Abstract
Surveys of gene expression reveal extensive variability both within and between a wide range of species. Compelling cases have been made for adaptive changes in gene regulation, but the proportion of expression divergence attributable to natural selection remains unclear. Distinguishing adaptive changes driven by positive selection from neutral divergence resulting from mutation and genetic drift is critical for understanding the evolution of gene expression. Here, we review the various methods that have been used to test for signs of selection in genomic expression data. We also discuss properties of regulatory systems relevant to neutral models of gene expression. Despite some potential caveats, published studies provide considerable evidence for adaptive changes in gene expression. Future challenges for studies of regulatory evolution will be to quantify the frequency of adaptive changes, identify the genetic basis of expression divergence and associate changes in gene expression with specific organismal phenotypes.
Similar content being viewed by others
Introduction
When and where a gene is expressed, and how much of the gene product is made, can be as important as the biochemical function of the encoded RNA or protein (Raff, 1996; Carroll et al., 2001; Davidson, 2001). In 1969, Britten and Davidson (1969) proposed a theory of gene regulation for eukaryotic cells that included a central role for regulatory divergence in phenotypic evolution. Subsequent comparisons of molecular and morphological phenotypes indicated that protein divergence was insufficient to account for the extensive phenotypic differences observed between species, and prompted the proposal that many adaptations may have arisen from changes in gene regulation rather than from changes in gene function (Wilson et al., 1974; King and Wilson, 1975). Over the last 30 years, molecular studies of development and evolution, combined with studies of experimental evolution, have provided strong support for this hypothesis (for example Wray et al., 2003 and Gompel et al., 2005). Nonetheless, the relative importance of regulatory changes versus changes in protein function remains subject to debate.
Genetic and transgenic experiments have shown that changes in gene regulation often underlie morphological differences between species. Examples include changes in pelvic structures in threespine sticklebacks mediated by Pitx1 (Shapiro et al., 2004); trichome patterning in Drosophila mediated by Ubx (Stern, 1998); larval hairs in Drosophila mediated by ovo/shaven-baby (Sucena and Stern, 2000); pigmentation in Drosphila mediated by bric-a-brac (Kopp et al., 2000), yellow (Wittkopp et al., 2002; Gompel et al., 2005) and ebony (Wittkopp et al., 2003); butterfly eyespots mediated by Distal-less (Beldade et al., 2002) and beak size among Galapagos finches mediated by BMP4 (Abzhanov et al., 2004).
Experimental evolution in microorganisms also provides compelling evidence that regulatory evolution contributes to phenotypic divergence. Parallel mutations observed in replicate populations are likely to have been fixed by positive selection (Orr, 2005). Parallel expression divergence has been reported for experimental populations of both Saccharomyces cerevisiae and Escherichia coli (Ferea et al., 1999; Cooper et al., 2003; Riehle et al., 2003; Fong et al., 2005). In two populations of E. coli evolving independently for 20 000 generations in glucose-limited media, 59 genes acquired similar expression changes in both populations (Cooper et al., 2003). Similarly, three strains of S. cerevisiae grown in glucose-limited media for 250 generations showed over 50 genes whose expression changed in parallel in all three lines, suggesting that these changes were adaptive (Ferea et al., 1999).
Finally, studies elucidating the molecular basis of adaptations in domesticated crops also indicate a significant role for regulatory evolution in phenotypic evolution. For example, in maize, changes in expression of the teosinte branched 1 (tb1) gene have evolved since it shared a common ancestor with teosinte (Doebley et al., 1997). Mutations affecting tb1 expression alter branching, consistent with morphological differences between domesticated corn and its wild relative (Hubbard et al., 2002). Evidence of directional selection has been found in the upstream cis-regulatory regions of noncoding region of tb1, suggesting adaptive changes in expression of this gene (Clark et al., 2006). Regulatory changes have also been implicated in the evolution of naked grains in maize (Wang et al., 2005) and in the loss of seed shattering in rice (Konishi et al., 2006).
These compelling examples illustrate that regulatory changes can underlie adaptations. But, do all changes in gene expression contribute to phenotypic evolution and how often is regulatory divergence adaptive? One way to obtain insight into this question is by using DNA microarrays to examine genomic patterns of gene expression within and between species and using statistical methods to distinguish neutral and nonneutral patterns of gene expression divergence. Indeed, methods originally developed to detect signs of selection on morphological characters and DNA sequences have now been applied to genomic expression data. Here, we review these tests, their applications and assumptions of the underlying models. We discuss features of regulatory evolution that may complicate the interpretation of these tests, including correlated changes in expression among genes, the relationship between genotype and gene expression and the dependency of mutational effects on current expression levels. We conclude by anticipating future directions for studies investigating the evolution of gene expression, including the identification of specific phenotypes affected by genes showing evidence of adaptive regulatory divergence.
Genomic variation in gene expression
Genome-wide measurements have revealed high rates of genetic variation in gene expression (typically >10% of genes) in humans (Enard et al., 2002; Rockman and Wray, 2002; Bray et al., 2003; Lo et al., 2003; Whitney et al., 2003; Morley et al., 2004; Pastinen et al., 2004; Radich et al., 2004), mice (Cowles et al., 2002; Schadt et al., 2003; Shockley and Churchill, 2006), fish (Oleksiak et al., 2002, 2005), flies (Jin et al., 2001; Wayne and McIntyre, 2002; Meiklejohn et al., 2003; Rifkin et al., 2003; Nuzhdin et al., 2004; Ranz et al., 2004), yeast (Cavalieri et al., 2000; Brem et al., 2002; Townsend et al., 2003; Yvert et al., 2003; Fay et al., 2004), plants (Kirst et al., 2005; Vuylsteke et al., 2005; Lai et al., 2006) and bacteria (Le et al., 2005). Patterns of expression divergence have also been compared between sexes (Jin et al., 2001; Ranz et al., 2003; Gibson et al., 2004), across developmental stages (Rifkin et al., 2003), among tissue types (Enard et al., 2002; Whitehead and Crawford, 2005; Khaitovich et al., 2005a, 2005b) and over different environments (Fay et al., 2004; Landry et al., 2006).
The genetic architecture of population genetic variation in gene expression has also been examined in a number of organisms. Using standard quantitative genetics methods, gene expression has been shown to be a heritable, often polygenic, trait (Brem et al., 2002; Schadt et al., 2003; Monks et al., 2004; Brem and Kruglyak, 2005; Cheung et al., 2005). Like many other quantitative traits, variation in gene expression shows evidence of dominance and nonadditive (epistatic) interactions among loci (Gibson et al., 2004; Brem et al., 2005; Storey et al., 2005; Vuylsteke et al., 2005). Both cis- and trans-acting regulatory variants contribute significantly to regulatory variation within a species (Brem et al., 2002; Schadt et al., 2003; Yvert et al., 2003; Monks et al., 2004; Morley et al., 2004; Wayne et al., 2004; Doss et al., 2005; Kirst et al., 2005; Ronald et al., 2005). Although regulatory variation is clearly abundant within populations, its evolutionary significance is harder to ascertain.
The role of selection and genetic drift in the evolution of gene expression
Evolution has many causes. Darwin (1876) poignantly argued that evolution is caused by natural selection. Since then, considerable efforts have been made to examine an alternative explanation: genetic drift, or the stochastic changes that occur as a consequence of finite population size (Kimura, 1983). Three primary methods have been used to distinguish between these possibilities. First, the comparative method examines the evolution of a character in relation to the evolution of other characters or environmental variables in a phylogenetic context (Martins, 2000). Second, neutral models of evolutionary change can be used to test for selection on quantitative characters (Turelli et al., 1988). Finally, the empirical rate of expression divergence across lineages can be used to distinguish between selection and drift (Rifkin et al., 2005). Each of these methods has recently been applied to gene expression data with the hope of addressing the long-standing idea that changes in gene regulation play a substantial role in adaptive evolution.
Comparative methods
The comparative method is based on the correlated evolution of a phenotype with some other trait or environmental variable. Two classic examples are the association between white coat coloration and animals living in snowy environments, and between large testis size and polygynous primate species (Harvey and Pagel, 1991). Although the repeated evolution of a trait is indicative of adaptation, correlated patterns of change can also arise for other reasons. First, shared ancestry causes correlations among characters in the absence of selection (Felsenstein, 1985). If the evolutionary history of the species under study is known, it can be taken into account by using independent contrasts that examine subsets of the tree (Felsenstein, 1985), by partitioning phenotypic changes into shared and unique regions of the lineage (Cheverud et al., 1985) or by using statistical techniques such as regression and general linear models (Grafen, 1989; Martins and Hansen, 1997; Rohlf, 2001). If there is uncertainty in the phylogeny, a Bayesian approach can be used to integrate overall plausible evolutionary histories (Huelsenbeck and Rannala, 2003). Second, correlations between characters can arise as a result of genetic, developmental or environmental constraints, unrelated to natural selection (Arnold, 1992). This is of particular concern for gene expression data, since the a priori expectation is that many changes in gene expression will be correlated with each other or with other phenotypes (for other issues related to the comparative method, see Harvey and Purvis, 1991; Martins and Garland, 1991; Diaz-Uriarte and Garland, 1998; Martins, 2000).
Comparative models
As described above, multiple approaches have been developed to account for phenotypic correlations resulting from phyogenetic relationships among taxa (Cheverud et al., 1985; Felsenstein, 1985; Grafen, 1989; Martins and Hansen, 1997; Rohlf, 2001). While most of these methods do not posit a particular model of phenotypic evolution, the Brownian motion model satisfies many of their assumptions. The Brownian motion model describes the evolution of a phenotype without specifying an explicit genetic model (Felsenstein, 1988). The model assumes (1) a Gaussian (normal) distribution of phenotypes within a population and (2) constant genetic variance regardless of the mean value of the trait (Lande, 1976). The first assumption holds when a large number of unlinked loci make an equally small and independent contribution to a trait. Although this is a common genetic model used in quantitative genetic theory, regulatory networks have a scale-free structure (that is, few genes with many connections and many genes with few connections) that suggests this is not an adequate description of gene regulation. Indeed, empirical studies have shown that some genetic changes have more widespread effects on gene expression than others (Brem et al., 2002; Yvert et al., 2003). The second assumption holds when the phenotypic value of a trait is unconstrained. The validity of this assumption is also a concern since many traits cannot evolve indefinitely without reaching some form of genetic, developmental or physical constraint (Arnold, 1992; Diaz-Uriarte and Garland, 1998). Gene expression levels are undoubtedly limited, which may cause them to appear to be under stabilizing selection, especially over long time periods. Over short periods, expression of most genes may evolve without constraints (Whitehead and Crawford, 2006a, 2006b).
Comparative data
Comparative studies of gene regulation have identified expression patterns correlated with both macroscopic phenotypes and environmental variables. For example, expression of some genes in Fundulus species was found to be correlated with temperature rather than with the phylogenetic relationship of the sampled populations (Oleksiak et al., 2002). Of 329 metabolic genes, 22% retained a significant correlation with temperature after correcting for phylogenetic correlations using a generalized least squares method and a phylogeny derived from five microsatellite loci (Whitehead and Crawford, 2006a, 2006b). The generalized least squares method (Martins and Hansen, 1997) is the same as the method of independent contrasts (Felsenstein, 1985) when characters evolve under a Brownian motion model (Rohlf, 2001). In another study, a subset of expression differences among strains of S. cerevisiae were found to be correlated with resistance to copper sulfate rather than with DNA sequence divergence at three loci (Fay et al., 2004). Finally, pathogenic and commensal strains of E. coli and Shigella species showed an incongruence between DNA-based and transcript-based phylogenies, suggesting convergent evolution (Le et al., 2005).
Interpreting comparative analyses
A number of comparative studies have examined gene expression differences within and between primate species. Some studies found a high rate of expression changes in the brain along the human lineage (Enard et al., 2002; Gu and Gu, 2003; Khaitovich et al., 2005a, 2005b), while other studies found little or no evidence of an accelerated rate (Hsieh et al., 2003; Uddin et al., 2004; Gilad et al., 2006a, 2006b). The reasons for these differences may be technical or methodological and have been discussed elsewhere (Gilad et al., 2006a, 2006b).
Three issues confound the interpretation of comparative studies of gene expression. The first is accounting for nonindependence among samples (Cavalli-Sforza and Edwards, 1967). When species are sampled, their shared evolutionary histories can be accounted for through phylogenetic reconstruction (Cheverud et al., 1985; Felsenstein, 1985; Grafen, 1989; Martins and Hansen, 1997; Rohlf, 2001). However, the situation is more complicated when samples are taken from the same species because different regions of the genome have different evolutionary histories (assuming mating and recombination occur). This makes intraspecific comparative studies (Oleksiak et al., 2002; Fay et al., 2004) subject to correlated patterns of change that cannot easily be controlled for.
A second issue related to comparative data is that many changes in gene expression can be correlated with a phenotype simply because of an inherent genetic or developmental program. For example, genes that show a correlation with resistant to copper sulfate are known to respond to general cellular stresses. Differential expression of these genes is likely a consequence of cells being sensitive or resistant to copper sulfate, rather than because these genes play a direct role in mediating copper resistance (Fay et al., 2004). Similarly, many genes associated with aerobic metabolism have changed expression along the human lineage (Uddin et al., 2004). These changes are likely a response to increased energy consumption rather than selection for changes in gene expression. Single-gene studies are subject to the same caveat; observing a change in gene expression that correlates with a phenotype does not prove that this is the molecular change responsible for adaptive divergence. The difficulty of separating cause and effect in gene expression studies can confound evolutionary interpretations of regulatory changes.
Nonindependence among coregulated genes, caused by the structure of regulatory networks, can also complicate comparative studies. Mutation accumulation studies indicate that groups of functionally related genes often acquire regulatory changes together (Denver et al., 2005), suggesting that many changes in gene expression are not independent. If so, then observing a large number of genes correlated with a character provides just as much evidence as a small number of genes. With some assumptions, this problem can be treated by reducing the number of traits using principle component analysis (Oleksiak et al., 2005) or by estimating the genetic variance and covariance matrices of the traits from family (Lande, 1979; Lande and Arnold, 1983) or population data (Cheverud, 1988). Recent work in this area has generated quantitative genetic methods for distinguishing selection and drift by detecting differences in the genetic variance and covariance matrices from two or more species (Roff, 2000). Even if covariant patterns of gene expression can be taken into account, the number of genes showing particular patterns of variation among samples may be relatively uninformative.
Tests of neutrality
The development of neutral models for the evolution of polygenic characters has provided a quantitative framework to understand the evolution of gene expression. Expression differences within or between species can be compared to those expected given an estimate of the mutational variance. Alternatively, changes in gene expression can be tested for rate heterogeneity across phylogenetic lineages. These models are similar to the models underlying the comparative approach.
Neutral models
A number of neutral models have been proposed for the evolution of polygenic traits (Lande, 1976; Chakraborty and Nei, 1982; Lynch and Hill, 1986; Khaitovich et al., 2005a, 2005b). The first class of models require no explicit genetic basis: given a trait with both a genetic and an environmental contribution, if heritable variations are normally distributed, the evolution of the trait (in the presence or in the absence of selection) can be modeled as a Gaussian process (Lande, 1976). In the absence of selection, the evolution of a trait follows a random walk, as described by Brownian motion models (Felsenstein, 1988), with a mean of zero and variance equal to h2σ2t/N, where h2σ2 is the heritable phenotypic variance, t is the number of generations and N is the effective population size (Lande, 1976). This model led to the first statistical test of neutrality based on the rate of phenotypic evolution (Lande, 1977). The test compares the observed phenotypic variance among lineages to that expected given an estimate of the heritable variation in a population, the effective population size and time. However, not all quantities need to be estimated because the equilibrium between mutation and drift determines the expected amount of phenotypic variation in a population. The input of variation by mutation and the loss of variation by drift is equal to 2NVm, where Vm is the mutational variance or the amount of phenotypic variation introduced into a population each generation by mutation. Substituting 2NVm for the heritable variation in a population leads to the classic result that the rate of phenotypic evolution is independent of the population size and depends only on the mutation rate over time (Lande, 1979; Lynch and Hill, 1986). This result is also characteristic of molecular evolution (Kimura, 1968).
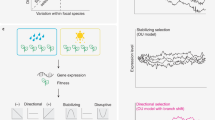
Neutral models of phenotypic evolution can also be formulated based on an explicit genetic model (Chakraborty and Nei, 1982; Lynch and Hill, 1986; Khaitovich et al., 2005a, 2005b). These models use population genetics theory (Crow and Kimura, 1970) to describe the mean and variance in the number of neutral mutations segregating within a population and the rate of substitution between species. The main difference among models is in the mutational variance of a trait, Vm, which is determined by how mutations and the genotype–phenotype relationship are modeled. Most models assume that the phenotypic effects of a mutation are normally distributed, for example continuum of alleles model, while others assume that all mutations have a single phenotypic effect, for example step-wise mutation model (Figure 1). Since Vm is a parameter in all of these models, results can be generalized and are quite intuitive. First, the rate of change in gene expression is equal to the rate of mutations that affect a gene's expression. The reason for this is that the difference in expression between two species is the sum of the effects of all the substitutions that have occurred between species. Since the number of neutral substitutions between species is independent of population size (Kimura, 1968), the rate of divergence in gene expression is equal to the mutation rate for a trait times the average effect of a mutation. If the average effect of a mutation is centered at zero, the variance of the difference between species is 2tVm, where t is the time since two species split, measured in generations (Lande, 1979, 1980; Chakraborty and Nei, 1982; Lynch and Hill, 1986). Second, the amount of heritable phenotypic variation within a population is a function of the population size and the mutational variance. Because the variance in the number of neutral mutations carried by each individual in a population is also a function of the population size and mutation rate (Crow and Kimura, 1970), the equilibrium level of genetic variance for a trait is again approximately 2NVm (assuming a large effective population size and no dominance) (Chakraborty and Nei, 1982; Lynch and Hill, 1986).
Mutation models
Tests of neutrality require an underlying mutational model. The most commonly used model is the continuum of alleles model, which assumes an infinite number of alleles (Kimura and Crow, 1964) with a continuous range of effects on phenotype (Figure 1). Although any distribution of phenotypic effects can be used, they are usually assumed to follow a Gaussian (normal) distribution (Lynch and Hill, 1986). A Poisson process was used to model the special case of the continuous state model that occurs when mutations are rare (Khaitovich et al., 2005a, 2005b). Khaitovich et al. (2005a, 2005b) also considered an asymmetric mutation model where mutational effects follow an extreme value distribution. When mutations are common and their effect size small and symmetric, both models converge to the Gaussian process described by the Brownian motion model.
The multistep mutation model, an extension of the single-step mutation model (Ohta and Kimura, 1973; Kimura and Ohta, 1978), assumes that each mutation causes a finite increase or decrease in some number of steps from the current allelic state. The multistep mutation model was used to model the evolution of a neutral character following an infinite but discrete distribution of states (Chakraborty and Nei, 1982). Under this model, the phenotype is determined by the sum of the allelic states across all loci, and mutations cause binomial deviations in the number of steps from the current state at each locus. When the binomial deviations are large, they are approximately normally distributed and the discrete state model approaches the continuous state model.
There are a number of concerns in applying any of the above-mentioned models to gene expression data. The first concern involves counting alleles and estimating their effects. One must assume that each mutation is detectable or else classify expression patterns into allelic states. Both options are problematic. Mutations with small effects may be missed because of the error inherent to measuring gene expression. At the same time, small differences resulting from imprecise measurements of the same allele may be erroneously considered different alleles. Without incorporating these sources of error into tests of neutrality, these problems are left unresolved. Furthermore, functionally equivalent alleles cannot be distinguished from those that are identical by descent. The infinite alleles model assumes no back mutations and so does not account for different alleles with the same function. This complicates the interpretation of studies using comparisons of regulatory polymorphism and divergence to infer adaptive divergence (for example Rifkin et al., 2003).
A second concern with applying neutral models to gene expression data is that all of the models assume the mutational variance is constant over time. Put differently, both the discrete and continuous phenotype models assume that the phenotype can evolve without mutational constraints such that the distribution of mutational effects is independent of the phenotype. Although this may be valid for modeling fold changes in gene expression levels over short time periods, this assumption will be violated if the absolute effect of a mutation depends on the current value of the phenotype. This seems likely, as mutations that decrease gene expression levels may be more common when gene expression levels are high and mutations that increase gene expression may be more common when gene expression is low. Theoretical models have been developed to accommodate such biased walks (Lande, 1976; Felsenstein, 1988; Kimmel et al., 1996); however, distinguishing selection from a biased mutation process may not be possible from polymorphism and divergence data alone because of the higher likelihood of convergence.
Rates tests
Neutral models estimate the rate at which mutation and genetic drift create variation within a population and divergence between species. When comparing observed data to neutral expectations, too little variation implies that purifying selection has constrained changes in gene expression, whereas too much variation implies that selection has either maintained variation within a population or driven divergence between species. Tests of neutrality based on the rate of expression changes are analogous to the tests of neutrality based on the rate of synonymous and nonsynonymous substitutions in protein coding sequences (Fay and Wu, 2003).
Studies comparing intra- and interspecific patterns of gene expression to neutral expectations indicate that the expression levels of most genes are selectively constrained; relatively few genes appear to be subject to adaptive evolution (Hsieh et al., 2003; Rifkin et al., 2003; Lemos et al., 2005). The key parameter in these models is the mutational variance, which is determined by the mutation rate and the average effect size. In one of the first evolutionary comparisons of gene expression on a genomic scale, Rifkin et al. (2003) estimated the mutational variance from patterns of gene expression within species. However, estimates of the mutational variance from population data can be unreliable (Turelli et al., 1988; Lemos et al., 2005). If mutations affecting the expression of a gene are rare, the mutational variance estimated from equilibrium levels of variation within a population will vary greatly over time and can be zero if a trait becomes monomorphic (Lynch and Hill, 1986). More recently, Rifkin et al. (2005) have directly measured the mutational variance using mutation accumulation lines (see below). Using a range of reasonable values for the mutational variance, Lemos et al. (2005) also found that the majority of gene expression levels are stable over time.
An alternative to these methods, one that avoids estimating the mutational variance, is to simply rank genes based on the ratio of polymorphism within a species to divergence between species. For expression patterns driven by positive selection, large differences between species are expected despite little variation within a species. Although this approach does not distinguish between neutral and selected changes, it provides an interesting list of candidate genes for further study (Meiklejohn et al., 2003; Nuzhdin et al., 2004; Gilad et al., 2006a, 2006b). For example, using this ranking system, Gilad et al. (2006a, 2006b) showed that a significant fraction of candidate genes identified in a comparison of primate species were transcription factors. Transcription factors are often dose sensitive (Seidman and Seidman, 2002), suggesting that in the absence of directional selection, their expression should be tightly constrained.
Relative rates tests
Methods that detect changes in the rate of divergence on one lineage may be more powerful than methods that assume a constant rate of regulatory evolution on all branches. Selection can cause short periods of rapid evolution, which may rarely increase the overall rate of divergence above neutral expectations. Similar to relative rates tests for protein coding sequences, a change in the rate of expression divergence can be explained not only by positive selection, but also by a change in functional constraint, that is, purifying selection (Fay and Wu, 2003). Nonetheless, likelihood methods have been implemented to test for rate heterogeneity across lineages or for a rate shift at a specific point in a phylogeny (Gu, 2004; Oakley et al., 2005). These methods are based on the Brownian motion model and do not require estimates of the mutational variance because they simply test whether rates of divergence on different phylogenetic lineages differ from one another. Application of these methods to expression data from paralogous gene families has shown that gene duplication results in an increase in the rate of expression divergence, particularly right after gene duplication (Gu, 2004; Gu et al., 2005). Although these methods have not yet been applied to divergence in the expression of orthologous genes, evidence for lineage-specific rates of evolution across Drosophila species was found using a heuristic method (Rifkin et al., 2003).
Empirical patterns of neutral evolution
Theoretical models can be used to predict neutral patterns of evolution, but their accuracy depends upon the validity of the underlying assumptions. Often, these assumptions are difficult to evaluate. An alternative strategy for determining patterns of neutral evolution is to simply observe neutral evolution in action. Changes in the expression of pseudogenes and changes in gene expression observed in mutation accumulation lines have been used as neutral proxies for regulatory evolution.
Pseudogenes
To determine the baseline rate of neutral evolution between humans and chimpanzees, Khaitovich et al. (2004) examined patterns of pseudogene expression. Pseudogenes are sequences that resemble genes, but are not thought to have any genetic function. Pseudogenes should evolve neutrally because without a function, mutations in pseudogenes cannot be deleterious or advantageous. Expression of functional genes was found to evolve at a rate similar to that of pseudogenes, suggesting little constraint on gene expression levels. However, for pseudogenes to have been used in this study, they were required to be present and expressed in both species, suggesting they may not be evolving neutrally (Balakirev and Ayala, 2003; Svensson et al., 2006). Only 23 pseudogenes were suitable for this analysis and it is not clear whether this sample size affects the results.
Mutation accumulation
As discussed above, the mutational variance is a critical parameter for modeling the evolution of gene expression. Mutational variance describes the proportion of phenotypic variance added to a population by mutations each generation. Genes with a high mutational variance acquire regulatory changes often, whereas genes with low mutational variance rarely change without the influence of selection. Different genes may have a different propensity for regulatory mutations (Gompel et al., 2005), thus empirical measurements of mutational variance for individual genes will ultimately be needed.
Mutational variance can be directly measured by eliminating natural selection and drift, allowing all mutations (except those causing lethality or sterility) to be maintained. Mutation accumulation lines accomplish this by using either a single individual (for selfing organisms) or a single male–female pair (for sexual species) to found each generation. In C. elegans, mutation accumulation lines were maintained for 280 generations and expression divergence was observed for 9% of the 7014 genes examined (Denver et al., 2005). Expression differences between natural isolates separated for thousands of generations affected only about one fifth as many genes, indicating that new mutations are not limiting for expression divergence. Rather, purifying selection minimizes expression differences in wild populations. Analysis of gene expression in Drosophila melanogaster mutation accumulation lines maintained for 200 generations also suggests that stabilizing selection is the primary force shaping regulatory evolution (Rifkin et al., 2005). Fewer changes in gene expression exist among Drosophila species than expected based on mutation rates in D. melanogaster. Nonetheless, mutational input does appear to influence expression divergence. Genes with the largest mutational variance had the largest expression differences among species, and variability among functional groups was similar in the mutational accumulation lines and interspecific comparisons.
Conclusions and future directions
The discovery of abundant, heritable variation in gene expression segregating in natural populations has reinvigorated investigations into the evolution of gene regulation. A similar flurry of studies followed the discovery of allozyme diversity 40 years ago (Lewontin and Hubby, 1966). In both cases, the primary focus was on whether diversity observed at the molecular level was the result of natural selection. Experimental evolution and evolutionary comparisons of development provide strong evidence that adaptation often occurs by changes in gene regulation. Comparisons of genomic expression patterns may be able to provide definitive answers about the general role of adaptation in the evolution of gene regulation; however, with existing models, observed patterns of regulatory variation are often consistent with both neutral and adaptive models. Elucidating basic parameters of regulatory mutations (for example, their frequency and distributions of mutational effects) will improve these models and create reliable tests for natural selection that can be applied to gene expression data.
Current and future investigations of regulatory evolution will identify polymorphisms underlying population genetic variation in gene expression (Ronald et al., 2005; Stranger et al., 2005; Tao et al., 2006). With these data in hand, we will be able to characterize genetic changes responsible for divergent expression, and design tests for signs of positive selection in regulatory DNA. However, these steps are not straightforward. For example, changes in gene expression, especially between species, are often caused by divergent cis-regulatory sequences (Brem et al., 2002; Yan et al., 2002; Schadt et al., 2003; Wittkopp et al., 2004), which are not well understood. The uncoupling of cis-regulatory sequence and function limits our ability to predict the phenotypic consequences of individual base substitutions (Tautz, 2000; Wittkopp, 2006). However, detailed studies of cis-regulatory regions are in progress for many model systems, including humans (ENCODE, 2004). In the near future, we may be able to predict which bases within a cis-regulatory element are functional and nonfunctional, allowing neutrality tests analogous to those used in coding regions to be developed for cis-regulatory sequences. Progress has already been made on this front (Hahn, 2006), although it is not yet clear which approach will be most reliable.
We are still at the early stages of understanding the molecular, genetic and evolutionary forces underlying divergent gene expression. Over the next few years, it will be exciting to discover how natural selection has shaped patterns of gene expression within and between species on a genomic scale. With regulatory diversity now documented in many systems, future studies can begin dissecting the genetic basis and biological functions of adaptive changes in gene expression.
References
Abzhanov A, Protas M, Grant BR, Grant PR, Tabin CJ (2004). Bmp4 and morphological variation of beaks in Darwin's finches. Science 305: 1462–1465.
Arnold SJ (1992). Constraints on phenotypic evolution. Am Nat 140: S85–S107.
Balakirev ES, Ayala FJ (2003). Pseudogenes: are they ‘junk’ or functional DNA? Annu Rev Genet 37: 123–151.
Beldade P, Brakefield PM, Long AD (2002). Contribution of distal-less to quantitative variation in butterfly eyespots. Nature 415: 315–318.
Bray NJ, Buckland PR, Owen MJ, O’Donovan MC (2003). Cis-acting variation in the expression of a high proportion of genes in human brain. Hum Genet 113: 149–153.
Brem RB, Kruglyak L (2005). The landscape of genetic complexity across 5700 gene expression traits in yeast. Proc Natl Acad Sci USA 102: 1572–1577.
Brem RB, Storey JD, Whittle J, Kruglyak L (2005). Genetic interactions between polymorphisms that affect gene expression in yeast. Nature 436: 701–703.
Brem RB, Yvert G, Clinton R, Kruglyak L (2002). Genetic dissection of transcriptional regulation in budding yeast. Science 296: 752–755.
Britten RJ, Davidson EH (1969). Gene regulation for higher cells: a theory. Science 165: 349–357.
Carroll SB, Grenier JK, Weatherbee SD (2001). From DNA to Diversity: Molecular Genetics and the Evolution of Animal Design. Blackwell: New York.
Cavalieri D, Townsend JP, Hartl DL (2000). Manifold anomalies in gene expression in a vineyard isolate of Saccharomyces cerevisiae revealed by DNA microarray analysis. Proc Natl Acad Sci USA 97: 12369–12374.
Cavalli-Sforza LL, Edwards AWF (1967). Phylogenetic analysis: models and estimation procedures. Evolution 21: 550–570.
Chakraborty R, Nei M (1982). Genetic differentiation of quantitative characters between populations or species. I. Mutation and random genetic drift. Genet Res 39: 303–314.
Cheung VG, Spielman RS, Ewens KG, Weber TM, Morley M, Burdick JT (2005). Mapping determinants of human gene expression by regional and genome-wide association. Nature 437: 1365–1369.
Cheverud JM (1988). A comparison of genetic and phenotypic correlations. Evolution 42: 958–968.
Cheverud JM, Dow MM, Leutenegger W (1985). The quantitative assessment of phylogenetic constraints in comparative analyses: sexual dimorphism in body weight among primates. Evolution 39: 1335–1351.
Clark RM, Wagler TN, Quijada P, Doebley J (2006). A distant upstream enhancer at the maize domestication gene tb1 has pleiotropic effects on plant and inflorescent architecture. Nat Genet 38: 594–597.
Cooper TF, Rozen DE, Lenski RE (2003). Parallel changes in gene expression after 20 000 generations of evolution in Escherichia coli. Proc Natl Acad Sci USA 100: 1072–1077.
Cowles CR, Hirschhorn JN, Altshuler D, Lander ES, Green P, Miller W et al. (2002). Detection of regulatory variation in mouse genes. Nat Genet 32: 432–437.
Crow JF, Kimura M (1970). An Introduction to Population Genetics Theory. Harper and Row: New York, NY.
Darwin C (1876). On the Origin of Species by Means of Natural Selection; or, the Preservation of Favored Races in the Struggle for Life. D. Appleton and Co.: New York, NY.
Davidson EH (2001). Genomic Regulatory Systems: Development and Evolution. Academic Press: San Diego, CA.
Denver DR, Morris K, Streelman JT, Kim SK, Lynch M, Thomas WK (2005). The transcriptional consequences of mutation and natural selection in Caenorhabditis elegans. Nat Genet 37: 544–548.
Diaz-Uriarte R, Garland TJ (1998). Effects of branch length errors on the performance of phylogenetically independent contrasts. Syst Biol 19: 654–672.
Doebley J, Stec A, Hubbard L (1997). The evolution of apical dominance in maize. Nature 386: 485–488.
Doss S, Schadt EE, Drake TA, Lusis AJ (2005). Cis-acting expression quantitative trait loci in mice. Genome Res 15: 681–691.
Enard W, Khaitovich P, Klose J, Zollner S, Heissig F, Giavalisco P et al. (2002). Intra- and interspecific variation in primate gene expression patterns. Science 296: 340–343.
ENCODE (2004). The ENCODE (ENCyclopedia Of DNA Elements) Project. Science 306: 636–640.
Fay JC, McCullough HL, Sniegowski PD, Eisen MB (2004). Population genetic variation in gene expression is associated with phenotypic variation in Saccharomyces cerevisiae. Genome Biol 5: R26.
Fay JC, Wu CI (2003). Sequence divergence, functional constraint, and selection in protein evolution. Annu Rev Genomics Hum Genet 4: 213–235.
Felsenstein J (1985). Phylogenies and the comparative method. Am Nat 125: 1–15.
Felsenstein J (1988). Phylogenies from molecular sequences: inference and reliability. Annu Rev Genet 22: 521–565.
Ferea TL, Botstein D, Brown PO, Rosenzweig RF (1999). Systematic changes in gene expression patterns following adaptive evolution in yeast. Proc Natl Acad Sci USA 96: 9721–9726.
Fong SS, Joyce AR, Palsson BO (2005). Parallel adaptive evolution cultures of Escherichia coli lead to convergent growth phenotypes with different gene expression states. Genome Res 15: 1365–1372.
Gibson G, Riley-Berger R, Harshman L, Kopp A, Vacha S, Nuzhdin S et al. (2004). Extensive sex-specific nonadditivity of gene expression in Drosophila melanogaster. Genetics 167: 1791–1799.
Gilad Y, Oshlack A, Rifkin SA (2006a). Natural selection on gene expression. Trends Genet 22: 456–461.
Gilad Y, Oshlack A, Smyth GK, Speed TP, White KP (2006b). Expression profiling in primates reveals a rapid evolution of human transcription factors. Nature 440: 242–245.
Gompel N, Prud’homme B, Wittkopp PJ, Kassner VA, Carroll SB (2005). Chance caught on the wing: cis-regulatory evolution and the origin of pigment patterns in Drosophila. Nature 433: 481–487.
Grafen A (1989). The phylogenetic regression. Philos Trans R Soc Lond B Biol Sci 326: 119–157.
Gu J, Gu X (2003). Induced gene expression in human brain after the split from chimpanzee. Trends Genet 19: 63–65.
Gu X (2004). Statistical framework for phylogenomic analysis of gene family expression profiles. Genetics 167: 531–542.
Gu Z, David L, Petrov D, Jones T, Davis RW, Steinmetz LM (2005). Elevated evolutionary rates in the laboratory strain of Saccharomyces cerevisiae. Proc Natl Acad Sci USA 102: 1092–1097.
Hahn MW (2006). Detecting natural selection on cis-regulatory DNA. Genetica 129: 7–18.
Harvey PH, Pagel MD (1991). The Comparative Method in Evolutionary Biology. Oxford University Press: New York, NY.
Harvey PH, Purvis A (1991). Comparative methods for explaining adaptations. Nature 351: 619–624.
Hsieh WP, Chu TM, Wolfinger RD, Gibson G (2003). Mixed-model reanalysis of primate data suggests tissue and species biases in oligonucleotide-based gene expression profiles. Genetics 165: 747–757.
Hubbard L, McSteen P, Doebley J, Hake S (2002). Expression patterns and mutant phenotype of teosinte branched1 correlate with growth suppression in maize and teosinte. Genetics 162: 1927–1935.
Huelsenbeck JP, Rannala B (2003). Detecting correlation between characters in a comparative analysis with uncertain phylogeny. Evolution 57: 1237–1247.
Jin W, Riley RM, Wolfinger RD, White KP, Passador-Gurgel G, Gibson G (2001). The contributions of sex, genotype and age to transcriptional variance in Drosophila melanogaster. Nat Genet 29: 389–395.
Khaitovich P, Hellmann I, Enard W, Nowick K, Leinweber M, Franz H et al. (2005a). Parallel patterns of evolution in the genomes and transcriptomes of humans and chimpanzees. Science 309: 1850–1854.
Khaitovich P, Paabo S, Weiss G (2005b). Toward a neutral evolutionary model of gene expression. Genetics 170: 929–939.
Khaitovich P, Weiss G, Lachmann M, Hellmann I, Enard W, Muetzel B et al. (2004). A neutral model of transcriptome evolution. PLoS Biol 2: E132.
Kimmel M, Chakraborty R, Stivers DN, Deka R (1996). Dynamics of repeat polymorphisms under a forward–backward mutation model: within- and between-population variability at microsatellite loci. Genetics 143: 549–555.
Kimura M (1968). Evolutionary rate at the molecular level. Nature 217: 624–626.
Kimura M (1983). The Neutral Theory of Molecular Evolution. Cambridge University Press: Cambridge, UK.
Kimura M, Crow JF (1964). The number of alleles that can be maintained in a finite population. Genetics 49: 561–576.
Kimura M, Ohta T (1978). Stepwise mutation model and distribution of allelic frequencies in a finite population. Proc Natl Acad Sci USA 75: 2868–2872.
King MC, Wilson AC (1975). Evolution at two levels in humans and chimpanzees. Science 188: 107–116.
Kirst M, Basten CJ, Myburg AA, Zeng ZB, Sederoff RR (2005). Genetic architecture of transcript-level variation in differentiating xylem of a eucalyptus hybrid. Genetics 169: 2295–2303.
Konishi S, Izawa T, Lin SY, Ebana K, Fukuta Y, Sasaki T et al. (2006). An SNP caused loss of seed shattering during rice domestication. Science 312: 1392–1396.
Kopp A, Duncan I, Godt D, Carroll SB (2000). Genetic control and evolution of sexually dimorphic characters in Drosophila. Nature 408: 553–559.
Lai Z, Gross BL, Zou Y, Andrews J, Rieseberg LH (2006). Microarray analysis reveals differential gene expression in hybrid sunflower species. Mol Ecol 15: 1213–1227.
Lande R (1976). Natural selection and random genetic drift in phenotypic evolution. Evolution 26: 314–334.
Lande R (1977). Statistical tests for natural selection on quantitative characters. Evolution 31: 442–444.
Lande R (1979). Quantitative genetic analysis of multivariate evolution, applied to brain: body size allometry. Evolution 33: 402–416.
Lande R (1980). Sexual dimorphism, sexual selection, and adaptation in polygenic characters. Evolution 34: 292–305.
Lande R, Arnold SJ (1983). The measurement of selection on correlated characters. Evolution 37: 1210–1226.
Landry CR, Oh J, Hartl DL, Cavalieri D (2006). Genome-wide scan reveals that genetic variation for transcriptional plasticity in yeast is biased towards multi-copy and dispensable genes. Gene 366: 343–351.
Le GT, Darlu P, Escobar-Paramo P, Picard B, Denamur E (2005). Selection-driven transcriptome polymorphism in Escherichia coli/Shigella species. Genome Res 15: 260–268.
Lemos B, Meiklejohn CD, Caceres M, Hartl DL (2005). Rates of divergence in gene expression profiles of primates, mice, and flies: stabilizing selection and variability among functional categories. Evolution 59: 126–137.
Lewontin RC, Hubby JL (1966). A molecular approach to the study of genic heterozygosity in natural populations. II. Amount of variation and degree of heterozygosity in natural populations of Drosophila pseudoobscura. Genetics 54: 595–609.
Lo HS, Wang Z, Hu Y, Yang HH, Gere S, Buetow KH et al. (2003). Allelic variation in gene expression is common in the human genome. Genome Res 13: 1855–1862.
Lynch M, Hill WG (1986). Phenotype evolution by neutral mutation. Evolution 40: 915–935.
Martins E (2000). Adaptation and the comparative method. Trends Ecol Evol 15: 296–299.
Martins EP, Garland Jr T (1991). Phylogenetic analyses of the correlated evolution of continuous characters: a simulation study. Evolution 45: 534–557.
Martins EP, Hansen TF (1997). Phylogenies and the comparative method: a general approach to incorporating phylogenetic information into the analysis of interspecific data. Am Nat 149: 646–667.
Meiklejohn CD, Parsch J, Ranz JM, Hartl DL (2003). Rapid evolution of male-biased gene expression in Drosophila. Proc Natl Acad Sci USA 100: 9894–9899.
Monks SA, Leonardson A, Zhu H, Cundiff P, Pietrusiak P, Edwards S et al. (2004). Genetic inheritance of gene expression in human cell lines. Am J Hum Genet 75: 1094–1105.
Morley M, Molony CM, Weber TM, Devlin JL, Ewens KG, Spielman RS et al. (2004). Genetic analysis of genome-wide variation in human gene expression. Nature 430: 743–747.
Nuzhdin SV, Wayne ML, Harmon KL, McIntyre LM (2004). Common pattern of evolution of gene expression level and protein sequence in Drosophila. Mol Biol Evol 21: 1308–1317.
Oakley TH, Gu Z, Abouheif E, Patel NH, Li WH (2005). Comparative methods for the analysis of gene-expression evolution: an example using yeast functional genomic data. Mol Biol Evol 22: 40–50.
Ohta T, Kimura M (1973). A model of mutation appropriate to estimate the number of electrophoretically detectable alleles in a finite population. Genet Res 22: 201–204.
Oleksiak MF, Churchill GA, Crawford DL (2002). Variation in gene expression within and among natural populations. Nat Genet 32: 261–266.
Oleksiak MF, Roach JL, Crawford DL (2005). Natural variation in cardiac metabolism and gene expression in Fundulus heteroclitus. Nat Genet 37: 67–72.
Orr HA (2005). The probability of parallel evolution. Evolution 59: 216–220.
Pastinen T, Sladek R, Gurd S, Sammak A, Ge B, Lepage P et al. (2004). A survey of genetic and epigenetic variation affecting human gene expression. Physiol Genomics 16: 184–193.
Radich JP, Mao M, Stepaniants S, Biery M, Castle J, Ward T et al. (2004). Individual-specific variation of gene expression in peripheral blood leukocytes. Genomics 83: 980–988.
Raff RA (1996). The Shape of Life: Genes, Development, and the Evolution of Animal Form. University of Chicago Press: Chicago.
Ranz JM, Castillo-Davis CI, Meiklejohn CD, Hartl DL (2003). Sex-dependent gene expression and evolution of the Drosophila transcriptome. Science 300: 1742–1745.
Ranz JM, Namgyal K, Gibson G, Hartl DL (2004). Anomalies in the expression profile of interspecific hybrids of Drosophila melanogaster and Drosophila simulans. Genome Res 14: 373–379.
Riehle MM, Bennett AF, Lenski RE, Long AD (2003). Evolutionary changes in heat-inducible gene expression in lines of Escherichia coli adapted to high temperature. Physiol Genomics 14: 47–58.
Rifkin SA, Houle D, Kim J, White KP (2005). A mutation accumulation assay reveals a broad capacity for rapid evolution of gene expression. Nature 438: 220–223.
Rifkin SA, Kim J, White KP (2003). Evolution of gene expression in the Drosophila melanogaster subgroup. Nat Genet 33: 138–144.
Rockman MV, Wray GA (2002). Abundant raw material for cis-regulatory evolution in humans. Mol Biol Evol 19: 1991–2004.
Roff D (2000). The evolution of the G matrix: selection or drift? Heredity 84: 135–142.
Rohlf FJ (2001). Comparative methods for the analysis of continuous variables: geometric interpretations. Evolution 55: 2143–2160.
Ronald J, Brem RB, Whittle J, Kruglyak L (2005). Local regulatory variation in Saccharomyces cerevisiae. PLoS Genet 1: e25.
Schadt EE, Monks SA, Drake TA, Lusis AJ, Che N, Colinayo V et al. (2003). Genetics of gene expression surveyed in maize, mouse and man. Nature 422: 297–302.
Seidman JG, Seidman C (2002). Transcription factor haploinsufficiency: when half a loaf is not enough. J Clin Invest 109: 451–455.
Shapiro MD, Marks ME, Peichel CL, Blackman BK, Nereng KS, Jonsson B et al. (2004). Genetic and developmental basis of evolutionary pelvic reduction in threespine sticklebacks. Nature 428: 717–723.
Shockley KR, Churchill GA (2006). Gene expression analysis of mouse chromosome substitution strains. Mamm Genome 17: 598–614.
Stern DL (1998). A role of Ultrabithorax in morphological differences between Drosophila species. Nature 396: 463–466.
Storey JD, Akey JM, Kruglyak L (2005). Multiple locus linkage analysis of genomewide expression in yeast. PLoS Biol 3: e267.
Stranger BE, Forrest MS, Clark AG, Minichiello MJ, Deutsch S, Lyle R et al. (2005). Genome-wide associations of gene expression variation in humans. PLoS Genet 1: e78.
Sucena E, Stern DL (2000). Divergence of larval morphology between Drosophila sechellia and its sibling species caused by cis-regulatory evolution of ovo/shaven-baby. Proc Natl Acad Sci USA 97: 4530–4534.
Svensson O, Arvestad L, Lagergren J (2006). Genome-wide survey for biologically functional pseudogenes. PLoS Comput Biol 2: e46.
Tao H, Cox DR, Frazer KA (2006). Allele-specific KRT1 expression is a complex trait. PLoS Genet 2: e93.
Tautz D (2000). Evolution of transcriptional regulation. Curr Opin Genet Dev 10: 575–579.
Townsend JP, Cavalieri D, Hartl DL (2003). Population genetic variation in genome-wide gene expression. Mol Biol Evol 20: 955–963.
Turelli M, Gillespie JH, Lande R (1988). Rate tests for selection on quantitative characters during macroevolution and microevolution. Evolution 42: 1085–1089.
Uddin M, Wildman DE, Liu G, Xu W, Johnson RM, Hof PR et al. (2004). Sister grouping of chimpanzees and humans as revealed by genome-wide phylogenetic analysis of brain gene expression profiles. Proc Natl Acad Sci USA 101: 2957–2962.
Vuylsteke M, van EF, Van HP, Kuiper M, Zabeau M (2005). Genetic analysis of variation in gene expression in Arabidopsis thaliana. Genetics 171: 1267–1275.
Wang H, Nussbaum-Wagler T, Li B, Zhao Q, Vigouroux Y, Faller M et al. (2005). The origin of the naked grains of maize. Nature 436: 714–719.
Wayne ML, McIntyre LM (2002). Combining mapping and arraying: an approach to candidate gene identification. Proc Natl Acad Sci USA 99: 14903–14906.
Wayne ML, Pan YJ, Nuzhdin SV, McIntyre LM (2004). Additivity and trans-acting effects on gene expression in male Drosophila simulans. Genetics 168: 1413–1420.
Whitehead A, Crawford DL (2005). Variation in tissue-specific gene expression among natural populations. Genome Biol 6: R13.
Whitehead A, Crawford DL (2006a). Neutral and adaptive variation in gene expression. Proc Natl Acad Sci USA 103: 5425–5430.
Whitehead A, Crawford DL (2006b). Variation within and among species in gene expression: raw material for evolution. Mol Ecol 15: 1197–1211.
Whitney AR, Diehn M, Popper SJ, Alizadeh AA, Boldrick JC, Relman DA et al. (2003). Individuality and variation in gene expression patterns in human blood. Proc Natl Acad Sci USA 100: 1896–1901.
Wilson AC, Maxson LR, Sarich VM (1974). Two types of molecular evolution: evidence from studies of interspecific hybridization. Proc Natl Acad Sci USA 71: 2843–2847.
Wittkopp PJ (2006). Evolution of cis-regulatory sequences and function in diptera. Heredity 97: 139–147.
Wittkopp PJ, Haerum BK, Clark AG (2004). Evolutionary changes in cis and trans gene regulation. Nature 15: 85–88.
Wittkopp PJ, Vaccaro K, Carroll SB (2002). Evolution of yellow gene regulation and pigmentation in Drosophila. Curr Biol 12: 1547–1556.
Wittkopp PJ, Williams BL, Selegue JE, Carroll SB (2003). Drosophila pigmentation evolution: divergent genotypes underlying convergent phenotypes. Proc Natl Acad Sci USA 100: 1808–1813.
Wray GA, Hahn MW, Abouheif E, Balhoff JP, Pizer M, Rockman MV et al. (2003). The evolution of transcriptional regulation in eukaryotes. Mol Biol Evol 20: 1377–1419.
Yan H, Yuan W, Velculescu VE, Vogelstein B, Kinzler KW (2002). Allelic variation in human gene expression. Science 297: 1143.
Yvert G, Brem RB, Whittle J, Akey JM, Foss E, Smith EN et al. (2003). Trans-acting regulatory variation in Saccharomyces cerevisiae and the role of transcription factors. Nat Genet 35: 57–64.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Fay, J., Wittkopp, P. Evaluating the role of natural selection in the evolution of gene regulation. Heredity 100, 191–199 (2008). https://doi.org/10.1038/sj.hdy.6801000
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6801000
Keywords
This article is cited by
-
Molecular and evolutionary processes generating variation in gene expression
Nature Reviews Genetics (2021)
-
The strength and pattern of natural selection on gene expression in rice
Nature (2020)
-
Regulatory and sequence evolution in response to selection for improved associative learning ability in Nasonia vitripennis
BMC Genomics (2018)
-
Differential gene expression revealed with RNA-Seq and parallel genotype selection of the ornithine decarboxylase gene in fish inhabiting polluted areas
Scientific Reports (2018)
-
Differences in brain gene transcription profiles advocate for an important role of cognitive function in upstream migration and water obstacles crossing in European eel
BMC Genomics (2015)