Abstract
We investigated transmission ratio distortion within an Icelandic population of Arabidopsis lyrata using 16 molecular markers unlinked to the S-locus. Transmission ratio distortion was found more often than expected by chance at the gametic level, but not at the genotypic or zygotic level. The gametic effect may be due to meiotic drive or selection acting postmeiotically. At the gametic level, 10.9% of the tests were significant, which is substantially lower than earlier observed in an interpopulation cross (allowing for differences in power)—suggesting that the high level of transmission ratio distortion in the interpopulation cross is due to population divergence. It is also substantially lower than previously observed in intrapopulation crosses at the self-incompatibility locus, suggesting inherent fitness differences of the self-incompatibility alleles. We discuss the possible role of deleterious alleles accumulating at loci under balancing selection. Zygotic effects play a larger role in the interpopulation cross than in the intrapopulation crosses suggesting that Dobzhansky–Muller incompatibilities may be accumulating between the widely diverged populations.
Similar content being viewed by others
Introduction
Transmission ratio distortion, or non-Mendelian segregation, is often observed in both animals and plants. There are various factors that may give rise to distorted segregation. Meiotic drive causes uneven representation of the alleles of a locus in gametes. While meiotic drive may be an important evolutionary force, it may often go undetected because it is expected to be evolutionarily transient, spreading to fixation unless halted by antagonistic effects (Taylor and Ingvarsson, 2003). Another possible factor causing transmission ratio distortion is selection at the gametophytic or zygotic level, including pollen competition and selective abortion of embryos (Korbecka et al., 2002, and references therein). Epistatic interactions between two or more loci (Dobzhansky–Muller incompatibilities) or chromosomal differentiation related to divergence may give rise to transmission ratio distortion in crosses between species or subspecies (reviewed in Fishman and Willis, 2001). Severe transmission ratio distortion has been found in progeny of both interspecific crosses (Fishman and Willis, 2001; Myburg et al., 2004; Yin et al., 2004) and intraspecific crosses between diverged populations (Hall and Willis, 2005; Törjék et al., 2006). Earlier studies have generally reported a larger transmission ratio distortion in interspecific crosses than in intraspecific crosses (Zamir and Tadmor, 1986; Jenczewski et al., 1997).
Arabidopsis lyrata, a member of the Brassicaceae family, has become an important model species for evolutionary plant genomics and its genome sequencing is underway (Mitchell-Olds, 2001; Clauss and Koch, 2006). In contrast to the closely related model species A. thaliana, it is outcrossing and thus has some advantages as a model for population genetic analysis of life history evolution (Riihimäki and Savolainen, 2004; Clauss and Mitchell-Olds, 2006). Furthermore, it serves as a model for the evolution of sporophytic self-incompatibility systems (Kusaba et al., 2001; Schierup et al., 2001; Nasrallah et al., 2002; Bechsgaard et al., 2006).
Kuittinen et al. (2004) constructed a genetic map of A. lyrata by crossing individuals from two genetically diverged populations (Karhumäki, Russia and Mjällom, Sweden) that show substantial morphological and genetic differentiation (Nei's genetic distance estimated by different markers: 0.51 by Jonsell et al., 1995; 0.39–0.51 by van Treuren et al., 1997 and 1.31 by Muller et al., 2007). Segregation patterns of about 50% of the molecular markers used to construct the map deviated significantly from Mendelian expectations.
Transmission ratio distortion has also been observed at the sporophytic self-incompatibility (S-) locus in crosses within a single natural population of A. lyrata (Bechsgaard et al., 2004). The S-locus consists of two closely linked genes, coding for two recognition proteins: the S-locus receptor kinase (SRK) expressed at the stigmatic surface and the S-locus cystein rich protein (SCR) expressed at the pollen surface which acts as a ligand to the SRK. The SRK and SCR of an S-allele express the same self-incompatibility type. If the self-incompatibility type of the pollen matches with that of the stigma, a self-incompatibility response will occur and the pollen will not germinate on the stigma. The S-locus is under strong, negative frequency-dependent selection, because rare self-incompatibility types have a mating advantage compared to more common ones (Wright, 1939). The S-locus has been shown to reside in a region of the genome with very low recombination rates (Casselman et al., 2000; Kamau and Charlesworth, 2005; Hagenblad et al., 2006). The low recombination rate is thought to have evolved to maintain the integrity of the system, because recombination between SRK and SCR would lead to a nonfunctional and self-compatible haplotype.
Theory predicts that genomic regions with low recombination and under balancing selection may accumulate a genetic load within an allelic class, since frequency-dependent selection will shelter them when their frequency gets low in the population (Uyenoyama, 2005). It is therefore possible that some S-haplotypes have accumulated a more severe genetic load than others, which will lead to segregation disadvantage. Yet, they are not expected to be lost easily as long as this disadvantage is offset by the mate availability advantage of a rare S-haplotype.
Bechsgaard et al. (2004) investigated the segregation of the S-haplotypes in A. lyrata and reported that in 6 of 19 (31.5%) crosses, significant segregation distortion was found between the S-haplotypes of the parental plants. Certain S-alleles consistently enjoyed a transmission advantage against others. This was interpreted as accumulation of a dominant genetic load linked to the S-locus in some S-haplotypes. Since only gametic selection could be demonstrated, it was assumed to be at the haploid stage. It was furthermore suggested that the S-locus may act as a trap for factors causing meiotic drive. Meiotic drive variants are generally expected to be rapidly fixed in a species, but if they are completely linked to an S-haplotype they will only be fixed within this S-haplotype and thus remain polymorphic in the species.
It is also possible that segregation in A. lyrata is very distorted throughout the genome for unknown reason. The distortion seen at the S-locus, or when crossing diverged populations, could just reflect this general high distortion.
There are few previous studies of deviations from Mendelian segregation in crosses within a single plant population. In conifers mapping with haploid megagametophytes has revealed segregation distortion. The within-species level of segregation distortion in the markers is highly variable in different species—from 2 to 79% of markers exhibit distortion, on average 20% (Krutovskii et al., 1998, and references therein). This high level of segregation distortion has been explained by high genetic loads in conifers. Direct observation of segregation of gametes is not feasible in angiosperms. Progeny arrays from crosses can be used, but gametic and zygotic selection can be difficult to disentangle (see below). Most results on transmission ratio distortion within populations derive from studies on inbreeding depression. The crosses have typically been conducted between highly divergent lines of cultivated species or outcrossing plants have been selfed (Carr and Dudash, 2003 and references therein). In a study of selfed progenies of A. lyrata deviations of marker segregation from Mendelian segregation were assumed to be due to linkage to viability loci (Kärkkäinen et al., 1999). That study however did not describe the general level of transmission ratio distortion in outcrossing A. lyrata populations.
To test whether there is an excess of transmission ratio distortion in the cross between diverged populations and intrapopulation crosses, and the S-locus compared to the rest of the genome, we examined Mendelian segregation throughout the genome in intrapopulation crosses by using some of the progeny arrays of Bechsgaard et al. (2004). We present data on the segregation of 14 microsatellites and 2 length polymorphisms in 8 of the families. Of these, 13 markers were also used in the interpopulation study by Kuittinen et al. (2004).
More specifically we address the following questions: (1) what is the proportion of distorted loci in the intrapopulation crosses, (2) is the proportion of distorted loci higher in the interpopulation cross than in the intrapopulation crosses, (3) does selection act at zygotic or gametic level and is there a difference between inter- and intrapopulation crosses and (4) is the level of distortion in the S-locus higher than the general level in the genome?
Materials and methods
Materials
We investigated segregation from intrapopulation crosses in eight full-sib families from an Icelandic population previously used by Bechsgaard et al. (2004). The parental plants originated from a single Icelandic population near Reykjavik named Mt. Esja. Plants from the parental population were denoted by prefix ‘00B’. Three of the eight families are from crosses between half sibs (families 8, 12 and 13), and two of the parental plants were used in several crosses (00B 17/3 to produce families 12, 13 and 14, and 00B 27/3 to produce families 6 and 23).
In total 404 plants were used in the current study. The number of offspring in each family varied between 26 and 70. The numbers in each family were as follows: family 6: 30 (from only one reciprocal cross), family 8: 49 (22 in reciprocal 1 and 27 in reciprocal 2), family 12: 60 (30 and 30), family 13: 53 (36 and 17), family14: 26 (16 and 10), family 21: 70 (25 and 45), family 23: 66 (13 and 53) and family 31: 36 (from only 1 reciprocal). Plants from the offspring population were denoted by prefix ‘01B’. For details about growth conditions see Bechsgaard et al. (2004).
Genotyping
A total of 30 loci were screened for polymorphisms from the 13 parental plants of the 8 families. Fourteen microsatellites and two indel polymorphisms located on all eight chromosomes were chosen because of their high polymorphism in the parental plants for complete genotyping in the families. The number of plants genotyped varies between loci because some loci were monomorphic in some families. The genetic distance between loci located on the same chromosome varied from 10 to 50 cM, based on the Karhumäki–Mjällom map (Kuittinen et al., 2004). The genomic locations of the markers are shown in Figure 1. Four of these loci were described earlier by Bell and Ecker (1994), eight by Clauss et al. (2002) and four by Kuittinen et al. (2004). Typing was performed according to the following protocol. DNA extraction was done in a previous study by Bechsgaard et al. (2004). Loci were amplified in a 10 μl PCR reaction containing 1 × Taq buffer, 2.5 mM MgCl2, 0.1 μM of each deoxyribonucleotide triphosphate, 0.25 μM of unlabeled primer, 0.25 μM of 5′ labeled primer (either FAM, PET, NED or VIC), 0.5 U Taq polymerase (Promega, Madison, WI, USA) and approximately 3 ng of template. Amplification profile was as following: 3 min denaturation at 94 °C, followed by 35 cycles of 20 s at 94 °C, 30 s at 50 °C and 10 s at 72 °C. Final extension was 45 min at 72 °C. Standard agarose gel electrophoresis was done to check the results from amplification and to be able to estimate the dilution for a capillary electrophoresis run. Dilutions from the PCR products were multiplexed and mixed with 10 μl of Hi-Di formamide and 0.2 μl of GeneScan 500 LIZ size standard (Applied Biosystems, Warrington, UK) for fragment analysis on an ABI 3730 DNA Analyzer. Sizes of the fragments were determined by GeneMapper software v. 3.7 (Applied Biosystems).
Data analysis
The maximum likelihood framework of Bechsgaard et al. (2004) was used to analyze the segregation results of the present study and to reanalyze the data from Bechsgaard et al. (2004) and Kuittinen et al. (2004). For each reciprocal cross, there are either four genotypic classes (type I: ab × cd or ab × ac), three genotypic classes (type II: ab × ab) or two genotypic classes (type III: aa × ab or aa × bc). Fully parameterized, type I has three free parameters, type II has two free parameters and type III has one free parameter (full model). Deviations from Mendelian expectation of genotype frequencies (type I and type II) were tested (the genotype test) by comparing the full model with a model constrained to equal genotype frequencies (type I) and 1:2:1 frequencies (type II) (Mendelian model (1)) using a standard likelihood ratio test. The test statistic is χ2 distributed with three d.f. (type I) or two d.f. (type II). Gametic selection can be demonstrated by testing the segregation of the alleles from each parent separately (gametic model) against Mendelian expectations (Mendelian model (2)), resulting in a χ2 (one d.f.) test (allele test). The segregation models and the tests used are summarized in Tables 1a and b. In type II crosses, it is not possible to resolve from which parent the alleles in the heterozygote class come from. To examine gametic selection we tested for deviation from Mendelian expectations considering total numbers of each allele in the offspring (the number of allele a: 2 × the number of genotype aa+the number of genotype ab, and the number of allele b: 2 × the number of genotype bb+the number of genotype ab).
For types I and II, a nested model assumes that any deviation from Mendelian segregation is due to distortion in the parents (gametic selection) and thus has only two free parameters (type I) or one free parameter (type II) (under the assumption that the alleles segregate equally in male and female gametophytes). Thus, interaction between the two parents, which we define as zygotic selection, can be tested by comparing the full model and the gametic model using a standard likelihood ratio test (zygotic test). The test statistic is χ2 distributed with one d.f., corresponding to the difference in the number of free parameters. We also tested whether the segregation patterns were equal in the two reciprocals by comparing a model that assumes different selection coefficients in the two reciprocals to a model assuming the same selection coefficient. This was done first by assuming that the selection was gametic (types I, II and III), and then zygotic (types I and II). All calculations were carried out using Mathematica (version 4.1) (Wolfram Research, Champaign, IL, USA).
Resampling the mapping family and S-locus data
The data set from Kuittinen et al. (2004) consists of a single family genotyped for 72 markers in both reciprocals. Between 35 and 99 individuals were genotyped in the first reciprocal and between 43 and 104 in the second reciprocal. The data set from Bechsgaard et al. (2004) consists of 11 families genotyped at the S-locus. The families consist of between 30 and 75 individuals. Due to larger sample sizes, the power to detect transmission ratio distortion in these two data sets is generally greater than in the intrapopulation crosses of the present study. To attain comparable levels of power, we resampled the data from Kuittinen et al. (2004) and Bechsgaard et al. (2004). The genotypes of 25 individuals were drawn without replacement from each reciprocal and tested using the maximum likelihood framework, and this process was repeated 1000 times. Since most of the intrapopulation crosses have more than 25 individuals genotyped in each reciprocal (average of 29.5), our comparison is conservative against the hypothesis that there is a difference in levels of transmission ratio distortion between inter- and intrapopulation crosses. Resampling was carried out using Mathematica (version 4.1) (Wolfram Research).
Results
Intrapopulation crosses
The full genotypic data at 14 microsatellite loci and 2 length polymorphisms in 8 families are supplied as Supplementary material. In total, the data consist of about 5300 genotypes with an average of more than 50 individuals per family. We performed 61 tests for deviations from Mendelian genotype frequencies (genotype test). Three of these tests (4.9%) deviated significantly from Mendelian segregation. A total of 192 tests were performed for deviations of segregation of each parental allele combination (allele test), and 21 (10.9%) were significantly distorted (Table 2).
All crosses of types I and II were examined for selection at the zygotic stage (zygotic test). In only 2 of 61 tests (3.3%) was a significant zygotic effect demonstrated (Table 2). Thus, gametic selection appears generally sufficient to explain the observed patterns of transmission ratio distortion. We also looked for evidence that some alleles were preferentially transmitted, but no obvious patterns were found between loci or families. The numbers and proportions of tests that significantly deviate from expected within each family and for each locus are presented in Tables 3a and b, respectively.
Only 3.9% (2/51) of the tests showed different segregation patterns in the reciprocals, assuming only gametic selection. The test was not done assuming zygotic selection, since zygotic selection seems to be weak (3.3%, see above).
Since one parental plant was used in three of the crosses and one was used in two of the crosses, the tests are not all independent. The alleles from 00B 17/3 in families 12, 13 and 14, and from 00B 27/3 in families 6 and 23 segregated in the same genetic background in the meiosis. This could influence our conclusions, if they were all distorted or not distorted. However, the segregation of these alleles does not show any consistent patterns among families, and about 10% of the alleles deviated from expected ratios.
Interpopulation cross
Segregation data (Kuittinen et al., 2004) were first resampled to obtain comparable sample sizes to the intrapopulation crosses described above (see ‘Materials and methods’ section). For the resampled data set, 20.1% of the genotype tests was significant and 19.1% of the allele tests was significant (Table 2). These fractions are much higher than the observed fractions in the intrapopulation crosses (4.9 and 10.9%, respectively).
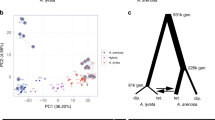
The larger number of markers in the interpopulation cross compared to intrapopulation crosses (72 and 16 markers, respectively) did not have an effect on the results. When comparing the 13 common markers (3 of the markers genotyped in the intrapopulation crosses were not genotyped in the interpopulation cross) between the crosses the percentages of significant tests are very close to the numbers reported for the full data set (data not shown). The differences at the allelic level between the crosses for the set of common markers are visualized in Figure 2.
Proportions of the allele tests significantly deviating from Mendelian ratios in the 13 common markers between the intra- and interpopulation crosses. For the interpopulation cross the proportions of the resampled data distorted are presented. Dark and light gray represent the first and the second reciprocal from the interpopulation cross, respectively, and white the intrapopulation crosses. The intrapopulation crosses cannot be split into first and second reciprocal, because the results are based on combining several families.
The loci segregating like types I and II were examined for a selective effect at the zygotic stage. In 10.5% of the tests a zygotic effect could be detected, which is about three times higher than in the intrapopulation crosses.
Furthermore, the full data of the 28 markers (of types I and II) reported to be distorted by Kuittinen et al. (2004) (7 of them in both reciprocals) were reanalyzed in the likelihood framework in order to disentangle gametic effects from zygotic effects (zygotic test). Altogether 35 tests were carried out when reciprocals were analyzed separately. Table 4 shows that in 27 of the 35 tests a separate gametic effect could be detected. In five of those, a zygotic effect could also be detected. In seven markers a zygotic effect only could be detected, and in one of the distorted markers neither a gametic nor a zygotic effect could be detected. That is, in one-third (12 of 35) of the tests, a significant zygotic effect could be demonstrated.
In the full data set (Kuittinen et al., 2004) 15.3% (11/72) of the markers differed between reciprocals when assuming gametic selection as the only force shaping the segregation patterns, and 18.5% (10/54) when assuming zygotic selection as the only force shaping the segregation patterns.
S-locus segregation
The segregation data from the S-locus (Bechsgaard et al., 2004) was first resampled to get comparable sample sizes to the intrapopulation crosses described above (see ‘Materials and methods’ section). For the resampled data set 32.2% of the 8 resampled families deviated from Mendelian expectation at the genotype level (Table 2). A total of 19 allele combinations (16 from type I and 3 from type III) were tested, and 17.7% were distorted (Table 2). These proportions are also much higher than observed in the intrapopulation crosses (4.9 and 10.9%, respectively). In 3.1% of the tests a zygotic effect was detectable (zygotic test).
Discussion
Intrapopulation crosses
At the 5% significance level, 10.9% of the tests at the allelic level deviated from the 1:1 expectation. This is an appreciable number of distortions considering that the size of progeny arrays limits the statistical power to detect weak distortion.
One possible explanation for this finding is meiotic drive, which has been demonstrated in several studies (for example, Fishman and Willis, 2005). Typically alleles causing meiotic drive would rapidly go to fixation in a population. If meiotic drive causes the transmission ratio distortion observed in this study, antagonistic deleterious effects must be present to prevent fixation. Strong deleterious effects have often been observed to be associated with segregation distorters, but whether this is a common phenomenon or just a bias in the detection of segregation distorters, due to rapid fixation without antagonistic deleterious effects, is not known (Taylor and Ingvarsson, 2003).
An alternative explanation could be inbreeding depression, since crosses were done between individuals from a single population. In a study of transmission ratio distortion with selfing in A. lyrata, some loci did show significant deviations from the expected ratio (Kärkkäinen et al., 1999). If both parents carry a given microsatellite allele linked to a deleterious recessive allele, this may result in a deviation from Mendelian expectations at both allele and genotype level (Hedrick and Muona, 1990). There is an indication that this is not the case in our results. If the three families with half-sib parents are surveyed and compared with the rest of the families, there is no difference in the proportion distorted (10/105 (half sib) vs 11/98 for allele tests, and 1/31 (half sib) vs 2/32 for genotype tests). Since the relationship between the parental plants is unknown it is of course possible that the ‘unrelated’ crosses are in fact genetically related as well.
This appreciable number of distortions at the gametic level supports the idea that meiotic drive is a powerful evolutionary force. It fixes alleles rapidly (which we most likely do not observe) or results in genetic conflict by creating antagonistic selection at other loci. Transmission ratio distortion at the gametic level might also be due to postmeiotic differences in the fitness of the gametes.
Zygotic selection seems to play a minor role in shaping the segregation pattern of the intrapopulation crosses. Only 3.3% of the tests were significant, which is not more than expected by chance. Most deleterious alleles at loci expressed postzygotically are recessive, and are thereby masked by dominant alleles. Deleterious alleles at loci expressed prezygotically on the other hand are not masked, and will cause transmission ratio distortion.
Since most of the markers were not distorted, it is not surprising that only two tests (3.9%) showed different reciprocal segregation. One of them (AGL20 in family 8) was due to one reciprocal being highly distorted, whereas for the other (F20D22 in family 12) neither of the reciprocals differed significantly from the expected ratios, but the reciprocals were slightly distorted in opposite directions.
Interpopulation cross
In the interpopulation cross, genotype tests, the proportion of loci exhibiting transmission ratio distortion was about four times as high as in the intrapopulation crosses. For the allele tests the difference was twofold when comparing the interpopulation cross with the intrapopulation crosses (Table 2). A biological explanation for the difference in transmission ratio distortion at the allelic level could be that segregation is more severely distorted in the Karhumäki and the Mjällom populations than in the Icelandic population, where the intrapopulation crosses originate from. A second alternative biological explanation could be that meiotic drive has fixed the alleles causing segregation distortion within populations. This would not lead to segregation distortion in intrapopulation crosses, but might lead to segregation distortion in interpopulation crosses at the gametic level (but not zygotic), when alleles causing meiotic drive become heterozygous.
In the intrapopulation crosses, transmission ratio distortion could barely be detected at the zygotic and genotypic level. Only 3.3 and 4.9% of the tests, respectively, showed a significant deviation from the expected. In the between-population cross, on the other hand, a sign of zygotic selection was detectable in 10.5% of the loci and 20.1% of the tests were significant at the genotypic level (Table 2). Interactions at the diploid stage therefore play a large role in the between-population cross. One explanation for the higher zygotic selection seen in the interpopulation cross could be epistatic interactions (Dobzhansky–Muller interactions), where individuals homozygous for alternative parental alleles at different loci have reduced fitness. To confirm this, further studies on interactions between markers should be conducted in between-population crosses.
Several loci segregated differently in the two reciprocals in the interpopulation cross at both gametic and zygotic level. This could be due to nuclear–cytoplasmic interactions. In the Karhumäki–Mjällom cross the reciprocals had different cytoplasms and it is feasible that cytoplasmic organelles originating from the Karhumäki population did not function well with some nuclear genes or alleles from the Mjällom population. This could have led to abortion of gametes carrying certain nuclear alleles or abortion of zygotes, but only with specific cytoplasmic background. There could also have been selection for particular allele combinations during gamete formation of the two different F1 parents. Interactions between pollen and stigma—leading to fertilization—are also known to be complicated (for example, reviewed by Chaudhury et al., 2001; Boavida et al., 2005). Pollen performance can vary between different genotypes and might lead to nonrandom fertilization (Snow and Spira, 1991). This could lead to differences between reciprocals, because only alleles from pollen donors have been affected by selection, not alleles from maternal parents.
S-locus segregation
The level of transmission ratio distortion observed in this study is also much lower than observed when reanalyzing the data on the S-locus from Bechsgaard et al. (2004). We interpret this as an inherently high level of transmission ratio distortion of the S-locus compared to the rest of the genome. This is consistent with theoretical predictions that genomic regions with low recombination rates harboring loci under negative frequency-dependent selection have the potential to accumulate deleterious mutations (Uyenoyama, 2005). Thus, it is likely that the complex organization and large physical extent of S-haplotypes with different number of genes (Sherman-Broyles et al., 2007) have fitness effects that may be elucidated by comparative sequencing of complete S-haplotypes.
Accumulation of genetic load within S-haplotypes will influence both the evolutionary dynamics of the S-locus and the frequencies of S-alleles in natural populations. Even though a genetic load will be counteracted by frequency-dependent selection, S-haplotypes with low frequencies are more exposed to drift, and thereby more likely to be lost. Nevertheless, the extent to which differential accumulation of genetic load will influence the dynamics of the S-locus and frequencies of the S-haplotypes is not clear. More segregation studies and simulation studies are needed to be able to set up predictions on, for example, S-haplotype frequencies, which could then be experimentally tested.
In summary, it was found that in intrapopulation crosses of A. lyrata, transmission ratio distortion at genotypic and zygotic level did not appear more often than expected by chance. At the gametic level, transmission ratio distortion was found more often than expected. This may be due to meiotic drive or selection acting postmeiotically. The proportion of distorted loci was higher at every level when comparing the interpopulation cross to the intrapopulation crosses. Selection seems to be stronger especially at zygotic level in the interpopulation cross. This could be due to Dobzhansky–Muller incompatibilities that have accumulated between the diverging populations crossed. The segregation distortion of the S-locus is shown to be substantially higher than for molecular markers suggesting different selection pressures at S-haplotypes potentially due to genetic load linked to the S-locus.
References
Bechsgaard J, Bataillon T, Schierup MH (2004). Uneven segregation of sporophytic self-incompatibility alleles in Arabidopsis lyrata. J Evol Biol 17: 554–561.
Bechsgaard JS, Castric V, Charlesworth B, Vekemans X, Schierup MH (2006). The transition to self-compatibility in Arabidopsis thaliana and evolution within S-haplotypes over 10 Myr. Mol Biol Evol 23: 1741–1750.
Bell CJ, Ecker JR (1994). Assignment of 30 microsatellite loci to the linkage map of Arabidopsis. Genomics 19: 137–144.
Boavida LC, Vieira AM, Becker JD, Feijo JA (2005). Gametophyte interaction and sexual reproduction: how plants make a zygote. Int J Dev Biol 49: 615–632.
Carr DE, Dudash MR (2003). Recent approaches into the genetic basis of inbreeding depression in plants. Phil Trans R Soc Lond B 358: 1071–1084.
Casselman AL, Vrebalov J, Conner JA, Singhal A, Giovannoni J, Nasrallah ME et al (2000). Determining the physical limits of the Brassica S locus by recombinational analysis. Plant Cell 12: 23–33.
Chaudhury AM, Koltunow A, Payne T, Luo M, Tucker MR, Dennis ES et al (2001). Control of early seed development. Annu Rev Cell Dev Biol 17: 677–699.
Clauss MJ, Cobban H, Mitchell-Olds T (2002). Cross-species microsatellite markers for elucidating population genetic structure in Arabidopsis and Arabis (Brassicaceae). Mol Ecol 11: 591–601.
Clauss MJ, Koch MA (2006). Poorly known relatives of Arabidopsis thaliana. Trends Plant Sci 11: 449–459.
Clauss MJ, Mitchell-Olds T (2006). Population genetic structure of Arabidopsis lyrata in Europe. Mol Ecol 15: 2753–2766.
Fishman L, Willis JH (2001). Evidence for Dobzhansky-Muller incompatibilities contributing to the sterility of hybrids between Mimulus guttatus and M. nasutus. Evolution 55: 1932–1942.
Fishman L, Willis JH (2005). A novel meiotic drive locus almost completely distorts segregation in Mimulus (monkeyflower) hybrids. Genetics 169: 347–353.
Hagenblad J, Bechsgaard J, Charlesworth D (2006). Linkage disequilibrium between incompatibility locus region genes in the plant Arabidopsis lyrata. Genetics 173: 1057–1073.
Hall MC, Willis JH (2005). Transmission ratio distortion in intraspecific hybrids of Mimulus guttatus: implications for genomic divergence. Genetics 170: 375–386.
Hedrick PW, Muona O (1990). Linkage of viability genes to marker loci in selfing organisms. Heredity 64: 67–72.
Jenczewski E, Gherardi M, Bonnin I, Prosperi JM, Olivieri I, Huguet T (1997). Insight on segregation distortions in two intraspecific crosses between annual species of Medicago (Leguminosae). Theor Appl Genet 94: 682–691.
Jonsell B, Kustås K, Nordal I (1995). Genetic variation in Arabis petraea, a disjunct species in northern Europe. Ecography 18: 321–332.
Kamau E, Charlesworth D (2005). Balancing selection and low recombination affect diversity near the self-incompatibility loci of the plant Arabidopsis lyrata. Curr Biol 15: 1773–1778.
Kärkkäinen K, Kuittinen H, van Treuren R, Vogl C, Oikarinen S, Savolainen O (1999). Genetic basis of inbreeding depression in Arabis petraea. Evolution 53: 1354–1365.
Korbecka G, Klinkhamer PGL, Vrieling K (2002). Selective embryo abortion hypothesis revisited—a molecular approach. Plant Biol 4: 298–310.
Krutovskii KV, Vollmer SS, Sorensen FC, Adams WT, Knapp SJ, Strauss SH (1998). RAPD genome maps of Douglas-fir. J Hered 89: 197–205.
Kuittinen H, de Haan AA, Vogl C, Oikarinen S, Leppälä J, Koch M et al (2004). Comparing the linkage maps of the close relatives Arabidopsis lyrata and A. thaliana. Genetics 168: 1575–1584.
Kusaba M, Dwyer K, Hendershot J, Vrebalov J, Nasrallah JB, Nasrallah ME (2001). Self-incompatibility in the genus Arabidopsis: characterization of the S locus in the outcrossing A. lyrata and its autogamous relative A. thaliana. Plant Cell 13: 627–643.
Mitchell-Olds T (2001). Arabidopsis thaliana and its wild relatives: a model sytem for ecology and evolution. Trends Ecol Evol 16: 693–700.
Muller M-H, Leppälä J, Savolainen O (2007). Genome wide effects of post-glacial colonization in Arabidopsis lyrata: reduced variation in Northern populations and high divergence. Heredity (e-pub ahead of print; doi:10.1038/sj.hdy.6801057).
Myburg AA, Vogl C, Griffin AR, Sederoff RR, Whetten RW (2004). Genetics of postzygotic isolation in Eucalyptus: whole-genome analysis of barriers to introgression in a wide interspecific cross of Eucalyptus grandis and E. globulus. Genetics 166: 1405–1418.
Nasrallah ME, Liu P, Nasrallah JB (2002). Generation of self-incompatible Arabidopsis thaliana by transfer of two S locus genes from A. lyrata. Science 297: 247–249.
Riihimäki M, Savolainen O (2004). Environmental and genetic effects on flowering differences between northern and southern populations of Arabidopsis lyrata (Brassicaceae). Am J Bot 91: 1036–1045.
Schierup MH, Mable BK, Awadalla P, Charlesworth D (2001). Identification and characterization of a polymorphic receptor kinase gene linked to the self-incompatibility locus of Arabidopsis lyrata. Genetics 158: 387–399.
Sherman-Broyles S, Boggs N, Farkas A, Liu P, Vrebalov J, Nasrallah ME et al (2007). S locus genes and the evolution of self-fertility in Arabidopsis thaliana. Plant Cell 19: 94–106.
Snow AA, Spira TP (1991). Pollen vigor and the potential for sexual selection in plants. Nature 352: 796–797.
Taylor DR, Ingvarsson PK (2003). Common features of segregation distortion in plants and animals. Genetica 117: 27–35.
Törjék O, Witucka-Wall H, Meyer RC, von Korff M, Kusterer B, Rautengarten C et al (2006). Segregation distortion in Arabidopsis C24/Col-0 and Col-0/C24 recombinant inbred line populations is due to reduced fertility caused by epistatic interaction of two loci. Theor Appl Genet 113: 1551–1561.
Uyenoyama MK (2005). Evolution under tight linkage to mating type. New Phytol 165: 63–70.
van Treuren R, Kuittinen H, Kärkkäinen K, Baena-Gonzalez E, Savolainen O (1997). Evolution of microsatellites in Arabis petraea and A. lyrata, outcrossing relatives of Arabidopsis thaliana. Mol Biol Evol 14: 220–229.
Wright S (1939). The distribution of self-sterility alleles in populations. Genetics 24: 538–552.
Yin TM, DiFazio SP, Gunter LE, Riemenschneider D, Tuskan GA (2004). Large-scale heterospecific segregation distortion in Populus revealed by a dense genetic map. Theor Appl Genet 109: 451–463.
Zamir D, Tadmor Y (1986). Unequal segregation of nuclear genes in plants. Bot Gaz 147: 355–358.
Acknowledgements
We thank Helmi Kuittinen and Thomas Bataillon for comments and suggestions, and Thomas Bataillon for his help to develop the Mathematica notebook to do the resampling. The study was supported by grant no. 107167 from the Biosciences and Environment Research Council to OS (JL and OS) and by grant no. 2052-01-0032 from the Danish Agricultural Sciences Research Council (MHS).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Supplementary Information accompanies the paper on Heredity website (http://www.nature.com/hdy)
Supplementary information
Rights and permissions
About this article
Cite this article
Leppälä, J., Bechsgaard, J., Schierup, M. et al. Transmission ratio distortion in Arabidopsis lyrata: effects of population divergence and the S-locus. Heredity 100, 71–78 (2008). https://doi.org/10.1038/sj.hdy.6801066
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6801066
Keywords
This article is cited by
-
The recombination landscape in Arabidopsis thaliana F2 populations
Heredity (2012)
-
An EST-derived SNP and SSR genetic linkage map of cassava (Manihot esculenta Crantz)
Theoretical and Applied Genetics (2012)