Abstract
Fibrolamellar hepatocellular carcinoma is a rare subtype of hepatocellular carcinoma with distinct clinical and histological features, and better survival compared with conventional hepatocellular carcinoma in some but not all series. We performed a comparative genomic hybridization analysis on 11 fibrolamellar carcinomas and correlated the findings with clinicopathologic features and survival. Chromosomal imbalances were identified in six cases (55%), whereas the other five (45%) yielded normal results. The mean number of aberrations per case was 3.9 for all cases and 7.2 in abnormal cases. Among the six abnormal cases, gains or losses were observed at 3 loci in two cases, 7 loci in one case, 8 loci in two cases and 14 loci in one case. The most common abnormalities were observed in chromosomes 7, 8 and 18, with 7q gain in five cases and 7p gain in four cases. Aberrations associated with intermediate or advanced conventional hepatocellular carcinomas, including losses at 3q, 4q and 13q were identified in 17–33% of fibrolamellar carcinomas. There was no correlation of chromosomal changes with age, gender and tumor size. The 5-year survival among the six patients with no chromosomal abnormalities was 80% (4/5) compared with 33% (2/6) in patients with chromosomal abnormalities (P=0.1). In conclusion, fibrolamellar carcinomas show fewer chromosomal abnormalities compared with those reported in literature for conventional hepatocellular carcinoma. The most common abnormalities occur in chromosomes 7 and 8. Fibrolamellar carcinomas with chromosomal changes appear to behave more aggressively compared with cases with no cytogenetic abnormalities. The favorable outcome in some fibrolamellar carcinomas may be due to absent or low number of cytogenetic aberrations.
Similar content being viewed by others
Main
Fibrolamellar hepatocellular carcinoma is a rare subtype of hepatocellular carcinoma with distinct clinical and pathologic features. The tumor is typically associated with young age, absence of cirrhosis, normal α-fetoprotein levels and absence of risk factors1, 2, 3, 4, 5, 6). Histologically, it is characterized by lamellar pattern of fibrosis, abundant granular cytoplasm and prominent nucleoli.1, 2, 4, 5, 7 Several studies have reported a favorable outcome in fibrolamellar carcinoma compared with conventional hepatocellular carcinoma.1, 3, 4, 5, 8, 9, 10, 11 However, other studies have shown that fibrolamellar carcinoma is an aggressive neoplasm with 5-year survival of around 50%.2, 6, 12, 13, 14, 15 These studies have suggested that the prognosis of fibrolamellar carcinomas may be better than conventional hepatocellular carcinomas arising in cirrhosis, but is similar to hepatocellular carcinoma arising in noncirrhotic liver.12, 15 The reason for variable outcomes in fibrolamellar carcinoma is not known.
Cytogenetic studies such as comparative genomic hybridization (CGH) and fluorescence in situ hybridization have demonstrated characteristic chromosomal aberrations in conventional hepatocellular carcinomas.16, 17, 18, 19, 20, 21, 22, 23, 24, 25 The earliest changes are gains at chromosomal arms 1q and 8q.24, 25 Other common abnormalities that occur during tumor progression are gains at 6q,7q, 20q and X, and losses at 4q, 8p, 13q, 16q and 17p.16, 17, 18, 19, 20, 21, 22, 23, 24, 25 Some of these chromosomal changes show distinct clinicopathologic associations. Elevated α-fetoprotein levels and p53 mutations correlate with loss of 4q.16 Gains of 8q and 20q have been observed in large tumors.17, 20 Hepatocellular carcinomas arising in noncirrhotic liver often show gain of 8q and loss of 13q.17, 18 Losses of 3q, 9p and 6q may be independent predictors of unfavorable outcome.19, 25 The number of cytogenetic aberrations also correlates with tumor differentiation and outcome.25
Unlike conventional hepatocellular carcinoma, cytogenetic changes and their relationship to outcome have not been widely studied in fibrolamellar carcinomas. A few case reports and two small series26, 27, 28, 29, 30 have examined chromosomal changes in fibrolamellar carcinomas, and it has been suggested that the better prognosis of fibrolamellar carcinomas compared with conventional hepatocellular carcinoma may be related to lower number of aberrations in fibrolamellar carcinoma. This study delineates the chromosomal changes in fibrolamellar carcinoma using CGH, and examines their relationship with clinicopathologic features and survival.
Materials and methods
Cases
The study group composed of 11 resected cases of fibrolamellar carcinomas. The slides were reviewed to confirm the diagnosis. All cases occurred in noncirrhotic liver and were characterized by large tumor cells with abundant granular cytoplasm, prominent nucleoli and lamellar pattern of fibrosis (Figure 1). The age, gender and tumor size were obtained from the pathology report. Clinical and pathologic records were reviewed to determine lymph node and distant metastasis. Information on recurrences and 5-year survival was obtained from the tumor registry. A minimum follow-up period of 5 years after initial surgery was available for all living patients. The institutional review board approved the study.
Array-Based CGH
The assay was performed using DNA extracted from formalin-fixed paraffin-embedded tissue using human 1.14 arrays obtained from the University of California at San Francisco Cancer Center Array Core.
DNA Extraction and Quantification
DNA from the tumor was extracted from 10 μm unbaked paraffin sections. A desired area of the tumor was selected on H&E-stained slides and the tissue was scraped, dissolved in PCR/proteinase K buffer and incubated overnight at 55°C Fresh proteinase K was added to the tubes at 24 and 48 h and the tissues was incubated for a total of 72 h. Proteinase K was inactivated at 95°C and genomic DNA was purified and concentrated using Amicon Microcon YM-30 columns to remove contaminating substances and small fragments. To assure consistency in the array hybridization, genomic DNA extracted was quantified using Taqman analysis using a CA-repeat fluorogenic probe as described.31
CGH Arrays
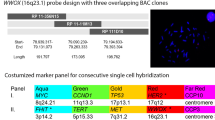
The arrays used in the study were prepared and hybridized as described previously.32 In brief, human 1.14 arrays were obtained from the UCSF Cancer Center Array Core (http://cc.ucsf.edu/microarray/). The arrays consisted of 2433 bacterial artificial chromosome (BAC) clones that covered the human genome at 1.5 Mb resolution. For hybridization, 1 μg of tumor DNA and 1 μg of gender-matched reference DNA (isolated from normal donor lymphocytes) was labeled by random priming using Cy3-dCTP and Cy5-dCTP, respectively, using Bioprime Kit (Invitrogen, Carlsbad, CA, USA). Unincorporated fluorescent nucleotides were removed using a Sephadex G-50 column (Amersham, Piscataway, NJ, USA). Sample and reference DNA were mixed with 100 μg Cot-1, precipitated and resuspended in hybridization solution. The hybridization solution was denatured for 10 min at 72°C before being incubated for 1 h at 37°C to allow blocking of repetitive sequences. Hybridization was performed for 72 h in a moist chamber on a slow rocking table. The arrays were washed for 10 min in 50% formamide and 2 × SSC at 45°C, and 10 min in phosphate buffer at room temperature. Slides were mounted in mounting solutions containing 0.3 μg/ml DAPI. Three single-color intensity images (DAPI, Cy3 and Cy5) were collected for each array using a charge-coupled device camera.
CGH Data Analysis
The UCSF SPOT software (http://jainlab.ucsf.edu/Downloads.html) was used to automatically segment the spots based on the DAPI images, perform local background correction and calculate various measurement parameters, including log 2 ratios of the total integrated Cy3 and Cy5 intensities for each spot.33 A second custom program SPROC (http://jainlab.ucsf.edu/Downloads.html) was used to associate clone identities and a mapping information file with each spot, so that the data could be plotted relative to the position of the BACs. Chromosomal aberrations were classified as a gain when the normalized log 2 Cy3/Cy5 ratio was >0.225 and as a loss when the ratio was <−0.225. This number was determined as threefold of the average s.d. of normal vs normal array CGH hybridization. Steep copy number changes with the graph showing a peak rather than a plateau, and a minimal normalized log 2 Cy3/Cy5 ratio of 0.9 and higher were classified as amplifications. Likewise, log 2 Cy3/Cy5 ratio of −0.8 and lower were classified as homozygous deletions. Multiple gains, losses and amplifications were counted as separate events. The threshold of gain or loss of an entire chromosome arm was defined as the median log 2 ratio of >0.225 or <−0.225 for all clones on the chromosome arm.
Statistical Analysis
The relationship of chromosomal abnormalities and clinicopathologic features was examined by Fisher exact test. For survival analysis, the starting point for the survival time was the date of surgery. Survival curves were calculated using the Kaplan–Meier method and statistical significance between curves was tested by the log-rank test. Multivariate analysis was performed using Cox proportional-hazards model; the hazard ratio and its 95% confidence intervals were assessed for each factor. P-values of <0.05 were considered statistically significant.
Results
Clinical Characteristics
Of the 11 cases of fibrolamellar carcinoma, 6 (55%) occurred in males. The mean age was 25.4 years (range 16–48 years). The age and gender were not known in one case. The mean tumor size was 14.3 cm (range 7.2–23 cm). All cases were unicentric primary liver tumors. The tumor was confined to the liver at diagnosis in seven cases, whereas metastatic disease was present in four cases. The overall 5-year survival was 55%. The 5-year survival in patients with tumor limited to the liver was 71% compared with 25% in patients with extrahepatic disease (P=0.1). Age, gender and tumor size did not correlate with survival.
Cytogenetic Aberrations
CGH revealed chromosomal imbalances in six cases (55%), whereas the other five (45%) yielded normal results. The mean number of aberrations was 3.6 per case when all cases were considered, and 7.2 in abnormal cases. Among the six abnormal cases, gains or losses were observed at 3 loci in two cases, 7 loci in one case, 8 loci in two cases and 14 loci in one case (Figure 2; Tables 1 and 2).
Chromosomal gains and losses in fibrolamellar carcinomas (FLM) and hepatocellular carcinomas (HCC). The HCC data (from Patil et al34) is shown for comparison.
Chromosome 7 abnormalities were most common with 7q gain in five cases and 7p gain in four cases. Abnormalities in chromosome 8 were seen in three cases and included 8q gain (two cases), 8p loss (two cases) and 8q loss (two cases). Abnormalities in chromosome 1 were seen in two cases (1p loss in one case and 1q gain in one case). Other recurrent abnormalities included gains at 19p (three cases), and losses at 18q (three cases), 4q, 13q, 14q and 21q (two cases each).
Cytogenetic Changes, Clinicopathologic Features and Survival
There was no correlation of CGH changes with age, gender and tumor size. The 5-year survival among the five patients with no CGH abnormalities was 80% (4/5) compared with 33% (2/6) in patients with chromosomal abnormalities (P=0.1; Figure 3) (Table 3).
Discussion
Chromosomal abnormalities in conventional hepatocellular carcinomas have been well characterized in several studies. Gains at 1q and 8q are the earliest changes in hepatocarcinogenesis and have been observed in 46–78% and 48–88% of conventional HCC, respectively.24, 25 Other common changes include gains of 6p (26–32%), 7q (32–40%), 17q (29–46%), 20q (25–37%) and Xq (25–50%), and losses of 4q (32–50%), 8p (29–58%), 13q (16–39%), 16q (30–63%) and 17p (31–60%).16, 17, 18, 19, 20, 21, 22, 23, 24, 25 Despite the differences in clinical and epidemiological features between fibrolamellar carcinoma and conventional hepatocellular carcinoma, this study and the limited literature on this subject indicate that conventional hepatocellular carcinomas and fibrolamellar carcinomas share many cytogenetic aberrations.
We observed chromosomal changes in 55% of fibrolamellar carcinomas, with abnormalities in chromosomes 7, 8 and 18 being the most common. Gains of chromosome 7p were observed in 33% of total cases and 67% of abnormal cases, whereas 7q gains were present in 25% of total cases and 50% of abnormal cases. This is the location of hepatocyte growth factor and its receptor (MET), and is similarly involved in conventional hepatocellular carcinoma where it has been associated with increased intrahepatic metastasis.21, 35 Chromosome 7 gain in a recurrent fibrolamellar carcinoma was also noted in a case report.26 However, none of the fibrolamellar carcinomas showed chromosome 7 changes in another series.28 Gains of 7p are less common in conventional hepatocellular carcinoma.36, 37 The epidermal growth factor receptor gene is located on 7p and may be involved in fibrolamellar carcinoma. Overexpression and increase in gene copy number of epidermal growth factor receptor has been shown to occur in nearly all cases of fibrolamellar carcinoma.38 The other common abnormality in our series was gain of 18q, which occurred in 25% of total cases and 50% of cases with cytogenetic changes. 18q losses suggest involvement of DPC4 and DCC genes that are typically involved in pancreatic and colorectal adenocarcinomas, respectively.39, 40 18q abnormalities are less common in conventional hepatocellular carcinoma. Hence, even though there is overlap in the chromosomal changes in fibrolamellar carcinoma and conventional hepatocellular carcinoma, some abnormalities such as gains on 7p and 18q may be more characteristic of fibrolamellar carcinoma.
Gains of 1q and 8q, the earliest abnormalities in conventional hepatocellular carcinoma, are seen in some but not the majority of fibrolamellar carcinomas. Marchio et al28 found gains of 1q in 60% of fibrolamellar carcinomas, but it occurred only in one case in our series. The reason for this discrepancy is not clear. Because fibrolamellar carcinoma typically occurs in the young and has better outcome compared with hepatocellular carcinoma arising in cirrhosis, it is possible that cytogenetic changes in fibrolamellar carcinoma are similar to early or well-differentiated conventional hepatocellular carcinoma. However, many of the chromosomal abnormalities seen in advanced conventional HCC are commonly present in fibrolamellar carcinoma. Loss of 3q is associated with poor survival in conventional hepatocellular carcinoma,25 and was identified in one-third of the cytogenetically abnormal fibrolamellar carcinomas in our series and also in two case reports.27, 28 Gains of 13q and 20q, and loss of 4q are considered intermediate to late steps in hepatocarcinogenesis20, 24, 25, 36 and occurred in 33, 16 and 33% of cytogenetically abnormal fibrolamellar carcinomas respectively in our study. Loss of 13q was also observed in 29% of cytogenetically abnormal fibrolamellar carcinomas by Marcio et al as well as in a case report.28, 30 Similarly, 20q gain has been reported in one case each in two reports of fibrolamellar carcinoma.26, 28 Gains of 7q and Xq also occur late in the evolution of conventional hepatocellular carcinoma and are often associated large tumor size and advanced stage.25 These changes were seen in 50 and 16%, respectively, of cytogenetically abnormal cases in our series. Several case reports of fibrolamellar carcinoma have also reported these abnormalities.26, 27, 30 Hence, based on cytogenetic changes, fibrolamellar carcinoma does not appear to be equivalent to early or well-differentiated conventional HCC, but shows many of the changes associated with advanced or poorly differentiated conventional hepatocellular carcinomas. The aggressive behavior of some fibrolamellar carcinomas may be related to these chromosomal abnormalities. The number of cases in our study was insufficient to test the significance of individual abnormalities.
In contrast to conventional hepatocellular carcinomas, a significant number of fibrolamellar carcinoma cases do not show any cytogenetic abnormalities as detected by CGH. In our series, chromosomal abnormalities were identified in 55% of fibrolamellar carcinomas. Marchio et al reported cytogenetic aberrations in 70% of fibrolamellar carcinomas.28 In contrast, more than 90% of conventional hepatocellular carcinomas show chromosomal abnormalities by CGH.17, 21, 25, 28, 36 An overwhelming majority of well-differentiated conventional hepatocellular carcinomas also show chromosomal abnormalities.36, 37 Fibrolamellar carcinoma cases show lower mean number of changes per case compared with conventional hepatocellular carcinoma. In our study, the mean number of abnormalities per case was 3.6, which is similar to 3.2 in the series of Marchio et al.28 In contrast, the mean number of aberrations per case in conventional hepatocellular carcinoma has ranged between 5.4 and 12.8.17, 21, 36, 34 Even early conventional hepatocellular carcinomas show more aberrations (mean 6.5) than fibrolamellar carcinomas.25 Along with other features such as young age and absence of cirrhosis, the absence or lower number of aberrations compared with conventional hepatocellular carcinoma may account for the favorable clinical outcome in some cases of fibrolamellar carcinomas. It is possible that fibrolamellar carcinomas do not have major chromosomal aberrations in the initial stage of evolution and develop multiple gains and losses with tumor progression. Minimal or no chromosomal aberrations in the primary tumor and numerous aberrations in subsequent recurrence or metastatic fibrolamellar carcinomas have been reported.26, 27 Other explanations at the genetic level have also been advanced to explain the less aggressive behavior of some fibrolamellar carcinomas. The frequency of allelic loss evaluated by polymerase chain reaction has been shown to be lower in fibrolamellar carcinoma compared with conventional hepatocellular carcinoma.41 These tumors have also been shown to lack intratumor heterogeneity as detected by DNA fingerprinting techniques, which may contribute to the less aggressive behavior in some studies.42
Resectability of the tumor is the most important prognostic factor in fibrolamellar carcinoma.4 Other features that have been associated with higher survival include solitary tumors, free margin status, normal liver enzymes, low stage, and absence of thrombosis or vascular invasion.2, 6, 8, 43, 44, 45 Tumor size, gender, cellular proliferation and atypia does not affect prognosis.45 Young age has been reported as a favorable as well as unfavorable prognostic factor.43, 45 The influence of chromosomal abnormalities on survival in fibrolamellar carcinoma is not fully understood. In our study, there was a trend toward higher 5-year survival in fibrolamellar carcinoma with no cytogenetic aberrations (80 vs 33%, P=0.1) and with no extrahepatic disease (71 vs 25%, P=0.1). None of the other clinicopathologic features correlated with survival.
In conclusion, chromosomal abnormalities are present in approximately half of fibrolamellar carcinomas and are similar to conventional hepatocellular carcinoma. Abnormalities at some loci such as 7p and loss of 18q may be more common in fibrolamellar carcinoma. The mean number of aberrations is lower compared with conventional hepatocellular carcinoma. Fibrolamellar carcinomas showing chromosomal changes by array-based CGH appear to behave more aggressively compared with cases with no cytogenetic aberrations. There is no correlation of genetic aberrations with age, gender, tumor size and metastatic disease.
References
Craig JR, Peters RL, Edmondson HA, et al. Fibrolamellar carcinoma of the liver: a tumor of adolescents and young adults with distinctive clinico-pathologic features. Cancer 1980;46:372–379.
Ruffin MT . Fibrolamellar hepatoma. Am J Gastroenterol 1990;85:577–581.
El-Serag HB, Davila JA . Is fibrolamellar carcinoma different from hepatocellular carcinoma? A US population-based study. Hepatology 2004;39:798–803.
Hodgson HJ . Fibrolamellar cancer of the liver. J Hepatol 1987;5:241–247.
Berman MA, Burnham JA, Sheahan DG . Fibrolamellar carcinoma of the liver: an immunohistochemical study of nineteen cases and a review of the literature. Hum Pathol 1988;19:784–794.
Ringe B, Wittekind C, Weimann A, et al. Results of hepatic resection and transplantation for fibrolamellar carcinoma. Surg Gynecol Obstet 1992;175:299–305.
Burgart LJ, Martinez CJM, Batts KP . Fibrolamllar hepatoma-importance of using a strict definition (abstract). Mod Pathol 1994;7:129A.
Starzl TE, Iwatsuki S, Shaw Jr BW, et al. Treatment of fibrolamellar hepatoma with partial or total hepatectomy and transplantation of the liver. Surg Gynecol Obstet 1986;162:145–148.
Wood WJ, Rawlings M, Evans H, et al. Hepatocellular carcinoma: importance of histologic classification as a prognostic factor. Am J Surg 1988;155:663–666.
Okuda K . Natural history of hepatocellular carcinoma including fibrolamellar and hepato-cholangiocarcinoma variants. J Gastroenterol Hepatol 2002;17:401–405.
Pinna AD, Iwatsuki S, Lee RG, et al. Treatment of fibrolamellar hepatoma with subtotal hepatectomy or transplantation. Hepatology 1997;26:877–883.
Nagorney DM, Adson MA, Weiland LH, et al. Fibrolamellar hepatoma. Am J Surg 1985;149:113–119.
Haas JE, Muczynski KA, Krailo M, et al. Histopathology and prognosis in childhood hepatoblastoma and hepatocarcinoma. Cancer 1989;64:1082–1095.
Katzenstein HM, Krailo MD, Malogolowkin MH, et al. Fibrolamellar hepatocellular carcinoma in children and adolescents. Cancer 2003;97:2006–2012.
Kakar S, Burgart LJ, Batts KP, et al. Clinicopathologic features and survival in fibrolamellar carcinoma: comparison with conventional hepatocellular carcinoma with and without cirrhosis. Mod Pathol 2005;18:1417–1423.
Rashid A, Wang JS, Qian GS, et al. Genetic alterations in hepatocellular carcinomas: association between loss of chromosome 4q and p53 gene mutations. Br J Cancer 1999;80:59–66.
Wong N, Lai P, Lee SW, et al. Assessment of genetic changes in hepatocellular carcinoma by comparative genomic hybridization analysis: relationship to disease stage, tumor size, and cirrhosis. Am J Pathol 1999;154:37–43.
Zondervan PE, Wink J, Alers JC, et al. Molecular cytogenetic evaluation of virus-associated and non-viral hepatocellular carcinoma: analysis of 26 carcinomas and 12 concurrent dysplasias. J Pathol 2000;192:207–215.
Laurent-Puig P, Legoix P, Bluteau O, et al. Genetic alterations associated with hepatocellular carcinomas define distinct pathways of hepatocarcinogenesis. Gastroenterology 2001;120:1763–1773.
Guan XY, Fang Y, Sham J, et al. Recurrent chromosome alterations in hepatocellular carcinoma detected by comparative genomic hybridization. Genes Chromosomes Cancer 2001;30:110–116.
Balsara BR, Pei J, De Rienzo A, et al. Human hepatocellular carcinoma is characterized by a highly consistent pattern of genomic imbalances, including frequent loss of 16q23.1–24.1. Genes Chromosomes Cancer 2001;30:245–253.
Rao UN, Gollin SM, Beaves S, et al. Comparative genomic hybridization of hepatocellular carcinoma: correlation with fluorescence in situ hybridization in paraffin-embedded tissue. Mol Diagn 2001;6:27–37.
Wilkens L, Bredt M, Flemming P, et al. Diagnostic impact of fluorescence in situ hybridization in the differentiation of hepatocellular adenoma and well-differentiated hepatocellular carcinoma. J Mol Diagn 2001;3:68–73.
Moinzadeh P, Breuhahn K, Stützer H, et al. Chromosome alterations in human hepatocellular carcinomas correlate with aetiology and histological grade—results of an explorative CGH meta-analysis. Br J Cancer 2005;92:935–941.
Poon TC, Wong N, Lai PB, et al. A tumor progression model for hepatocellular carcinoma: bioinformatic analysis of genomic data. Gastroenterology 2006;131:1262–1270.
Lowichik A, Schneider NR, Tonk V, et al. Report of a complex karyotype in recurrent metastatic fibrolamellar hepatocellular carcinoma and a review of hepatocellular carcinoma cytogenetics. Cancer Genet Cytogenet 1996;88:170–174.
Wilkens L, Bredt M, Flemming P, et al. Cytogenetic aberrations in primary and recurrent fibrolamellar hepatocellular carcinoma detected by comparative genomic hybridization. Am J Clin Pathol 2000;114:867–874.
Marchio A, Pineau P, Meddeb M, et al. Distinct chromosomal abnormality pattern in primary liver cancer of non-B, non-C patients. Oncogene 2000;19:3733–3738.
Terracciano L, Tornillo L . Cytogenetic alterations in liver cell tumors as detected by comparative genomic hybridization. Pathologica 2003;95:71–82.
Terracciano LM, Tornillo L, Avoledo P, et al. Fibrolamellar hepatocellular carcinoma occurring 5 years after hepatocellular adenoma in a 14-year-old girl: a case report with comparative genomic hybridization analysis. Arch Pathol Lab Med 2004;128:222–226.
Ginzinger DG, Godfrey TE, Nigro J, et al. Measurement of DNA copy number at microsatellite loci using quantitative PCR analysis. Cancer Res 2000;60:5405–5409.
Veltman JA, Fridlyand J, Pejavar S, et al. Array-based comparative genomic hybridization for genome-wide screening of DNA copy number in bladder tumors. Cancer Res 2003;63:2872–2880.
Jain AN, Tokuyasu TA, Snijders AM, et al. Fully automatic quantification of microarray image data. Genome Res 2002;12:325–332.
Patil MA, Gütgemann I, Zhang J, et al. Array-based comparative genomic hybridization reveals recurrent chromosomal aberrations and Jab1 as a potential target for 8q gain in hepatocellular carcinoma. Carcinogenesis 2005;26:2050–2057.
Ueki T, Fujimoto J, Suzuki T, et al. Expression of hepatocyte growth factor and its receptor c-met proto-oncogene in hepatocellular carcinoma. Hepatology 1997;25:862–866.
Kusano N, Shiraishi K, Kubo K, et al. Genetic aberrations detected by comparative genomic hybridization in hepatocellular carcinomas: their relationship to clinicopathological features. Hepatology 1999;29:1858–1862.
Wilkens L, Bredt M, Flemming P, et al. Differentiation of liver cell adenomas from well-differentiated hepatocellular carcinomas by comparative genomic hybridization. J Pathol 2001;193:476–482.
Buckley AF, Burgart LJ, Kakar S . Epidermal growth factor receptor expression and gene copy number in fibrolamellar hepatocellular carcinoma. Hum Pathol 2006;37:410–414.
Fearon ER, Cho KR, Nigro JM, et al. Identification of a chromosome 18q gene that is altered in colorectal cancers. Science 1990;247:49–56.
Hahn SA, Schutte M, Hoque AT, et al. DPC4, a candidate tumor suppressor gene at human chromosome 18q21.1. Science 1996;271:350–353.
Ding SF, Delhanty JD, Bowles L, et al. Infrequent chromosome allele loss in fibrolamellar carcinoma. Br J Cancer 1993;67:244–246.
Sirivatanauksorn Y, Sirivatanauksorn V, Lemoine NR, et al. Genomic homogeneity in fibrolamellar carcinomas. Gut 2001;49:82–86.
Zhao G, Su S, Borek D, et al. Long survival and prognostic factors in hepatocellular carcinoma. J Surg Oncol 1990;45:257–260.
McPeake JR, O’Grady JG, Zaman S, et al. Liver transplantation for primary hepatocellular carcinoma: tumor size and number determine outcome. J Hepatol 1993;18:226–234.
Moreno-Luna LE, Arrieta O, García-Leiva J, et al. Clinical and pathologic factors associated with survival in young adult patients with fibrolamellar hepatocarcinoma. BMC Cancer 2005;5:142.
Acknowledgements
This study was supported by grant from UCSF Research Education and Allocation Committee and UCSF Liver Center.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kakar, S., Chen, X., Ho, C. et al. Chromosomal changes in fibrolamellar hepatocellular carcinoma detected by array comparative genomic hybridization. Mod Pathol 22, 134–141 (2009). https://doi.org/10.1038/modpathol.2008.178
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2008.178
Keywords
This article is cited by
-
Fibrolamellar Hepatocellular Carcinoma
The American Journal of Gastroenterology (2009)