Abstract
Cholesterol 7α-hydroxylase (CYP7A1) regulates the balance between cholesterol supply and metabolism by catalyzing the rate-limiting step of bile acid biosynthesis. The transcriptional activity of CYP7A1 is tightly controlled by various nuclear receptors. A forkhead transcription factor O1 (FOXO1) plays a critical role in metabolism, and insulin inactivates FOXO1 through Akt-dependent phosphorylation and nuclear exclusion. We investigated the role of insulin-Akt-FOXO1 signaling pathway in CYP7A1 transcriptional regulation since we found putative insulin-response elements, FOXO1 binding sequences, in both rat and human CYP7A1 promoters. However, ectopic expression of FOXO1 increased the rat CYP7A1-, but mildly reduced human CYP7A1-promoter activities in a dose-dependent manner. Similarly to bile acids, insulin treatment increased small heterodimer partner (SHP) mRNA rapidly and transiently, leading to the suppression of CYP7A1 transcription in both human and rodents. Chromatin immunoprecipitation showed that FOXO1 directly bound to rat CYP1A1 promoter in the absence of insulin. FOXO1 binding to the rat promoter was diminished by insulin treatment as well as by expression of SHP. Our results suggest that the stimulation of insulin- signaling pathway of Akt-FOXO1 and SHP expression may regulate cholesterol/bile acid metabolisms in liver, linking carbohydrate and cholesterol metabolic pathways. A prolonged exposure of insulin in hyperinsulinemic insulin resistance or diabetic status represses CYP7A1 transcription and bile acid biosynthesis through SHP induction and FOXO1 inactivation, leading to impairment of the hepatic cholesterol/bile acid metabolisms.
Similar content being viewed by others
Introduction
Cholesterol 7α-hydroxylase (cytochrome P450 7A1, CYP7A1) plays an important role in maintaining cholesterol homeostasis by catalyzing the rate-limiting step in bile acid biosynthesis from hepatic cholesterol (Myant and Mitropoulos, 1977; Jelinek et al., 1990; Redinger, 2003). CYP7A1 is regulated mainly at the transcription level by various effectors such as diurnal rhythm, stress, xenobiotics, cholesterol, and bile acids, the direct end products of CYP7A1 (Yamagata et al., 2004). Transcriptional regulation of CYP7A1 is achieved by interactions among various nuclear hormone receptors, including hepatocyte nuclear factor-4α (HNF-4α) (Cooper et al., 1997), alpha(1)-fetoprotein transcription factor (FTF) (del Castillo-Olivares and Gil, 2000), small heterodimer partner (SHP) (Goodwin et al., 2000; Lu et al., 2000), and peroxisome proliferator activated receptor γ coactivator-1α (PGC-1α) (Shin et al., 2003; Ponugoti et al., 2007).
Forkhead transcription factor O1 (FOXO1) mediates the important effects of insulin as a member of insulin signaling cascade (Barthel et al., 2005) in liver, pancreas, skeletal muscle, adipose tissue and hypothalamus. In hypothalamus, FOXO1 controls food intake by regulating the transcription of orexigenic (NPY and AgRP) and antiorexigenic (POMC) neutopeptides (Kim et al., 2006). FOXO1 is exclusively expressed in β-cells in adult pancreas and involved in proliferation and differentiation of β-cell (Kitamura and Ido Kitamura, 2007). In adipose tissue and skeletal muscle, FOXO1 prevents differentiations of preadipocytes and myoblast (Farmer, 2003; Kamei et al., 2004). In liver, FOXO1 augments hepatic glucose production through transcription of gluconeogenic genes, PEPCK and G6Pase (Puigserver et al., 2003), and suppresses glycolysis and lipogenesis by inhibiting expressions of glucokinase and SREBP-1c. FOXO1 also enhances the apoprotein apoC-III expression (Altomonte et al., 2004), resulting in the increased plasma triglyceride levels. Thus, FOXO1 regulates multiple aspects of glucose and lipid metabolism in the liver. FOXO1 knockout mice are embryonic lethal due to incomplete vascular development (Furuyama et al., 2004). Transgenic mice expressing FOXO1 showed the impaired insulin sensitivity, glucose intolerance, and the age-related muscle atropy, all of which lead to obesity and diabetes (Kamei et al., 2004; Zhang et al., 2006).
FOXO1 activity is regulated by PI3K/Akt-mediated phosphorylation (van der Horst and Burgering, 2007). In response to insulin, PI3K phosphorylated Akt, and the phospho-Akt (pAkt) directly phosphorylates FOXO1 at three conserved residues (Thr24, Ser256, and Ser319). The phospho-FOXO1 (pFOXO1) is excluded from nucleus, and degraded by ubiquitin proteasome pathway in cytoplasm, which results in the suppression of target gene transcriptions. Therefore, mutations at phosphorylation sites of FOXO1 generate constitutively active form of FOXO1 (FOXO1-3A) which activates target gene expressions (Biggs et al., 1999; Tang et al., 1999). FOXO1 is a nuclear transcription factor which interacts with insulin response elements (IREs), T(G/A)TTT(T/G) (G/T), in the promoter of target genes involved in cell cycle, apoptosis, oxidative-stress resistance, and metabolism (Barthel et al., 2005; Kitamura and Ido Kitamura, 2007; van der Horst and Burgering, 2007). FOXO1 also regulates genes as a co-activator or co-repressor of many transcription factors.
We found eight and five putative IREs in the rat and human CYP7A1 promoters, respectively. There are several reports on differential regulation of mouse and human CYP7A1 (Chen et al., 2001; Goodwin et al., 2003; Kim et al., 2004). In the present study, we investigated whether FOXO1 might participate in the regulation of rat CYP7A1 transcription to control cholesterol/bile acid metabolisms as well as glucose metabolism in liver, possibly linking the two major metabolic pathways. Using co-transfection assays, we found the FOXO1-dependent rat CYP7A1 transcription, which was suppressed by the insulin/ PI3K/Akt pathway and bile acids. Besides the inactivation of the signaling pathway, up-regulation of SHP by insulin may lead to the suppression of CYP7A1 like bile acids.
Results
FOXO1 is the positive regulator of the rat CYP7A1 transcription
The sequences of rat CYP7A1 promoter region (between -402 and +4 from transcriptional start or between -436 and -30 from translation start codon) are rich in A-T and contain 8 putative FOXO binding consensus sequences (IRE-likes) (Figure 1A). To test if FOXO1 induces rat CYP7A1 transcription, HepG2 cells were co-transfected with pCYP7A1-luc and various amounts of FOXO1. Figure 1B showed that FOXO1 increased the CYP7A1-luc activity in a dose-dependent manner. CYP7A1 activation by the constitutively active (dominant positive) form of FOXO1 (FOXO1-3A) was stronger than by the wild type FOXO1, which suggested that CYP7A1 promoter activity might depend on FOXO1 phosphorylation. A well-known co-activator PGC-1α exhibited the dose-dependent increase of CYP7A1-luc with or without 100 ng FOXO1 (Figures 1C and 1D). The results suggest that FOXO1 may modulate CYP7A1 transcriptional activity onto its promoter and PGC-1α co-regulates the CYP7A1 transcription.
FOXO1 and PGC-1α-dependent activation of rat CYP7A1 promoter. (A) Eight putative FOXO1 binding sites (IRE consensus sequence, bold and underlined) in rat CYP7A1 promoter (-402 to +4 from transcriptional start + 1). (B-D). Induction of rat CYP7A1 promoter (rCYP7A1-luc) activity by FOXO1 and PGC-1α. HepG2 cells were transiently co-transfected with rCYP7A1-luc with or without increasing amounts of FOXO1, FOXO1-3A, or PGC-1α as indicated and harvested for luciferase assay. In D, 100 ng Foxo1 was co-transfected. Luciferase activities were expressed as a fold induction of the mock control. Values are mean ± SD of three independent assays of duplicates (n = 6).
Co-operative activation of rat CYP7A1 by FTF, HNF-4α and PGC-1α
Many nuclear receptors are involved in CYP7A1 transcriptional regulation. The transcriptional regulatory network of CYP7A1 is complex to clarify the interactions between nuclear receptors. This is partly because nuclear receptors differentially regulate rat and human CYP7A1 expression. Therefore, we confirmed the transcription activities of FTF (mouse homologue LRH-1, liver receptor homologue-1) and HNF-4α on rat CYP7A1-luc. As expected, FTF increased CYP7A1-luc in a dose-dependent manner and HNF-4α additionally increased the FTF-induced CYP7A1 (Figure 2A). PGC-1α did not co-activate FTF activity (Figure 2B), whereas it did strongly HNF-4α activity (Figure 2C) to the similar extend with FOXO1 (Figure 1D). When CYP7A1 is highly induced by FTF and HNF-4α, co-activation of PGC-1α was not as strong as in HNF-4α only (Figure 2D). The results suggest that a strong repressor may maintain the appropriate transcriptional level of CYP7A1 through controlling a positive regulator of FTF.
Co-activation of FTF- or HNF-4α-induced rCYP7A1 transcription by PGC-1α. HepG2 cells were transiently transfected with rCYP7A1-luc and nuclear receptors (100 ng FTF and/or 800 ng HNF-4α) with the indicated amounts of PGC-1α. Total plasmid amount remains 1.9 µg with mock DNA in all experiments. Luciferase activities were expressed as fold induction of mock control. Values are mean ± SD of three independent assays of duplicates.
Strong repression of rat CYP7A1 by SHP in the presence of FTF
Bile acid mediates feedback repression of CYP7A1 by enhancing the expression of SHP (Lu et al., 2000), since SHP hinders LRH-1 binding to CYP7A1 promoter by interacting directly with LRH-1, a mouse homologue of FTF (Goodwin et al., 2000). It is not clear whether SHP interacts with FOXO1 or PGC-1α to control CYP7A1 transcription. Again, we performed co-transfection analyses using rat CYP7A1-luc and combinations of nuclear receptors. SHP reduced the CYP7A1-luc activity (Figure 3A). Here, it should be noted that 0.1 µg of SHP inhibited 50% of CYP7A1 transcription and 1 µg of SHP decreased only 60% of the activity. On the other hand, SHP clearly repressed the FOXO1-induced CYP7A1 transcription in a dose-dependent manner (Figure 3B). Similarly to LRH-1, the FTF-induced CYP7A1 activity was strongly suppressed by co-expression of SHP (Figure 3C). The inhibitory effects of SHP on FTF were stronger than on FOXO1 or HNF-4α (Figure 3D), implying that SHP might preferentially interact with FTF rather than FOXO1 or HNF-4α. Therefore, the presence of FTF makes the CYP7A1 transcription control sensitive to SHP. It is not clear why a low concentration of SHP stimulates CYP7A1 a little further in the presence of HNF-4α (Figures 3D and 3F). In the presence of FTF, however, 100 ng of SHP almost completely suppressed the CYP7A1 promoter activity despite of co-expressions of positive regulators, HNF-4α and PGC-1α (Figure 3E). Maximum amount (1 µg) of PGC-1α co-expression did not restore the SHP-mediated repression to the initial level without SHP. In contrast, 100 ng of SHP was enough to abolish the CYP7A1 promoter activity induced by FTF, HNF-4α and PGC-1α (Figure 3F). To achieve the SHP-mediated repression in the presence of PGC-1α, an approximate molar ratio of SHP to PGC-1α was 1:40, which means only 1/40 of SHP is required to abolish the PGC-1α-mediated co-activation, implying that SHP may be the most critical control factor for CYP7A1 transcription.
Co-repression of rCYP7A1 by SHP. (A-E) HepG2 cells were transiently transfected with rCYP7A1-luc and nuclear receptors (100 ng FOXO1 or, 800 ng FTF, 800 ng HNF-4α) with the indicated amounts of PGC-1α and SHP. (F) FTF, HNF-4α, PGC-1α 800 ng each. Luciferase activities were expressed as fold induction of mock control. Values are mean ± SE of three independent assays of duplicates.
FOXO1 is a mild negative regulator of the human CYP7A1 transcription
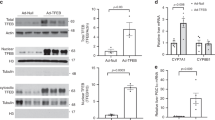
To compare the effects of FOXO1 on human CYP7A1 promoter, we have constructed the human CYP promoter (-326 to +104)-luciferase plasmids by PCR and performed the similar co-transfection assays (Figure 4A). Although the human CYP7A1 promoter contains three putative, but not clearly conserved, IREs, FOXO1 dose-dependently decreased the human CYP7A1 promoter activity in HepG2 cells (Figure 4B). Here, we have to note that rat CYP7A1 promoter was 10-fold more sensitive to FOXO1 than human CYP7A1 to confer the activity. Deletion of IRE-1 (ΔIRE-1) had no effects on hCYP7A1-luc activity (Figure 4C). Deletions of IRE-2 or IRE-3 (ΔIRE-2 or ΔIRE-3) enhanced hCYP7A1-luc but FOXO1-mediated repression was not abolished. Thus, IRE-1 is not functional and both IRE-2 and IRE-3 may be the binding sites of FOXO1. ChIP analysis of HepG2 cells revealed the direct binding of FOXO1 to human CYP7A1 promoter (Figure 4D). FOXO1-3A overexpression using adenovirus enhanced FOXO1 binding to human CYP7A1 promoter, but it did not increase PGC-1α binding to the promoter. Insulin treatment for 30 min increased phosphorylation and degradation of FOXO1, but did not alter the PGC-1α expression (Figure 4E).
(A) Human CYP7A1 promoter sequences. Three putative FOXO1 binding sites are shown in bold (IRE-1,2,3). The 430 bp fragment of human CYP7A1 promoter (-326 to +104 from transcriptional start +1) was cloned into pGL3-Basic vector. (B) Repression of human CYP7A1 promoter (hCYP7A1-luc) activity by FOXO1 and FOXO1-3A. (C) The wild type (WT) hCYP7A1-luc and the IRE-deleted-luc mutants were co-transfected into HepG2 cells with or without FOXO1. (D) Chromatin IP. HepG2 cells were infected with adenovirus containing constitutively active FOXO1-3A(+) or LacZ(-) for 36 h and chromatine IP was performed. Cross-linked chromatin solution was immunoprecipitated with anti-FOXO1 or anti-PGC1α. Normal rabbit IgG or acetylated H3 (Ac-H3) were utilized as negative and positive controls, respectively. Ten percent of the soluble chromatin used in the reaction was used as inputs. Precipitated DNA was analyzed by PCR using primer sets for the CYP7A1 promoter regions. Arrows in (A) are the primers for ChIP assay. (E) Western blots. HepG2 cells were stimulated with insulin for 30 min, and harvested for Western blot.
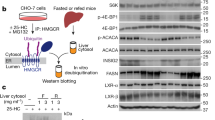
CYP7A1 expression was down-regulated by insulin in vitro and in vivo
Diabetic patients and experimental diabetic animals showed the enhanced levels of bile acids pool and excretion. Insulin normalized the elevated bile acids in diabetes and reduced the CYP7A1 transcription in rat primary hepatocytes (Twisk et al., 1995). Reflecting insulin-activated PI3K-Akt protein kinase cascade and FOXO1 phosphorylation, we expected that insulin would repress CYP7A1 expression via inactivation of FOXO1. The insulin treatment down-regulated the mRNA levels of CYP7A1 in HepG2 cells (Figure 5) and murine liver (Figure 6). Insulin decreased hCYP7A1 mRNA within 4 h, while it rapidly increased hSHP mRNA up to 17-fold in 2 h in HepG2 cells, which were determined by semi-qRT-PCR (Figure 5B) or real-time PCR (Figure 5C). In order to determine the effects of insulin on mRNAs of rodent CYP7A1 and SHP in vivo, we intraperitoneally injected insulin into C57BL/6 mice (n = 3) and isolated livers after 2 h. Insulin also down-regulated the mCYP7A1 mRNA and enhanced mSHP mRNA in mouse liver (Figure 6). The results suggest that the rapid increase of SHP may be involved in the insulin-mediated repression of CYP7A1 transcription.
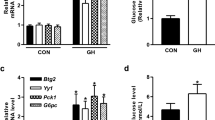
Opposite regulations of human hCYP7A1 and hSHP by insulin in HepG2 cells. Quiescent HepG2 cells in SFM were treated with insulin for 2 h or 4 h. Total RNA was isolated and the mRNA levels of hCYP7A1 or hSHP were determined by semi-qRT-PCR. (A) Representative gel pictures of semi-qRT-PCR. (B) The band intensities of the gel were quantified by densitometry and graphed. (C) Real time qPCR of hCYP7A1 and hSHP mRNAs using the insulin-treated HepG2 cells. *P < 0.05, **P < 0.01 vs. PBS.
Opposite regulations of mouse mCYP7A1 and mSHP by insulin in mouse liver. The fed C57BL/6 mice (9 weeks old, n = 3) were injected (1 U/kg in 100 µl, i.p.). After 2 h, hepatic total RNA was isolated and the levels of mCYP7A1 and mSHP were determined. (A) Representative gel pictures of semi-qRT-PCR. (B) The band intensities of the gel were quantified by densitometry and graphed. (C) Real time qPCR of mCYP7A1 mRNAs in the insulin-treated mouse liver. **P < 0.01 vs. PBS.
Rat CYP7A1 promoter activity was regulated by insulin-PI3K-Akt-FOXO1 signaling pathway
To determine if FOXO1 phosphorylation by insulin is involved in rat CYP7A1 transcriptional activity, constitutively active FOXO1, FOXO1-3A, was co-transfected with rCYP7A1-luc. Figure 7A shows that FOXO1 phosphorylation also contributes to the insulin-mediated decrease of rat CYP7A1 because FOXO1-3A transfection-induced CYP7A1 was not affected by insulin differently from the FOXO1-treated. FOXO1 activity is negatively regulated by Akt and PI3K. PI3K consists of p85 regulatory and p110 catalytic subunits. The labile p110 is stabilized by p85. As expected, the FOXO1-induced rat CYP7A1 was inactivated by Akt or PI3K-p110 (dominant positives) and further activated by PI3K-Δp85 (dominant negative) co-transfection, supporting that the effects of FOXO1 on rat CYP7A1 might be under the control of PI3K and Akt (Figure 7B). Effects of PI3K-Akt were abolished in the FOXO1-3A transfected cells. ChIP analysis of rat hepatoma H4IIE cells demonstrated that FOXO1 was bound directly to CYP7A1 promoter and insulin displaced its binding when FOXO1 was overexpressed (Figure 7C). Co-transfection of SHP also reduced FOXO1 bound to the promoter, suggesting that augmentation of SHP expression by insulin suppressed the FOXO1-dependent CYP7A1 expression. CDCA is a hydrophobic primary bile acid which activates SHP transcription (Goodwin et al., 2000). CDCA also decreased rat CYP7A1 promoter activity of the FOXO1- or the FOXO1-3A-transfected cells in a dose-dependent manner (Figure 7D). Hence, both insulin and CDCA may down-regulate the FOXO1-dependent rat CYP7A1 transactivation through SHP/FOXO1-dependent pathway.
FOXO1 phosphorylation-dependent repression of rat CYP7A1 transcription. (A) FOXO1-3A abolished the insulin-mediated repression rat CYP7A1 promoter activity. HepG2 cells transfected with rCYP7A1-luc, 100 ng FOXO1 (or FOXO1-3A) in SFM were treated with insulin (100 nM) for 16 h and harvested for luciferase assays (n = 6). (B) Regulation of rCYP7A1 transcriptional activity through insulin/PI3K/Akt/FOXO1 signaling pathway. HepG2 cells were transiently co-transfected with rCYP7A1-luc, FOXO1 (or FOXO1-3A) in the presence of Δp85, p110 or Akt expression vectors as indicated. (C) Chromatin IP. Rat H4IIE cells were transfected with FOXO1 and/or SHP for 48 h, treated with insulin for 1 h and ChIP was performed. Cross-linked chromatin solution was immunoprecipitated with anti-FOXO1. Normal mouse IgG or anti-RNA polymerase II (RNA pol II) were utilized as negative and positive controls, respectively. Ten percent of the soluble chromatin used in the reaction was used as inputs. Precipitated DNA was analyzed by PCR using primer sets for the rat CYP7A1 promoter regions. FOXO bound to the promoter was quantified and normalized by RNA pol II. (D) FOXO1 phosphorylation-independent inhibition of rCYP7A1 by bile acids. HepG2 cells transfected with rCYP7A1-luc, FOXO1 (or FOXO1-3A) were treated with CDCA (0, 25, 50, 100 µM) in SFM for 24 h. Luciferase activities were expressed as a fold induction of the mock control (n = 6, **P < 0.01 vs. PBS).
Discussion
In the present study, we demonstrated that FOXO1 was a transcriptional activator of rat CYP7A1 under the tight control of insulin-PI3K-Akt signaling cascade. Insulin and bile acid suppress rat CYP7A1 transcription via both FOXO1 inactivation and SHP induction. PGC-1α and SHP were co-regulators of rat CYP7A1 transcription. On the other hand, FOXO1 was not, but SHP was a critical modulator of human CYP7A1 transcription in response to insulin.
The transcriptional regulation of CYP7A1 is complex to understand. The confusion had been caused by the differential regulation of bile acid and cholesterol metabolisms between rodents and humans. First, FTF stimulated the rat CYP7A1 transcription (del Castillo-Olivares and Gil, 2000; Goodwin et al., 2000; Lu et al., 2000), but it repressed the human CYP7A1 activity (Chen et al., 2001). Second, LXRα stimulated the rat CYP7A1, but not human CYP7A1 promoter activity because human CYP7A1 promoter lacks LXR elements (Chen et al., 2002; Goodwin et al., 2003). Here, we added another transcription factor, FOXO1, showing differences in CYP7A1 regulation between rodent and human.
There are several contradictory reports on the CYP7A1 gene regulation by insulin. It has been reported that insulin inhibits the transcription of mouse CYP7A1 because the insulin-induced SREBP-1c blocks the functional interaction between HNF-4α and PGC-1α as a non-DNA binding inhibitor (Ponugoti et al., 2007). Furthermore, microarray analysis of liver from the transgenic mice overexpressing FOXO1 revealed 2-fold increase of CYP7A1 and 40% decrease of SREBP-1c (Zhang et al., 2006). Feeding (physiological increase of insulin) decreased CYP7A1 mRNA by 90% compared to the fasting state in mice. The fasting induction of CYP7A1 was blunted in liver-specific FOXO1 knock-out mice (Shin and Osborne, 2009). In contrast, insulin at physiological concentrations showed dual effects on human CYP7A1 gene expression; a rapid stimulation of human CYP7A1 through FOXO1 inactivation and a late repression of CYP7A1 mediated by prolonged insulin exposure-induced SREBP-1c (Li et al., 2006). It has been reported that FOXO1 is a strong repressor in human CYP7A1 transcription because siRNA knockdown of FOXO1 induced 6-fold CYP7A1 mRNA and dominant positive FOXO1 inhibited it in human primary culture (Li et al., 2009).
It is confusing because insulin treatment decreases both human and rodent CYP7A1 mRNA expressions despite of the opposite role of FOXO1 in two species. Here, we have to note that insulin rapidly and strongly induced SHP mRNA in both human cells and mice. SHP is an interesting and powerful negative regulator of various nuclear receptors in vivo and in vitro because it lacks the DNA binding domain (Lee et al., 2010). SHP directly binds to FOXO1 or HNF-4α to dissociate them from promoters of G6Pase or PEPCK, respectively (Yamagata et al., 2004). SHP also suppressed the FTF-mediated transactivation in the strongest manner. Therefore, it is possible that insulin-induced SHP hinders the binding of FOXO1, HNF-4α, and FTF to rat and human CYP1A1 promoters by interacting with them, resulting in suppression of transcription. It was confirmed that SHP overexpression dissociated FOXO1 from rat CYP7A1 promoter (Figure 6C). Alternatively, the transcriptional regulation of human CYP7A1 by insulin may be achieved mainly by transcription factor other than FOXO1, like SREBP-1c differently from rodent CYP7A1. Thus, SREBP1c would link the insulin action to lipid/bile acid homeostasis in human. In conclusion, the results suggest that a prolonged exposure of insulin in hyperinsulinemic insulin resistance or diabetic status represses CYP7A1 transcription and bile acid biosynthesis through SHP induction and FOXO1 inactivation, leading to impairment of the cholesterol/bile acid metabolisms. The insulin-mediated regulatory network of rat CYP7A1 expression is summarized in Figure 8. Further study on the relation among FOXO1, SHP, and CYP7A1 using clinical samples may enhance the understanding of pathogenesis of cardiovascular complications in diabetic patients. Again, our study draws an attention to rodent study when we investigate the molecular mechanism of insulin resistance and the effects of therapeutic interventions on bile acid or cholesterol metabolism.
Methods
Cell culture and transient transfection
HepG2 cells (ATCC, HB-8085) were cultured in Minimum Essential Medium (MEM) supplemented with 10% fetal bovine serum (FBS) and 100 µg/ml penicillin and 100 µg/ml streptomycin at 37℃/5% CO2. For transient transfections, the cells (4 × 105 cells/well) in 6- well plates were transfected with the indicated plasmid(s) and expression vector of β-galactosidase (pcDNA3.1/LacZ) by calcium phosphate co-precipitation method (Jeong et al., 2009). The total amounts of expression vectors were kept constant by adding the pcDNA3.1/HisC expression vector to transfection. After 36 h incubation in MEM containing 10% FBS, the cells were harvested and luciferase activity was measured using luciferase assay kit (Promega, Madison, WI) and luminometer (Berthold, German). The transfection efficiencies were normalized by the β-galactosidase activity.
To determine the effects of insulin or chenodeoxycholic acid (CDCA), HepG2 cells were cultured in serum free MEM for 16 h and treated with 100 nM insulin for 2 h, 4 h or 16 h or with 0-100 µM CDCA for 24 h. The cells were harvested for western blot or semi-quantitative RT-PCR analysis.
Animals
C57BL6 mice (9 weeks old, n = 3) were injected intraperitoneally (1 U/kg in 100 µl). After 2 h, hepatic total RNA was isolated using Trizol (Invitrogen, Carlsbad, CA), and the CYP7A1 mRNA was determined by semi-quantitative and real time qRT-PCR using mouse specific primers.
Plasmids
FOXO1 and constitutively active FOXO1 (FOXO1-3A) mammalian expression vectors were provided by Dr. Guan K.L. (Michigan University) (Kim et al., 2006). Rat CYP7A1 promoter (-402 to +4 bp)-luciferase construct (pCYP7A1-luc), HNF-4α, and FTF expression vectors were obtained from Dr. Choi H.S. (Chonnam University, Korea). Human CYP7A1 promoter (-326 to +105 from transcriptional start +1) were amplified by PCR and cloned between Kpn I and Hind III of pGL3-basic and its sequences were verified (Figure 4). Three deletion mutants (ΔIRE-1, ΔIRE-2, ΔIRE-3) that lacks each putative FOXO1 binding site were generated using oligonucleotide-directed mutagenesis and a two-step PCR procedures as described (Jeong et al., 2009). The primer sequences for two-step PCR are following: hCYP7A1 promoter (5'-cgg ggt acc ttt atg tat tta tta gct at-3' and 5'-ccc aag ctt ctg cta tag caa tcc ccc aa-3'), ΔIRE-1 (5'-taaacaggtt acacatacca-3' and 5'-tggtatgtgt aacctgttta-3'), ΔIRE-2 (5'-aatggctaat tgtcaaccaa-3' and 5'-ttggttgaca attagccatt-3'), ΔIRE-3 (5'-agcttgagtc agtggcatcc-3' and ggatgccact gactcaagct-3'). Mammalian expression plasmids of Akt, PI3K p110, PI3K Δp85 were generously provided by Drs. Park HS (University of Seoul, Korea) and Ha JH (Kyung Hee University, Korea).
Chromatin immunoprecipitation (ChIP)
For rat CYP7A1 promoter binding, rat hepatoma H4IIE cells in 100 mm plate were transfected with 10 µg FOXO1 and/or 2.5 µg SHP using Superfect (Qiagen, Hilden, Germany). After 48 h, the transfected cells were treated with 100 nM insulin for 1 h. For human CYP7A1 promoter binding, HepG2 cells were infected 14 h after plating with adenoviruses expressing either LacZ (Ad-LacZ) or dominant positive FOXO1-3A (Ad-FOXO1) at a multiplicity of infection (moi) of 50 (Kim et al., 2006). At 4 h after infection, fresh growth medium was added and incubated for 36 h. ChIP assay was performed using the transfected or infected cells. Briefly, the cells were subjected to formaldehyde cross-linking for 15 min. The cross-linked protein-DNA complex in the soluble chromatin, which was prepared by sonication, was immunoprecipitated with anti-FOXO1, or anti-PGC-1α (Santa Cruz Biotechnology, Santa Cruz, CA). IgG was used as a negative control. Anti-acetyl histone 3 (Ac-H3, Santa Cruz, CA) or anti-RNA polymerase II (RNA pol II, Millipore, MA) antibodies was used as a positive control. Ten percent of the soluble chromatin used in the reaction was used as inputs. The precipitated DNA fragments were analyzed by 38-40 cycles of PCR using primers directed against human CYP7A1 promoter (forward 5'-tat gcc cat ctt aaa cag g-3' and reverse 5'-atc tag gcc aaa atc tct g-3') or rat CYP7A1 promoter (forward 5'-gct gtg gct tcc tgg tag atg-3' and reverse 5'-ccg aaa cag tgg gtc tga cta g-3').
Semi-quantitative and real time RT-PCR
Total RNA was prepared from cultured cells or livers by Trizol (Invitrogen). Semi-quantitative RT-PCR (semi-qRT-PCR) was performed to check primer dimer formation. RT-PCR products were visualized on 1.5% agarose gel and quantified by densitometry. Real time RT-PCR for CYP7A1 and SHP was performed using SYBR green dye on Light Cycler1.5 (Roche, Basel, Switzerland) with SYBR® Premix Ex Taq™ (TaKaRa, Otsushiga, Japan) at 95℃ for 10 sec, followed by 40 cycles of 95℃ for 5 s and 60℃ for 34 s according to the manufacturer's protocol. Measurement was performed in duplicate for each sample. Primer sequences, product size, annealing temperature and the number of cycles for semi-qRT-PCR were following: human CYP7A1 (101 bp, 61℃, 25 cycles), 5'- cca tta ggt gtt gtg cca cg-3' and 5'- cat cca tcg ggt caa tgc tt-3'; mouse CYP7A1 (102 bp, 65℃, 26 cycles), 5'- cca ggg aga tgc tct gtg ttc-3' and 5'- acc cag aca gcg ctc ttt gat -3'; human SHP (107 bp, 61℃, 25 cycles), 5'-gac ttc ctt ggt ttg gac aca-3' and 5'-gct ttc att ctc atc cca aga-3'; mouse SHP (101 bp, 65℃, 26 cycles), 5'- acg atc ctc ttc aac cca gat g-3' and 5'- ggc tcc aag act tca cac agt-3'; human and mouse 18S rRNA (102 bp, 60℃, 26 cycles), 5'-gag cga aag cat ttg cca ag-3' and 5'-ggc atc gtt tat ggt cgg aa-3'.
Statistical analysis
Student's t-tests were performed using Instat or Prism (GraphPad Software, San Diego, CA) with significance set at P < 0.05. Data represent mean ± standard deviation (SD) for experiments repeated 3-6 times on independent occasions.
Abbreviations
- CDCA:
-
chenodeoxycholic acid
- CYP7A1:
-
cholesterol 7α-hydroxylase
- FOXO1:
-
forkhead transcription factor O1
- FOXO1-3A:
-
constitutively active form of FOXO1
- FTF:
-
alpha(1)-fetoprotein transcription factor
- HNF-4α:
-
hepatocyte nuclear factor-4α
- IRE:
-
insulin-response elements
- PGC-1α:
-
peroxisome proliferator activated receptor γ coactivator-1α
- SHP:
-
small heterodimer partner
References
Altomonte J, Cong L, Harbaran S, Richter A, Xu J, Meseck M, Dong HH . Foxo1 mediates insulin action on apoC-III and triglyceride metabolism . J Clin Invest 2004 ; 114 : 1493 - 1503
Barthel A, Schmoll D, Unterman TG . FoxO proteins in insulin action and metabolism . Trends Endocrinol Metab 2005 ; 16 : 183 - 189
Biggs WH, Meisenhelder J, Hunter T, Cavenee WK, Arden KC . Protein kinase B/Akt-mediated phosphorylation promotes nuclear exclusion of the winged helix transcription factor FKHR1 . Proc Natl Acad Sci USA 1999 ; 96 : 7421 - 7426
Chen JY, Levy-Wilson B, Goodart S, Cooper AD . Mice expressing the human CYP7A1 gene in the mouse CYP7A1 knock-out background lack induction of CYP7A1 expression by cholesterol feeding and have increased hypercholesterolemia when fed a high fat diet . J Biol Chem 2002 ; 277 : 42588 - 42595
Chen W, Owsley E, Yang Y, Stroup D, Chiang JY . Nuclear receptor-mediated repression of human cholesterol 7alpha-hydroxylase gene transcription by bile acids . J Lipid Res 2001 ; 42 : 1402 - 1412
Cooper AD, Chen J, Botelho-Yetkinler MJ, Cao Y, Taniguchi T, Levy-Wilson B . Characterization of hepatic-specific regulatory elements in the promoter region of the human cholesterol 7alpha-hydroxylase gene . J Biol Chem 1997 ; 272 : 3444 - 3452
del Castillo-Olivares A, Gil G . Role of FXR and FTF in bile acid-mediated suppression of cholesterol 7alpha-hydroxylase transcription . Nucleic Acids Res 2000 ; 28 : 3587 - 3593
Farmer SR . The forkhead transcription factor Foxo1: a possible link between obesity and insulin resistance . Mol Cell 2003 ; 11 : 6 - 8
Furuyama T, Kitayama K, Shimoda Y, Ogawa M, Sone K, Yoshida-Araki K, Hisatsune H, Nishikawa S, Nakayama K, Ikeda K, Motoyama N, Mori N . Abnormal angiogenesis in Foxo1 (Fkhr)-deficient mice . J Biol Chem 2004 ; 279 : 34741 - 34749
Goodwin B, Jones SA, Price RR, Watson MA, McKee DD, Moore LB, Galardi C, Wilson JG, Lewis MC, Roth ME, Maloney PR, Willson TM, Kliewer SA . A regulatory cascade of the nuclear receptors FXR, SHP-1, and LRH-1 represses bile acid biosynthesis . Mol Cell 2000 ; 6 : 517 - 526
Goodwin B, Watson MA, Kim H, Miao J, Kemper JK, Kliewer SA . Differential regulation of rat and human CYP7A1 by the nuclear oxysterol receptor liver X receptor-alpha . Mol Endocrinol 2003 ; 17 : 386 - 394
Jelinek DF, Andersson S, Slaughter CA, Russell DW . Cloning and regulation of cholesterol 7 alpha-hydroxylase, the rate-limiting enzyme in bile acid biosynthesis . J Biol Chem 1990 ; 265 : 8190 - 8197
Jeong JH, Cho S, Pak YK . Sterol-independent repression of low density lipoprotein receptor promoter by peroxisome proliferator activated receptor gamma coactivator-1alpha (PGC-1alpha) . Exp Mol Med 2009 ; 41 : 406 - 416
Kamei Y, Miura S, Suzuki M, Kai Y, Mizukami J, Taniguchi T, Mochida K, Hata T, Matsuda J, Aburatani H, Nishino I, Ezaki O . Skeletal muscle FOXO1 (FKHR) transgenic mice have less skeletal muscle mass, down-regulated Type I (slow twitch/red muscle) fiber genes, and impaired glycemic control . J Biol Chem 2004 ; 279 : 41114 - 41123
Kim HJ, Kim JY, Kim JY, Park SK, Seo JH, Kim JB, Lee IK, Kim KS, Choi HS . Differential regulation of human and mouse orphan nuclear receptor small heterodimer partner promoter by sterol regulatory element binding protein-1 . J Biol Chem 2004 ; 279 : 28122 - 28131
Kim MS, Pak YK, Jang PG, Namkoong C, Choi YS, Won JC, Kim KS, Kim SW, Kim HS, Park JY, Kim YB, Lee KU . Role of hypothalamic Foxo1 in the regulation of food intake and energy homeostasis . Nat Neurosci 2006 ; 9 : 901 - 906
Kitamura T, Ido Kitamura Y . Role of FoxO proteins in pancreatic beta cells . Endocr J 2007 ; 54 : 507 - 515
Lee KM, Seo HY, Kim MK, Min AK, Ryu SY, Kim YN, Park YJ, Choi HS, Lee KU, Park WJ, Park KG, Lee IK . Orphan nuclear receptor small heterodimer partner inhibits angiotensin II-stimulated PAI-1 expression in vascular smooth muscle cells . Exp Mol Med 2010 ; 42 : 21 - 29
Li T, Kong X, Owsley E, Ellis E, Strom S, Chiang JY . Insulin regulation of cholesterol 7alpha-hydroxylase expression in human hepatocytes: roles of forkhead box O1 and sterol regulatory element-binding protein 1c . J Biol Chem 2006 ; 281 : 28745 - 28754
Li T, Ma H, Park YJ, Lee YK, Strom S, Moore DD, Chiang JY . Forkhead box transcription factor O1 inhibits cholesterol 7alpha-hydroxylase in human hepatocytes and in high fat diet-fed mice . Biochim Biophys Acta 2009 ; 1791 : 991 - 996
Lu TT, Makishima M, Repa JJ, Schoonjans K, Kerr TA, Auwerx J, Mangelsdorf DJ . Molecular basis for feedback regulation of bile acid synthesis by nuclear receptors . Mol Cell 2000 ; 6 : 507 - 515
Myant NB, Mitropoulos KA . Cholesterol 7 alpha-hydroxylase . J Lipid Res 1977 ; 18 : 135 - 153
Ponugoti B, Fang S, Kemper JK . Functional interaction of hepatic nuclear factor-4 and peroxisome proliferator-activated receptor-gamma coactivator 1alpha in CYP7A1 regulation is inhibited by a key lipogenic activator, sterol regulatory element-binding protein-1c . Mol Endocrinol 2007 ; 21 : 2698 - 2712
Puigserver P, Rhee J, Donovan J, Walkey CJ, Yoon JC, Oriente F, Kitamura Y, Altomonte J, Dong H, Accili D, Spiegelman BM . Insulin-regulated hepatic gluconeogenesis through FOXO1-PGC-1alpha interaction . Nature 2003 ; 423 : 550 - 555
Redinger RN . The coming of age of our understanding of the enterohepatic circulation of bile salts . Am J Surg 2003 ; 185 : 168 - 172
Shin DJ, Campos JA, Gil G, Osborne TF . PGC-1alpha activates CYP7A1 and bile acid biosynthesis . J Biol Chem 2003 ; 278 : 50047 - 50052
Shin DJ, Osborne TF . FGF15/FGFR4 integrates growth factor signaling with hepatic bile acid metabolism and insulin action . J Biol Chem 2009 ; 284 : 11110 - 11120
Tang ED, Nunez G, Barr FG, Guan KL . Negative regulation of the forkhead transcription factor FKHR by Akt . J Biol Chem 1999 ; 274 : 16741 - 16746
Twisk J, Hoekman MF, Lehmann EM, Meijer P, Mager WH, Princen HM . Insulin suppresses bile acid synthesis in cultured rat hepatocytes by down-regulation of cholesterol 7 alpha-hydroxylase and sterol 27-hydroxylase gene transcription . Hepatology 1995 ; 21 : 501 - 510
van der Horst A, Burgering BM . Stressing the role of FoxO proteins in lifespan and disease . Nat Rev Mol Cell Biol 2007 ; 8 : 440 - 450
Yamagata K, Daitoku H, Shimamoto Y, Matsuzaki H, Hirota K, Ishida J, Fukamizu A . Bile acids regulate gluconeogenic gene expression via small heterodimer partner-mediated repression of hepatocyte nuclear factor 4 and Foxo1 . J Biol Chem 2004 ; 279 : 23158 - 23165
Zhang W, Patil S, Chauhan B, Guo S, Powell DR, Le J, Klotsas A, Matika R, Xiao X, Franks R, Heidenreich KA, Sajan MP, Farese RV, Stolz DB, Tso P, Koo SH, Montminy M, Unterman TG . FoxO1 regulates multiple metabolic pathways in the liver: effects on gluconeogenic, glycolytic, and lipogenic gene expression . J Biol Chem 2006 ; 281 : 10105 - 10117
Acknowledgements
Authors thank Sunny Lim and Jun Woo Kim for their excellent technical assistance. This study was supported by the Kyung Hee University Research Fund in 2007 (KHU-20071406).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0/) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Park, WH., Pak, Y. Insulin-dependent suppression of cholesterol 7α-hydroxlase is a possible link between glucose and cholesterol metabolisms. Exp Mol Med 43, 571–579 (2011). https://doi.org/10.3858/emm.2011.43.10.064
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.3858/emm.2011.43.10.064
Keywords
This article is cited by
-
Extracting proteins involved in disease progression using temporally connected networks
BMC Systems Biology (2018)
-
Insulin Regulates Adrenal Steroidogenesis by Stabilizing SF-1 Activity
Scientific Reports (2018)