transfection of 293FT cells using TransIT-293 (Mirus). Infections were performed by incubating 105 MDDCs in 96-well round-bottomed plates in the presence of 5 µg ml−1 polybrene. Cell-surface staining of activation markers was performed 48 h after infection. shRNA vectors carrying GFP were transduced into fresh monocytes together with SIV-VLP(G) and dendritic cell differentiation was induced. More than 90% of cells were routinely transduced and cells were challenged at day 4 with HDV-IRES-RFP(G) or other control pathogen-associated molecular patterns.
Online Methods
Cells
GHOST28 and 293FT (Invitrogen) cells were cultured in DMEM, 10% fetal bovine serum (FBS) (HyClone) and antibiotics. Peripheral blood mononuclear cells were isolated from Institutional Review Board (IRB)-approved buffy coats from normal donors. CD14+ cells were isolated by two sequential positive selections with anti-human CD14 magnetic beads (Miltenyi). Purity was at least 99%. CD14+ cells were cultured in RPMI medium, 10% FBS, antibiotics and HEPES in the presence of recombinant human GM-CSF at 10 ng ml−1 and IL-4 at 50 ng ml−1 (eBioscience). Fresh media was added at day 3, and cells were stimulated or infected at day 4. To isolate CD11c+ dendritic cells, CD14-depleted peripheral blood mononuclear cells were further depleted using biotin-labelled antibodies against CD3, CD16, CD19 and CD56 and streptavidin magnetic beads (Miltenyi). The negative fraction was stained and sorted on a FACSAria (BD Biosciences) as CD3−CD14−CD16−CD19−CD20−CD56−HLA-DR+CD11c+ (Supplementary Table 2). Purity was at least 98%. Total CD4+ T cells were isolated using human CD4 magnetic beads (Miltenyi). Naive CD4+ T cells were further sorted as CD4+CD25−CD45RA+CD45RO−.
T-cell clones
T-cell clones were expanded as previously described29. The clone DR4LI15 (LGLNKIVRMYSPTSI) was obtained from N. Bhardwaj and the clones B81TL9 (TPQDLNTML) and B14DA9 (DRFYKTLRA) were obtained from B. Walker.
Reagents
LPS and poly(I:C) were from Sigma and were used at 1 µg ml−1 and 10 µg ml−1, respectively. Curdlan (CM-Curdlan) was form Wako and was used at 1 µg ml−1. Cyclosporin A and FK506 were from Calbiochem. AZT, raltegravir, lopinavir, saquinavir and tipranavir were obtained through the NIH AIDS Research & Reference Reagent Program. SCY (SCYX00011867717) is similar to SCY-635 and was a gift from Scynexis.
Infection and stimulation
At day 4 of MDDC differentiation, cells were harvested, counted and resuspended in their own media at a concentration of one million per ml with 5 µg ml−1 polybrene, and 100 µl was aliquoted in round-bottomed 96-well plates. For infection, 50 µl of media or SIV-VLP(G) was first added. One-hundred microlitres of media or dilutions of various HIV-1-derived viral preparations were then added. Saquinavir, tripranavir, AZT and raltegravir were added at 10 µM. Neutralizing anti-IFN-α and anti-IFN-β were added at 20 µg ml−1. B18R (eBioscience) was added at 0.2 µg ml−1. IL28RA-Fc (R&D Systems) was added at 1 µg ml−1. Neutralizing anti-IFNAR was added at 1 µg ml−1. For shRNA-transduced dendritic cells, cells were harvested, counted and resuspended in fresh media containing GM-CSF, IL-4 and 1 µg ml−1 polybrene at a concentration of one million per ml. One-hundred microlitres was aliquoted in round-bottomed 96-well plates and 100 µl of media or virus was added.
Western blot analysis
Cells were lysed in 1% NP-40, 50 mM Tris pH 8, 120 mM NaCl, 4 mM EDTA, 50 mM NaF, 1 mM NA3VO4 and a protease inhibitor cocktail (Roche). Total lysates were resolved on SDS–PAGE, transferred to polyvinylidene fluoride (PVDF) membranes and probed with primary antibodies and corresponding horseradish peroxidase (HRP)-conjugated secondary antibodies (GE Healthcare).
Cytoplasmic and nuclear fractionation
MDDCs were harvested at 24 h after infection or treatment. 4 × 106 cells were washed once with room temperature PBS, gently pelleted and resuspended in 400 µl of cold cytoplasmic lysis buffer (CL buffer) containing 10 mM HEPES pH 7.9, 10 mM sodium potassium, 1.5 mM magnesium chloride, 1 mM sodium orthovanadate, 2 mM sodium pyrophosphate, 2 mM sodium β-glycerophosphate, 5 mM sodium fluoride, complete EDTA-free protease inhibitor cocktail (Roche) and phosphatase inhibitor cocktail (SIGMA P2850). Cells in cold CL buffer were immediately pelleted at 4 °C, the supernatant was discarded, 40 µl CL was added, and buffer and cells were gently resuspended by slow pipetting and soft flicking and left on ice for 15 min. Two and a half microlitres of 10% NP-40 was added and cells were lysed by gentle flicking. Nuclei were pelleted at 15,900g for 5 min at 4 °C. Forty microlitres of supernatant was harvested and saved as the cytoplasmic fraction, and remaining liquid was discarded. Forty microlitres of cold nuclear lysis buffer (NL buffer) containing 420 mM sodium chloride, 20 mM HEPES pH 7.9, 1.5 mM magnesium chloride, 0.2 mM EDTA, 25% glycerol and protease and phosphatase inhibitors as in CL buffer was added. Nuclei were resuspended by vigorous flicking and incubated on ice for 15 min, with occasional flicking. Nuclei were vortexed for 10 s and sonicated for 10 min in a 4 °C bath sonicator (30 s on, 30 s off). The nuclear lysate was cleared by centrifugation at 15,900g for 5 min at 4 °C, and the resulting supernatant was saved as the nuclear extract. Western blot loading buffer with dithiothreitol was added to the cytoplasmic and nuclear extracts, and the samples were heated at 70 °C for 15 min. Ten microlitres of each sample was run on a 7.5% SDS–PAGE gel and transferred to PVDF membrane (Roche). Membranes were blocked with 5% non-fat dry milk in Tris-buffered saline (TBS) containing 0.1% Tween-20 (TBST) and probed with primary antibody overnight while rocking at 4 °C, washed six times for 5 min with TBST, probed with secondary HRP-conjugated antibody (GE Healthcare) for one hour at 19 °C (room temperature), washed six times for 5 min in TBST, and incubated with ECL reagents (Pierce Pico or Pierce Femto). Chemiluminescence signal was visualized using Kodak film.
Quantitative bioassay for IFNs
293FT and THP-1 cells were infected with HIV-GFP(G) and SIV-VLP(G) or transfected with poly(I:C) or total RNA from NDV-infected A549 cells harvested in Trizol (Invitrogen) 8 h after infection using lipofectamine 2000 (Invitrogen). NDV viral stock was produced by inoculating 10-day-old embryonated chicken eggs (Charles River). CD4+ T cells were expanded with 5 µg ml−1 phytohemagglutinin-L (Sigma) and 10 U ml−1 human IL-2 for 4 days and infected with 100 haemagglutinin (HA) units ml−1 of Sendai virus (Charles River) or infected with HIV-GFP(G) and SIV-VLP(G). Media were replaced after 24 h and culture supernatants were harvested after another 24 h. Cell culture supernatants were ultraviolet-irradiated to inactivate traces of Sendai virus. Supernatants were assayed for IFN activity using a recombinant COS-1 cell line, which carries a luciferase reporter containing multiple repeats of IFN-stimulated response element (ISRE). In brief, the reporter cells were exposed to cell culture supernatants for 8 h to overnight and assayed for luciferase activities, which were then translated to IFN activities by using a standard curve generated from a serial dilution of human IFNα-2a.
Microarray analysis
MDDCs were infected with HIV-GFP(G), SIV-VLP(G), both, or treated with LPS. Cells were harvested after 48 h and a subset was analysed by flow cytometry. RNA was prepared with TRIZOL and microarray data generation was done using standard protocols on Human Genome U133A 2.0 arrays (Affymetrix). Microarray analysis was performed using the Bioconductor package in R and Genespring GX10 (Agilent). Probes were filtered based on at least a twofold change in expression and P < 0.05. Promoter analysis was performed using PRIMA30 in EXPANDER31. TLR pathway analysis was performed using SPIKE32.
Quantitative PCR
Quantitative PCR analysis was performed as described33 using the standard curve method or the ΔCT method (for primer sets see Supplementary Table 3).
Plasmids
HIV-GFP, which is env−vpu−vpr−vif−nef− , with the GFP open reading frame in place of nef, has already been described34. HIV-GFP ΔRev was generated by mutating the start codon of rev. HIV-GFP PTAP− was generated by mutating the PTAP motif in p6 to LIRL. HIV-GFP ΔGag was generated by inserting a stop codon after seven amino acids of Gag. Vpx–Vpr fusion protein was generated by fusing SIVmac251 Vpx with HIV-1 NL4-3 Vpr using the linker ANYAAAAAAADPS in pIRES2-EGFP (Clontech). LKO1gfp was generated by replacing the puroR open-reading frame in pLKO1puro35 with the EGFP coding region. shRNAs were designed as described previously (Supplementary Table 4), except that a partial mir30 sequence CTGTGAAGCCACAGATGGG was used for the loop. shRNAs were then cloned as described35. T54A/N57A, Q63A/Q67A, G89V and the parental vector pLaiΔEnv-GFP3 are env–nef– , with GFP in place of Nef, and were previously described12. HDV-IRES-RFP is described elsewhere36. HIV-2 ROD9 Δenv GFP was generated from an HIV-2 ΔEnv construct37 by inserting the GFP-coding sequence in Nef, thus disrupting Nef. All plasmid DNA was prepared with Invitrogen HiPure plasmid kit. Plasmid DNA did not induce dendritic cell maturation, and viral-producing cells were washed after DNA transfection.
Virus production
Viral particles were produced by transfection of 293FT cells with 3 µg DNA and 8 µl TransIT-293 (Mirus Bio); for shRNA vectors, we used 0.4 µg CMV-VSVG, 1 µg pCMV-ΔR8.91 and 1.6 µg shRNA; for SIV-VLP(G), 0.4 µg CMV-VSVG and 2.6 µg pSIV3+ (ref. 38); for HIV-GFP(G), 0.4 µg CMV-VSVG and 2.6 µg HIV-GFP; for HIV2 ROD9 Δenv GFP(G), 0.4 µg CMV-VSVG and 2.6 µg HIV2 ROD9 ΔEnv GFP; for NL4-3-ΔE-EGFP, 0.4 µg pCMV-VSVG and 2.6 µg pNL4-3-ΔE-EGFP39. HIV-GFP ΔRev and HIV–GFP PTAP− were produced with 0.4 µg CMV-VSVG, 0.5 µg pCMV-ΔR8.91 and 2.1 µg HIV plasmid. HIV-GFP ΔGag and HIV-1 capsid mutants were produced with 0.4 µg CMV-VSVG, 1 µg pCMV-ΔR8.91 and 1.6 µg HIV plasmid. R5-GFP is NL4-3 encoding for the BAL envelope and GFP in Nef34,36. One day after transfection, media was removed, cells were washed out once and fresh media was added. Viral supernatants were harvested one day later and filtered at 0.45 µM. In some experiments, p24 concentration was measured by p24 enzyme-linked immunosorbent assay (ELISA).
RNAi
Synthetic short interfering RNA can be delivered in MDDCs by electroporation, but this is highly and rapidly toxic and renders difficult the interpretation of dendritic cell activation, which can be altered by the presence of apoptotic or necrotic cells. In addition, we have not been able to achieve significant knockdown using synthetic siRNA with lipid-based reagents in MDDCs. In fact, fluorescently labelled siRNA appeared to be simply endocytosed with all the lipid-based reagents that we tested (data not shown). We thus used shRNA vectors. Using this method, we routinely obtained >90% transduction efficiency, alleviating the need for cell sorting or selection. Five-million freshly isolated CD14+ cells were cultured in 5 ml of media containing GM-CSF, IL-4, and 1 µg ml−1 polybrene. One millilitre of SIV-VLP(G) supernatant and 2.5 ml of shRNA vector supernatant were added to cells. At days 1 and 3, 2 ml of fresh media was added. At day 4, cells were transduced at more than 90% based on GFP expression and were used for further infections and stimulation as above.
HIV-specific T-cell-clone stimulation
Forty-eight hours after infection of MDDCs, 105 rested HIV-specific T-cell clones were added in the presence of GolgiStop (BD Biosciences). Where indicated, T cells were activated with 50 ng ml−1 PMA (Sigma) and 0.5 µg ml−1 ionomycin (Sigma). Cells were incubated for 6 h and processed for intracellular staining33.
Naive T-cell proliferation assay
Forty-eight hours after infection, naive T cells were labelled with CFSE (eBioscience) as descried previously18. Dendritic cells were infected with dilutions of SIV-VLP(G) and pLaiΔEnv-GFP3(G) wild type (indicated as HIV-GFP(G)) or G89V. AZT was added at the time of infection and SCY was added from 3 h to 8 h after infection. Forty-eight hours after infection, half of the dendritic cells were processed for surface staining and cytometry. The other half were washed with media and resuspended in fresh media without cytokines. Twenty-thousand T cells were mixed with dendritic cells at a dendritic cell to T cell ratio of 1:5 and 1:15. Cells were stimulated by dilutions of anti-CD3 (OKT3 hybridoma supernatant, approximately 1–100 ng ml−1) in a total volume of 150–200 µl in round-bottomed 96-well plates. Cells were harvested and analysed by flow cytometry at day 4 or day 5 after activation.
Trans-enhancement
105 day 4 MDDCs were infected with dilutions of HDV-IRES-RFP(G) and SIV-VLP(G) in 96-well round-bottomed plates in the presence of 5 µg ml−1 polybrene. Type-I-IFN-neutralizing antibodies and recombinant proteins were maintained throughout the experiment in some samples. Media was replaced after 24 h. Another 24 h later, half of the cells were processed for surface staining and RFP and CD86 expression were measured by flow cytometry. The other half was mixed with a preparation of replication- competent R5–GFP and 5 × 105 CD4+ T cells at day 4 after activation with PHA-L (Sigma) and IL-2, in the absence of polybrene. GFP expression in CD4+ T cells was measured another 48 h later.
References
Steinman, R. M. & Hemmi, H. Dendritic cells: translating innate to adaptive immunity. Curr. Top. Microbiol. Immunol. 311, 17–58 (2006)
Takeuchi, O. & Akira, S. Innate immunity to virus infection. Immunol. Rev. 227, 75–86 (2009)
Stetson, D. B., Ko, J. S., Heidmann, T. & Medzhitov, R. Trex1 prevents cell-intrinsic initiation of autoimmunity. Cell 134, 587–598 (2008)
Nègre, D. et al. Characterization of novel safe lentiviral vectors derived from simian immunodeficiency virus (SIVmac251) that efficiently transduce mature human dendritic cells. Gene Ther. 7, 1613–1623 (2000)
Cameron, P. U. et al. Dendritic cells exposed to human immunodeficiency virus type-1 transmit a vigorous cytopathic infection to CD4+ T cells. Science 257, 383–387 (1992)
Kwon, D. S., Gregorio, G., Bitton, N., Hendrickson, W. A. & Littman, D. R. DC-SIGN-mediated internalization of HIV is required for trans-enhancement of T cell infection. Immunity 16, 135–144 (2002)
Mangeot, P. E. et al. High levels of transduction of human dendritic cells with optimized SIV vectors. Mol. Ther. 5, 283–290 (2002)
Goujon, C. et al. With a little help from a friend: increasing HIV transduction of monocyte-derived dendritic cells with virion-like particles of SIV(MAC). Gene Ther. 13, 991–994 (2006)
Honda, K. et al. Selective contribution of IFN-α/β signaling to the maturation of dendritic cells induced by double-stranded RNA or viral infection. Proc. Natl Acad. Sci. USA 100, 10872–10877 (2003)
Malim, M. H., Hauber, J., Fenrick, R. & Cullen, B. R. Immunodeficiency virus rev trans-activator modulates the expression of the viral regulatory genes. Nature 335, 181–183 (1988)
Demirov, D. G., Orenstein, J. M. & Freed, E. O. The late domain of human immunodeficiency virus type 1 p6 promotes virus release in a cell type-dependent manner. J. Virol. 76, 105–117 (2002)
Yamashita, M., Perez, O., Hope, T. J. & Emerman, M. Evidence for direct involvement of the capsid protein in HIV infection of nondividing cells. PLoS Pathog. 3, e156 (2007)
Yoo, S. et al. Molecular recognition in the HIV-1 capsid/cyclophilin A complex. J. Mol. Biol. 269, 780–795 (1997)
Luban, J., Bossolt, K. L., Franke, E. K., Kalpana, G. V. & Goff, S. P. Human immunodeficiency virus type 1 Gag protein binds to cyclophilins A and B. Cell 73, 1067–1078 (1993)
Boggiano, C., Manel, N. & Littman, D. R. Dendritic cell-mediated trans-enhancement of human immunodeficiency virus type 1 infectivity is independent of DC-SIGN. J. Virol. 81, 2519–2523 (2007)
Sato, M. et al. Distinct and essential roles of transcription factors IRF-3 and IRF-7 in response to viruses for IFN-α/β gene induction. Immunity 13, 539–548 (2000)
Moris, A. et al. Dendritic cells and HIV-specific CD4+ T cells: HIV antigen presentation, T-cell activation, and viral transfer. Blood 108, 1643–1651 (2006)
Antons, A. K., Wang, R., Kalams, S. A. & Unutmaz, D. Suppression of HIV-specific and allogeneic T cell activation by human regulatory T cells is dependent on the strength of signals. PLoS ONE 3, e2952 (2008)
Gett, A. V., Sallusto, F., Lanzavecchia, A. & Geginat, J. T cell fitness determined by signal strength. Nature Immunol. 4, 355–360 (2003)
Langenkamp, A. et al. T cell priming by dendritic cells: thresholds for proliferation, differentiation and death and intraclonal functional diversification. Eur. J. Immunol. 32, 2046–2054 (2002)
Hopkins, S. et al. SCY-635, a novel nonimmunosuppressive analog of cyclosporine that exhibits potent inhibition of hepatitis C virus RNA replication in vitro . Antimicrob. Agents Chemother. 54, 660–672 (2010)
Chatterji, U. et al. The isomerase active site of cyclophilin A is critical for hepatitis C virus replication. J. Biol. Chem. 284, 16998–17005 (2009)
de Silva, T. I., Cotten, M. & Rowland-Jones, S. L. HIV-2: the forgotten AIDS virus. Trends Microbiol. 16, 588–595 (2008)
Neagu, M. R. et al. Potent inhibition of HIV-1 by TRIM5-cyclophilin fusion proteins engineered from human components. J. Clin. Invest. 119, 3035–3047 (2009)
Price, A. J. et al. Active site remodeling switches HIV specificity of antiretroviral TRIMCyp. Nature Struct. Mol. Biol. 16, 1036–1042 (2009)
Janeway, C. A., Jr Approaching the asymptote? Evolution and revolution in immunology. Cold Spring Harb. Symp. Quant. Biol. 54, 1–13 (1989)
Kosmrlj, A. et al. Effects of thymic selection of the T-cell repertoire on HLA class I-associated control of HIV infection. 465, 350–354 Nature . (2010)
Morner, A. et al. Primary human immunodeficiency virus type 2 (HIV-2) isolates, like HIV-1 isolates, frequently use CCR5 but show promiscuity in coreceptor usage. J. Virol. 73, 2343–2349 (1999)
Fonteneau, J. F. et al. Generation of high quantities of viral and tumor-specific human CD4+ and CD8+ T-cell clones using peptide pulsed mature dendritic cells. J. Immunol. Methods 258, 111–126 (2001)
Elkon, R., Linhart, C., Sharan, R., Shamir, R. & Shiloh, Y. Genome-wide in silico identification of transcriptional regulators controlling the cell cycle in human cells. Genome Res. 13, 773–780 (2003)
Shamir, R. et al. EXPANDER—an integrative program suite for microarray data analysis. BMC Bioinformatics 6, 232 (2005)
Elkon, R. et al. SPIKE—a database, visualization and analysis tool of cellular signaling pathways. BMC Bioinformatics 9, 110 (2008)
Manel, N., Unutmaz, D. & Littman, D. R. The differentiation of human TH-17 cells requires transforming growth factor-β and induction of the nuclear receptor RORγt. Nature Immunol. 9, 641–649 (2008)
Unutmaz, D., KewalRamani, V. N., Marmon, S. & Littman, D. R. Cytokine signals are sufficient for HIV-1 infection of resting human T lymphocytes. J. Exp. Med. 189, 1735–1746 (1999)
Moffat, J. et al. A lentiviral RNAi library for human and mouse genes applied to an arrayed viral high-content screen. Cell 124, 1283–1298 (2006)
Oswald-Richter, K. et al. HIV infection of naturally occurring and genetically reprogrammed human regulatory T-cells. PLoS Biol. 2, e198 (2004)
Griffin, S. D., Allen, J. F. & Lever, A. M. The major human immunodeficiency virus type 2 (HIV-2) packaging signal is present on all HIV-2 RNA species: cotranslational RNA encapsidation and limitation of Gag protein confer specificity. J. Virol. 75, 12058–12069 (2001)
Mangeot, P. E. et al. Development of minimal lentivirus vectors derived from simian immunodeficiency virus (SIVmac251) and their use for gene transfer into human dendritic cells. J. Virol. 74, 8307–8315 (2000)
Zhang, H. et al. Novel single-cell-level phenotypic assay for residual drug susceptibility and reduced replication capacity of drug-resistant human immunodeficirency virus type 1. J. Virol. 78, 1718–1729 (2004)
Acknowledgements
We thank members of the Littman laboratory for valuable discussions and critical reading of the manuscript, and T. Egawa, E. Kurz, X. Gong and L. Kozyhaya for assistance with experiments. We thank S. Hopkins at Scynexis for the gift of the SCY compound, J. Zavadil and the genomic core facility of New York University Medical Center for performing the array studies, M. Emerman, M. Yamashita and W. Sundquist for HIV constructs and critical reading of the manuscript, B. Walker, A. Piechocka-Trocha, D. Kwon, S. M. Vine, N. Bardhwaj and E. Miller for T-cell clones and reagents, and S. Schwab and R. Medzhitov for critical reading of the manuscript. N.M. thanks M. Sitbon, N. Taylor and J.-M. Blanchard for continuous support. The work was supported sequentially by EMBO and Cancer Research Institute fellowships (N.M.), by the Institut National de la Santé et de la Recherche Médicale (N.M.), by the Howard Hughes Medical Institute (D.R.L.), the Helen and Martin Kimmel Center for Biology and Medicine (D.R.L.), and National Institutes of Health (NIH) grants AI33856 (D.R.L.), AI28900 (D.E.L.), U54AI57168 (D.E.L.) and R01AI065303 (D.U.).
Author information
Authors and Affiliations
Contributions
N.M. and D.R.L. designed the study and wrote the manuscript. N.M. performed the experiments and analyses. B.H. provided technical help. D.U. provided expertise and contributed to experiments with human T-cell proliferation assays. D.E.L. provided expertise in identifying the IFN response. D.E.L. and Y.W. designed the quantitative bioassay for IFNs and Y.W. performed the assay. All authors discussed results and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1- 4 and Supplementary Figures 1 – 18 with legends. (PDF 2086 kb)
Rights and permissions
About this article
Cite this article
Manel, N., Hogstad, B., Wang, Y. et al. A cryptic sensor for HIV-1 activates antiviral innate immunity in dendritic cells. Nature 467, 214–217 (2010). https://doi.org/10.1038/nature09337
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature09337
This article is cited by
-
Early antiretroviral therapy favors post-treatment SIV control associated with the expansion of enhanced memory CD8+ T-cells
Nature Communications (2024)
-
The Hippo signaling component LATS2 enhances innate immunity to inhibit HIV-1 infection through PQBP1-cGAS pathway
Cell Death & Differentiation (2022)
-
HIV-1 capsid variability: viral exploitation and evasion of capsid-binding molecules
Retrovirology (2021)
-
Vpx enhances innate immune responses independently of SAMHD1 during HIV-1 infection
Retrovirology (2021)
-
SUMOylation of SAMHD1 at Lysine 595 is required for HIV-1 restriction in non-cycling cells
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.