Abstract
Alternative splicing (AS) of pre-messenger RNA is a common phenomenon that creates different transcripts from a single gene, and these alternative transcripts affect phenotypes. The majority of AS research has examined tissue and developmental specificity of expression of particular AS transcripts, how this specificity affects cell function, and how aberrant AS is related to disease. Few studies have examined quantitative between-individual variation in AS within a cell or tissue type, or in relation to phenotypes, but the results are compelling: quantitative variation in AS affects plastic traits such as stress, anxiety, fear, egg production, muscle performance, energetics and plant growth. Genomic analyses of AS are also at a nascent stage, but have revealed a number of significant evolutionary patterns. Growing knowledge of upstream genes and kinases that regulate AS provides the as-yet little explored potential to examine how these genes and pathways respond to environmental and genotype variables. Research in this area can provide glimpses of a labyrinth of genetic architectures that have rarely been considered in evolutionary and organismal biology, or in quantitative genetics. The scarcity of contribution to knowledge about AS from these fields is illustrated by the fact that heritability of quantitative variation in AS has not yet been determined for any gene in any organism. New research tactics that incorporate quantitative analyses of AS will allow organismal and evolutionary biologists to attain a fuller mechanistic understanding of many of the traits they study, and may lead to more rapid discovery of functionally important polymorphisms.
Similar content being viewed by others
Main
A major aim in biology is to determine how genes give rise to phenotypes. Associating genes and transcripts with phenotypes is easiest for single amino-acid polymorphisms in structural proteins or enzymes. The task becomes more difficult when a quantitative trait is affected by multiple genes and by the environment, and this is typically the case for the traits most interesting to organismal and evolutionary biologists. For example, egg production and mating behavior or the growth form of plants in different light environments are plastic quantitative traits with complex genetic architectures. Understanding these types of adaptive plasticity and the evolution of underlying gene and gene-by-environment interactions are primary challenges in contemporary biology.
Recent studies continue to advance the theoretical (Gabriel et al., 2005; Izem and Kingsolver, 2005; Leimar, 2005; West-Eberhard, 2005; Zhang, 2005) and empirical (Pigliucci et al., 2003; Nussey et al., 2005; Wayne et al., 2005) understanding of the evolution of phenotypic plasticity and reaction norms, but for the most part evolutionary biology has not yet incorporated rapidly accumulating knowledge of genetic determinants of plastic quantitative traits (e.g. Alonso-Blanco et al., 2005; Mackay and Lyman, 2005; but see Knies et al., 2006).
Alternative splicing as a mechanism for generating phenotypic plasticity
To date, most of the research aimed at understanding mechanistic bases of complex phenotypic changes in response to environmental variables has been aimed at changes in gene expression levels (i.e. quantitative polymerase chain reaction, subtractive hybridization and microarrays). The on–off status or quantitative expression level of transcripts is certainly a potent way to affect phenotypes, but not for genes that encode proteins that must always be present, or must be present in a certain stoichiometric ratio with other proteins. For such genes, having more or less of a certain protein is not always a viable or functionally meaningful option. Alternative splicing (AS) presents another avenue for plasticity, by allowing structurally and functionally different proteins to be produced from a single locus, independently of or in addition to changes in expression level. Also, AS allows a tremendous amplification of protein diversity, thereby providing functional diversity that far exceeds what can be accomplished only by varying expression levels.
AS is a form of transcriptional regulation in which alternative 5′ and 3′ splice sites and/or exon inclusion/skipping give rise to a variety of messenger RNAs (mRNAs) from a single gene. Estimates of the percentage of alternatively spliced genes in mammals range from 22 to 74% (Baek and Green, 2005), and the percentage appears to be similarly high in Drosophila (http://proline.bic.nus.edu.sg/dedb/cgi-bin/altsplice.py). AS in plants appears to be somewhat less prevalent than in animals (8–14% in Arabidopsis and rice; Wang and Brendel, 2006). AS is not entirely restricted to eukaryotes, as retroviruses depend on AS for critical events in their life cycle (Pollard and Malim, 1998). Patterns of AS vary among viral strains, perhaps in functionally important ways (Stoltzfus and Madsen, 2006).
A percentage of alternative transcripts are degraded by nonsense-mediated mRNA decay (Baek and Green, 2005), but the majority of AS transcripts create functionally diverse protein isoforms. Examples of functional diversity among AS-produced proteins include developmental switches that control sex determination in Drosophila (reviewed by Lopez, 1998), membrane pumps that have different directionality of net transport (Lazaridis et al., 2000), and the 38 016 variants of the single Drosophila Dscam gene, which encode a set of immunoglobulin superfamily receptors that function in axon guidance and immune responses (Watson et al., 2005; Chen et al., 2006a). This is a perfunctory list, and knowledge of how AS affects healthy and pathological cellular phenotypes grows almost daily (for recent reviews see Lipscombe, 2005; Stamm et al., 2005; Stetefeld and Ruegg, 2005).
AS from the perspective of evolutionary biology
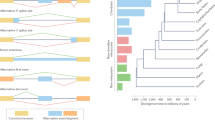
A recent upswing of attention to AS in genomics studies has revealed a number of interesting results. Recently diverged genes within species (Su et al., 2006) and homologous genes across species (Cusack and Wolfe, 2005; Malko et al., 2006) show different patterns of AS, which indicates that AS tends to diverge rapidly after branching events and may affect speciation. AS exons are significantly less conserved, in terms of presence/absence status, between species of flies (Drosophila versus Anopheles) than are constitutive exons (Malko et al., 2006). Patterns of AS can also be convergent, as has been shown for ion channel genes of flies and humans, in which alternative mutually exclusive exons have evolved independently in homologous genes following exon duplication events (Copley, 2004). Coevolution of AS patterns (Herranz et al., 2005) has been inferred from the finding that, across related species, changes in alternative exons in one protein are accompanied by changes in the alternative exons of interacting proteins. Genomic studies have even revealed glimpses of the ‘ghost of splicing past’. This intriguing result comes from a study showing that the human genome contains pseudogenes that arose from incorporation into the genome (reverse transcription) of mature, spliced mRNAs; some of those appear to be alternative transcripts that were expressed ancestrally but are not presently expressed by the parent gene (Shemesh et al., 2006).
Most evolutionary studies of AS have used only bioinformatics data with little knowledge of, or attention to phenotypes. An exception is a study (Wada et al., 2004) in which differential expression and AS of glutamate receptor genes was examined in the brain of birds that are capable or incapable of song learning. Only three avian taxa are capable of song learning (passerines, hummingbirds and parrots) and each of those lineages shows different patterns of expression of glutamate receptor superfamily genes in the vocal learning centers compared to adjacent brain regions. One vocal nonlearner, a dove, was also examined; it showed a conspicuous absence of differential gene expression in the brain areas that contain vocal nuclei in song-learning taxa. Some of these glutamate receptor genes are alternatively spliced, and in the one glutamate receptor subfamily for which expression probes were designed to be splice-form specific, there was differential expression of alternatively spliced forms in vocal nuclei compared to adjacent brain regions. The different splice forms of these receptors are known to differ in how rapidly they desensitize after channel opening, and thus are likely to have significant phenotypic effects that might play a part in vocal learning capability.
On a microevolutionary level, analyses of AS genes show that there is an increase in nonsynonymous substitution (Ka) and a decrease in synonymous substitution (Ks) in regions within and flanking alternative exons (Xing and Lee, 2005), and the ratio of nonsynonymous to synonymous substitution (Ka/Ks) is higher in AS exons compared to constitutively spliced exons (Chen et al., 2006b). These finding suggest that, because they are not always included in the mature transcript and therefore functionally expendable in proteins, alternative exons are less subject to deleterious mutation and therefore are hotspots for the evolution of protein diversity. Comparisons among genes in which AS is present in mouse or human, but not both (i.e. genome-specific AS) show that there is also an increase in nonsynonymous divergence rate in constitutive exons of AS genes (Cusack and Wolfe, 2005), perhaps because most of these instances of genome-specific AS involve new exon insertions that apparently caused evolutionary responses in other parts of the gene. Hence, the constitutive exons of genes respond evolutionarily to the advent of new alternative exons.
AS and the generation of plastic organism-level phenotypes
Mechanistic studies of AS have devoted little attention to variability among individuals. As a result, little interest in AS has been stimulated among organismal and evolutionary biologists, or quantitative geneticists. To appreciate this point, consider that searches of the journals Heredity and Evolution reveal only one research paper (Flood and Ruvinsky, 2001) to date on AS, and there is no mention of AS in the latest editions of leading texts for quantitative (Falconer and Mackay, 1996) and population genetics (Hartl and Clark, 1997). Few studies have examined how AS varies in a quantitative manner among individuals and populations (but see Marden et al., 2001), and to my knowledge, the heritability of quantitative variation in AS has not yet been determined for any gene in any organism. A large percentage of inherited human diseases are caused by mutations that alter AS (Garcia-Blanco et al., 2004), which indicates that there is a large genomic target for mutations that affect qualitative or quantitative patterns of AS. These diseases are likely to be extreme examples of an abundant class of polymorphisms that affect AS and create subtly different phenotypes. Among these loci are surely alleles that alter the way AS is connected to and responds to environmental variation, that is, plasticity genes.
Although quantitative variation in AS among individuals and organism-level phenotypes have not been prevalent research foci, the few studies that have taken this approach reveal a number of striking results. The sections below describe four case studies from this body of work.
Case study 1: neurological responses to stress and changes in behavior associated with AS of acetylcholinesterase
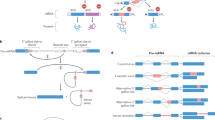
Acetylcholinesterase (AChE) is an enzyme that hydrolizes the neurotransmitter acetylcholine, thereby affecting the balance between transmission and termination of neural signals. AS at the 3′-end of the coding region results in protein isoforms that are enzymatically indistinguishable but differ in how they polymerize and localize. Inclusion of alternative exon 6 (Figure 1a) creates AChE-S, whose protein polymerizes as tetramers that bind to and localize at synaptic membranes. Exclusion of exon 6 and retention of a small intron at the 3′-end of exon 4 creates the AChE-R isoform (Figure 1a) that encodes a monomer that does not bind to synapses but instead moves freely throughout the brain and the body.
(a) Intron–exon structure and alternative exon inclusion at the 3′-end of transcripts of the mammalian AChE gene. (b and c) Fluorescent in situ hybridization with mRNA of AChE-S (b) and AChE-R (c) in the prefrontal cortex of mice before and 2 weeks after stress. (d) Relationship between human self-reported anxiety scores and the level of AChE enzyme activity in their blood serum; this reflects abundance of the freely circulating AChE-R isoform. (a–c) are adapted from Meshorer et al. (2005); (d) is adapted from Sklan et al. (2004).
Acute stress causes an increase in transcription of the AChE-R variant, which is normally rare in neuronal tissue (Figure 1b and c); this has an apparent protective effect of counteracting stress-induced neural damage, perhaps by downregulating brain-wide synaptic transmission. Transgenic mice that overexpress the AChE-S isoform show more stress-induced neurodeterioration, whereas transgenic mice overexpressing AChE-R show reduced neuronal damage compared to controls or to AChE-S overexpressing transgenics (Sternfeld et al., 2000). These results suggest that the AChE-R isoform plays a protective role during acute stress.
The switch to AChE-R transcription during acute stress is rapid (minutes) but persists for weeks (Meshorer et al., 2002). AChE-R has been shown (Birikh et al., 2003) to interact in neurons with the scaffold protein RACK1, and through it with protein kinase CII, a signaling molecule involved in fear conditioning. Mice exposed to mild stress, and transgenic mice overexpressing AChE-R, show fearful behavioral traits such as delayed willingness to emerge from a box into an open field. The AChE-R isoform also increases expression of the splicing factor (SC35) that favors AChE-R transcript formation, thereby forming a positive feedback loop that may be the mechanism underlying the prolonged behavioral impact of acute stress (Meshorer et al., 2005).
These studies of AS in the AChE gene provide both a mechanistic hypothesis and rudiments of a regulatory pathway for acute and chronic stress phenotypes in mammals. They also hint at a source for nonstress related phenotypic variation in behaviors such as fearfulness and boldness. Animal species that have existed for long periods of time on islands in the absence of predators show greatly reduced fear, and it might be productive to determine if they have different patterns of expression of AChE isoforms and splicing factors such as SC35. These molecules might also be good candidates to examine in relation to intraspecific variation in boldness and/or dispersiveness. AChE-R is rare in brains of nonstressed laboratory mice, but might be more common in free-living mammals that experience many types of stress on a regular basis. The hypothesis that variation in basal expression of the AChE-R isoform affects a variety of behavioral phenotypes is supported by the finding that in healthy human volunteers, blood serum levels of monomeric, circulating AChE-R protein show a significant inverse relationship with anxiety (Figure 1d; Sklan et al., 2004). Fearful behavior in mice maps to multiple loci on different chromosomes (Gershenfeld and Paul, 1997), and it would be interesting to determine if any of these loci encode or interact with upstream regulators of AChE or its splice factors. Finally, it is interesting to note that AChE is also alternatively spliced at the 5′-end of the gene (Meshorer and Soreq, 2006), but the phenotypic effects of these splice variants have not yet been examined.
This landmark body of work on AChE has opened many new avenues for exploring the basic biology, biomedicine, and genetics of stress, fear and anxiety. Scientific opinion regarding the importance and impact of new findings is always diverse, but a cautious, tentative and understated approach is typically the norm. Against that background it is eye-opening to find the following quote from a perspective (Rossi, 2004) in the journal Advanced Mind Body Medicine: ‘stress-induced AS of the AChE gene may play a pivotal role in the deep psychobiology of psychotherapy, meditation, spiritual rituals and the experiencing of positive humanistic values that have been associated with mind body medicine, such as compassion, beneficence, serenity, forgiveness and gratitude.’ I dare not add anything more to that.
Case study 2: nutritional status and control of egg fate by AS of the broad complex gene in insects
Fecundity of female insects is a plastic trait that is adjusted in response to nutrition. From a mechanistic standpoint, the best examined species is Drosophila melanogaster, in which it has been shown that egg development reaches a critical decision point at stages 8 and 9 of oogenesis (Terashima and Bownes, 2004). At these stages an egg either commits to and undergoes apoptosis and its nutrients are reabsorbed, or it begins to take up nutrients from the surrounding hemolymph and synthesize yolk proteins. Poor nutrition causes an increase in the titer of the hormone ecdysone, which stimulates an increase in the relative abundance of the Z2 and Z3 splice variants of the Broad Complex gene (BR-C; Figure 2). As demonstrated by flies carrying BR-C transgenes coupled to a heat-shock promoter, heat-shock induced overexpression of Z2 and Z3 constructs caused an increase in the percentage of eggs undergoing apoptosis, whereas heat-shock induced expression of the Z1 and Z4 constructs, or heat shock of the control line, had no significant effects on egg fate (Terashima and Bownes, 2004). In fed flies (access to yeast), experimental injection of ecdysone resulted in an increase in relative abundance of the Z2 and Z3 splice variants, and increased apoptosis of eggs. Experimental application of a juvenile hormone analog had effects that were opposite those of ecdysone. In addition to effects on apoptosis, higher expression of the Z3 splice variant also suppressed expression of three major yolk proteins (yp1, yp2 and yp3).
(a and b) Intron–exon structure and AS of exons containing zinc-finger motifs at the 3′-end of the BR-C gene in Drosophila. Adapted from Bayer et al. (1997). (c and d) Effects of access to nutrients from yeast (F) or no access (S) on relative expression of the Z2 and Z3 splice forms of BR-C, and on the percentage of Stage 9 eggs undergoing apoptosis. Adapted from Terashima and Bownes (2004).
These results show that a primary determinant of potential female fecundity and reproductive fitness, the number of eggs that advance beyond a critical stage of development, is in large part determined by a pathway in which nutritional cues are transduced into hormonal signals that lead to changes in AS of the BR-C gene. In addition to genes that interact in the transduction of nutritional state into a hormonal signal, there are likely to be numerous genes that affect how ecdysone and JH hormonal signals affect AS of BR-C, and the downstream processes by which isoforms of BR-C affect oocyte maturation. Polymorphism in any of these steps could potentially affect the reaction norm describing how female fecundity responds to nutritional signals. Rapidly growing knowledge of the target of rapamycin and insulin/phosphatidylinositol 3-kinase signaling pathways that animals use to monitor their internal nutritional state (Britton et al., 2002; Cota et al., 2006; Dann and Thomas, 2006) are likely to soon provide a much more complete understanding of links between food availability and fecundity. Evolutionary biologists should be greatly interested in these developments, as allocation of energy to maintenance versus reproduction is the central issue in life history evolution.
The general viewpoint in ecology and evolutionary biology is that variable life history outcomes such as egg production are dictated primarily by energy availability and allocation within an individual. What I aim to stress here is that there are known sensory mechanisms and tightly regulated molecular genetic pathways that govern these allocation phenotypes and create reaction norms that should be under intense selection pressures. Description of these genes and pathways by molecular biologists makes them available for examination from an ecological and evolutionary perspective.
Case study 3: nutritional status and the control of muscle contractile performance and energy consumption by AS of the troponin-t gene in dragonflies
Male dragonflies present an interesting counterpoint to the case described above for female insects. Rather than producing eggs as their major reproductive expenditure, male dragonflies engage in elaborate and often highly escalated aerial contests to establish breeding territories that provide them with access to mates. Power output from flight muscles varies by approximately threefold in mature male Libellula pulchella dragonflies, and this variation is positively related to lifetime territorial tenure and number of mates (Marden and Cobb, 2004). As with variation in female fecundity in Drosophila, the reason why such variation persists is because it is a plastic phenotype, determined at least in part by AS of a gene, troponin-t (Tnt), that encodes a protein that comprises part of the calcium-sensitive switch that turns muscle contraction on and off.
The 5′ region of the Tnt gene in L. pulchella contains three alternative exons (Figure 3a and b), one of which is a microexon that contains only one codon. These alternative exons are expressed in different combinations to create six distinct transcripts in flight muscles, along with a seventh that occurs in leg and body wall muscles but is absent from flight muscles. There is a strong correlation between the individual and the summed relative abundance of two of these transcripts and the sensitivity of skinned muscle fibers to activation by calcium (Figure 3c; Marden et al., 2001). There is nearly as strong a correlation between the summed relative abundance of the same two transcripts and the power output of intact muscle (Figure 3d). Experiments in other animals have repeatedly shown that manipulation of Tnt genes and/or protein isoforms affects muscle calcium sensitivity and contractile performance (Ogut et al., 1999; Gomes et al., 2002, 2004; MacFarland et al., 2002; Nassar et al., 2005; Brotto et al., 2006; Chandra et al., 2006), and single amino-acid replacement mutations in human Tnt also have these effects (Hernandez et al., 2005).
(a and b) Intron–exon structure and AS of exons near the 5′-end of the Troponin-t gene (Tnt) in Libellula pulchella dragonflies. Arrows represent additional constitutive exons (not shown) that extend to the 3′-end of the gene. (c and d) Relationship between the relative abundance (% of all Tnt transcripts) of two Tnt splice variants and the calcium sensitivity of skinned muscle fiber activation, and the mechanical power output of intact flight muscle. Adapted from Marden et al. (2001).
Comparisons of muscle power output with fat content (the major energy storage substrate in dragonflies) show that individuals with more fat have higher performing muscles (Marden and Cobb, 2004). Relative expression of one of the power-promoting Tnt alternative transcripts (Tnt 267) is tightly related to the amount of fat in the thorax (Marden, unpublished) and to changes in wingbeat frequency that affect the energetic cost of flight (Marden et al., 1999, 2001). These results suggest that AS of Tnt serves as a ‘fulcrum’ for a tradeoff between energy consumption and flight performance. Dragonflies with ample energy reserves use AS of Tnt to adjust their muscle energy consumption and performance upward, whereas those with low energy reserves settle for a less consumptive but less reproductively successful lifestyle. Here as in the case of Drosophila oogenesis, we see strong clues that AS is involved in a life history tradeoff between energy allocation to reproduction versus survival and somatic maintenance.
An interesting footnote to this dragonfly example is that there are significant differences in the mean values of Tnt transcript abundances and muscle contractile performance variables between samples of dragonflies from two ponds that are separated by only 16 km (Marden et al., 2001). These two ponds occur in areas that have different soil chemistry, which affects the pH of the water and growth rates of plants in adjoining terrestrial areas where adult dragonflies hunt for flying insect prey (Marden and Cassarella, in preparation). Dragonflies at the acid soil site, where plant growth rates are radically lower, have lower muscle power and associated differences in troponin-t alternative transcript mixtures. A working hypothesis is that plant primary production affects the abundance of dragonfly prey, thereby affecting nutritional state and causing population level differences in the regulation of muscle energy consumption through nutrient-sensing systems that affect troponin-t splicing.
Earlier in this paper I stated that it is not always viable or possible to affect phenotypes by varying gene expression levels, and that in such cases AS is particularly important. Tnt is an excellent illustration of this principal because it must be present in a certain stoichiometric ratio with other proteins (in this case, with regularly spaced binding sites on actin and tropomyosin, which together with troponin make up muscle thin filaments). Tnt can only function when complexed with these other molecules at spatially defined and numerically limited binding sites, which appear to be always saturated with troponins and tropomyosin. Thus, there is little or no opportunity for changes in Tnt gene expression level to have a functional effect; rather it is changes in the type of protein that is made and incorporated into binding sites in muscle that determines functional variation.
Case study 4: plasticity of plant growth form in response to activity of an ethylene responsive kinase that affects AS
The tobacco gene PK12 encodes a kinase that is induced by ethylene, a signaling molecule that affects diverse aspects of plant growth, development and stress responses. PK12 belongs to a class of LAMMER kinases, of which all known homologs in plants and animals phosphorylate and affect the nuclear distribution of splicing factors, the proteins that control both constitutive and AS of other genes. These characteristics identified PK12 as a candidate gene in ethylene-controlled signaling pathways (Savaldi-Goldstein et al., 1996).
To examine the in vivo effects of PK12, a construct of the tobacco gene was introduced into Arabidopsis (which contains a homologous protein with 73% amino-acid identity) where it was shown to bind, phosphorylate and colocalize with the Arabidopsis splicing factor atSRp34/SR1 (Savaldi-Goldstein et al., 2003). Lines expressing high levels of the transgene had high PK12 kinase activity and showed altered patterns of AS in three of the four splicing factor genes that were examined (Figure 4a), i.e. the kinase had both broad and specific effects on AS of splicing factors. The same lines showed changes in growth form (Figure 4b) and lifespan under normal light/dark conditions, and differed in plasticity of growth under constant dark versus constant light conditions (Figure 4c).
(a) Intron–exon structure and relative abundance of alternatively spliced transcripts of three Arabadopsis genes (atSRp34/SR1; U1-70K; ANP1) in plants that have either wild type or transgenically increased expression of a gene (PK12:HA) that encodes a LAMMER kinase. Two of these genes (atSRp34/SR1; U1-70) and another not shown (asSrp30) had changed patterns of AS, whereas ANP1 showed no change. (b) Differences in growth form of seedlings and mature leaves of wild-type and PK12 transgenic plants grown in normal light:dark cycles. (c) Changes in the plastic response of growth form in different light regimens (3d constant dark or 5d constant light). Transgenic seedlings show decreased plasticity of growth form, especially their root length. Adapted from Savaldi-Goldstein et al. (2003).
AS of target genes downstream of the affected splicing factors was not examined (no such plant genes are yet known), but transcriptome-wide expression levels were examined using microarrays. This survey identified 48 genes that were up- or downregulated by at least 1.6-fold.
One of the things that this study nicely shows is that plasticity of organism-level traits can depend on how AS pathways are activated by and respond to environmental signals. Plant biologists need to look no farther than this study by Savaldi-Goldstein et al. (2003) to find some excellent candidates for ‘plasticity genes’.
Hypotheses regarding polymorphism in quantitative patterns of AS
At this point it is interesting to consider the possible existence of heritable variation that affects quantitative patterns of AS and phenotypic responses to environmental cues. Based on the two case studies presented above for insect egg production and muscle performance, it is easy to see that there might exist morphs that take more or less risk in terms of how they adjust energy expenditure according to nutritional status and the likelihood of starvation. A more risk-prone morph could have greater mating success in the short term, but reduced survival in the long term. Spatial and temporal variation in population density and environmental quality could favor different levels of risk and could possibly maintain genetic variation at the level of populations or metapopulations, in a manner somewhat analogous to what has recently been shown for polymorphisms in flight metabolic rate and dispersiveness in butterflies (Haag et al., 2005; Hanski and Saccheri, 2006). At this point such a scenario is purely hypothetical for polymorphisms in quantitative variation in AS, but the case studies presented here lay the ground work for pursuing such research.
Regulation of AS: heuristic illustration of opportunities for polymorphisms that could affect phenotypic plasticity and reaction norms
As described above in the plant growth example, loci that regulate AS provide attractive candidates for alleles that cause variation in regulation and expression of plastic responses and reaction norms. Knowledge of genes and pathways that control AS is beginning to accumulate (reviewed by Stamm, 2002; Black, 2003). Rather than rehash that material here, I instead present as an example the regulation of AS in vertebrate troponin-t. Not only is this one of the best understood genes in terms of AS regulation, it also overlaps with one of the case studies presented above, thereby providing an opportunity to examine this case in a broader perspective. The combination of regulatory and phenotypic knowledge accumulated for this gene provides insight regarding possible ways that genotypic variation within populations could cause heritable variation in AS phenotypes and reaction norms.
Recent studies have provided a fairly detailed understanding of the regulation of alternative exon 5 in mini-gene constructs of human and chicken cardiac troponin-t (Charlet et al., 2002; Ladd et al., 2005). The splicing factor CUG-binding protein (CUG-BP) binds to the hexanucleotide CUGCUG in the intron downstream of alternative exon 5. Binding of CUG-BP to that site acts to increase the inclusion of exon 5 in mature transcripts. The exon inclusion action of CUG-BP is antagonized by polypyrimidine tract binding protein (PTB), which binds to a number of intron sites upstream and downstream of alternative exon 5. Polymerization among neighboring PTBs is thought to block access to other intron sites and thereby prevent binding by CUG-BP.
A third family of RNA binding proteins (muscleblind; MBNL) also antagonizes CUG-BP and determines AS of not just Tnt but also of the vertebrate insulin receptor gene (Ho et al., 2004). Because of this coregulation, the composition of the transcript pools for vertebrate troponin-t and insulin receptor are highly correlated. This is particularly interesting in light of the observation that AS of the Tnt gene in dragonflies varies according to energy reserves. Nutritionally variable coregulation of AS in a muscle gene that affects energy expenditure, and an insulin receptor that affects cellular energy uptake, stimulates the novel hypothesis that coregulated AS may coordinate cellular energetics throughout the body.
There are many places in the Tnt AS pathway where allelic variation could create different reaction norms. For example, single nucleotide mutations in and around the CUGCUG binding motif of the intron flanking alternative exon 5 could alter the way that CUG-BP binds at this site and thereby affect inclusion of exon 5. This would presumably affect only troponin-t splicing, with few pleiotropic effects. No such polymorphisms are yet known, but an insertion/deletion polymorphism of a five base pair sequence (CTTCT) in intron 3 of human cardiac Tnt has been discovered (Komamura et al., 2004); it affects the relative abundance of transcripts containing exon 4 and is significantly associated with increased mass of the left ventricle and thickness of the interventricular septal wall. These phenotypes are not likely to be a direct result of changes in AS of Tnt; more likely they are a secondary response to changes in the contractile performance of muscle fibers carrying a different mix of Tnt protein isoforms (i.e. muscle hypertrophy to compensate for weaker contraction).
Allelic variation within CUG or PTB genes could have similar effects on troponin-t splicing, but would probably involve many more pleiotropic and epistatic effects as these transcription factors also control AS in other genes. This is probably true for splicing factors in general. Another place where allelic variation for plastic responses could generally occur is within an alternative exon. In that case, phenotypes would presumably appear only when the exon is included in the mature transcript. Polyphenisms that vary in occurrence among genotypes (Braendle et al., 2005) could potentially arise through such a mechanism. Another interesting feature of amino-acid variability within alternative exons is that such loci would not consistently associate with phenotypes in mapping studies if subjects vary in the extent to which they express the polymorphic alternative exon.
The splicing regulators CUG and PTB are themselves alternatively spliced, so there is much additional complexity in this regulatory pathway, with many potential sites for loci that cause variation in quantitative and plastic traits. It is common for AS pathways to feature AS at many levels, often in a cascading manner (e.g., sex determination in Drosophila; Lopez, 1998). This makes elucidation and experimental manipulation of AS pathways a challenging task, all the more so because little of the relevant information is contained at the level of genomic sequence, and much of it is not easily detected by conventional forms of gene expression profiling.
The future: systems level analyses of AS
An approaching frontier is the ability to use exon-specific arrays (e.g. Johnson et al., 2003; Srinivasan et al., 2005; Shai et al., 2006) to quantify AS across many genes and to determine how AS is connected to other forms of variation in gene expression, as well as to allelic variation, signaling pathways and environmental variables. The basic idea is that each AS gene must have a number of array probes that hybridize to constitutive exons only, AS exons only and/or junctions between exons. Comparisons of the amount of hybridization to these probes can be used to quantify AS patterns, although independent verification remains necessary for at least some of the genes or for genes of particular interest. Methods for designing and interpreting data from AS-specific arrays (e.g. Le et al., 2004; Shai et al., 2006) are still in their infancy, but it is encouraging to note that there are now companies that state this as their primary goal. Development of methods to examine transcriptome wide changes in AS, in parallel with global gene expression phenotypes, will enable new ways to examine genetic mechanisms and molecular pathways by which variable phenotypes are generated; this in turn will allow examinations of how allelic variation affects plastic responses. Studying ‘plasticity alleles’ in an evolutionary context will provide a mechanistic basis for more detailed investigations of the evolution of phenotypic plasticity and reaction norms, and will surely be a component of what has been hailed as a post-modern synthesis that fuses developmental and evolutionary biology (West-Eberhard, 2003).
General conclusions
The explosion of genomic, SNP and microarray expression data provide ample material to search for genetic polymorphisms that affect phenotypes and potentially respond to selection. However, investigations often begin not from bioinformatic data but from observations of organismal traits; this in turn leads to a search for the underlying genetic architecture. Material that I have presented here argues that there should be a general awareness that in many cases the phenotypic variation of interest could arise from the way genes are alternatively spliced. In order to discover such mechanisms it is not sufficient to amplify, clone and sequence a fragment of a candidate gene and subsequently look only for allelic variation or differences in expression level, or to perform subtractive hybridization or to analyze only genomic data; such approaches ignore and overlook the possibility of variation in the relative abundance of alternative transcripts. Evolutionary biologists and quantitative geneticists should be aware that quantitative variation in alternative transcripts among individuals is an interesting trait that is both plastic and likely to vary among genotypes. The fact that quantitative variation in AS is so seldom measured and has never been assessed for heritability in any organism identifies this area as an important lacuna in biology. In relation to well-trodden areas of modern biology, exploration of intraspecific variability in AS in relation to environmental and genetic variation is an open field that promises to be especially productive.
References
Alonso-Blanco C, Mendez-Vigo B, Koornneef M (2005). From phenotypic to molecular polymorphisms involved in naturally occurring variation of plant development. Int J Dev Biol 49: 717–732.
Baek D, Green P (2005). Sequence conservation, relative isoform frequencies, and nonsense-mediated decay in evolutionarily conserved alternative splicing. Proc Natl Acad Sci USA 102: 12813–12818.
Bayer CA, Kalm L, Fristrom JW (1997). Relationships between protein isoforms and genetic functions demonstrate functional redundancy at the Broad-Complex during Drosophila metamorphosis. Dev Biol 187: 267–282.
Birikh KR, Sklan EH, Shoham S, Soreq H (2003). Interaction of ‘readthrough’ acetylcholinesterase with RACK1 and PKCII correlates with intensified fear-induced conflict behavior. Proc Natl Acad Sci USA 100: 283–288.
Black DL (2003). Mechanisms of alternative pre-messenger RNA splicing. Annu Rev Biochem 72: 291–336.
Braendle C, Friebe I, Caillaud MC, Stern DL (2005). Genetic variation for an aphid wing polyphenism is genetically linked to a naturally occurring wing polymorphism. Proc Biol Sci 272: 657–664.
Britton J, Lockwood WK, Li L, Cohen SM, Edgar BA (2002). Drosophila's insulin/PI3-kinase pathway coordinates cellular metabolism with nutritional conditions. Dev Cell 2: 239–249.
Brotto MA, Biesiadecki BJ, Nosek TM, Jin JP (2006). Coupled expression of troponin T and troponin I isoforms in single skeletal muscle fibers correlates with contractility. Am J Physiol Cell Physiol 290: C567–C576.
Chandra M, Tschirgi ML, Rajapakse I, Campbell KB (2006). Troponin T modulates sarcomere length-dependent recruitment of cross-bridges in cardiac muscle. Biophys J 90: 2867–2876.
Charlet BN, Logan P, Singh G, Cooper TA (2002). Dynamic antagonism between ETR-3 and PTB regulates cell type-specific alternative splicing. Mol Cell 9: 649–658.
Chen BE, Kondo M, Garnier A, Watson FL, Puettmann-Holgado R, Lamar DR et al. (2006a). The molecular diversity of Dscam is functionally required for neuronal wiring specificity in Drosophila. Cell 125: 607–620.
Chen FC, Wang SS, Chen CJ, Li WH, Chuang TJ (2006b). Alternatively and constitutively spliced exons are subject to different evolutionary forces. Mol Biol Evol 23: 675–682.
Copley RR (2004). Evolutionary convergence of alternative splicing in ion channels. Trends Genet 20: 171–176.
Cota D, Proulx K, Smith KA, Kozma SC, Thomas G, Woods SC et al. (2006). Hypothalamic mTOR signaling regulates food intake. Science 312: 927–930.
Cusack BP, Wolfe KH (2005). Changes in alternative splicing of human and mouse genes are accompanied by faster evolution of constitutive exons. Mol Biol Evol 22: 2198–2208.
Dann SG, Thomas G (2006). The amino acid sensitive TOR pathway from yeast to mammals. FEBS Lett 580: 2821–2829.
Falconer DS, Mackay TFC (1996). Introduction to Quantitative Genetics, 4th edn. Pearson/Prentice Hall: Harlow.
Flood WD, Ruvinsky A (2001). Alternative splicing and expressivity of the Axin(Fu) allele in mice. Heredity 87: 146–152.
Gabriel W, Luttbeg B, Sih A, Tollrian R (2005). Environmental tolerance, heterogeneity, and the evolution of reversible plastic responses. Am Nat 166: 339–353.
Garcia-Blanco MA, Baraniak AP, Lasda EL (2004). Alternative splicing in disease and therapy. Nat Biotechnol 22: 535–546.
Gershenfeld HK, Paul SM (1997). Mapping quantitative. Trait loci for fear-like behaviors in mice. Genomics 46: 1–8.
Gomes AV, Guzman G, Zhao J, Potter JD (2002). Cardiac troponin T isoforms affect the Ca2+ sensitivity and inhibition of force development. Insights into the role of troponin T isoforms in the heart. J Biol Chem 277: 35341–35349.
Gomes AV, Venkatraman G, Davis JP, Tikunova SB, Engel P, Solaro RJ et al. (2004). Cardiac troponin T isoforms affect the Ca(2+) sensitivity of force development in the presence of slow skeletal troponin I: insights into the role of troponin T isoforms in the fetal heart. J Biol Chem 279: 49579–49587.
Haag CR, Saastamoinen M, Marden JH, Hanski I (2005). A candidate locus for variation in dispersal rate in a butterfly metapopulation. Proc R Soc Lond B 272: 2449–2456.
Hanski I, Saccheri I (2006). Molecular-level variation affects population growth in a butterfly metapopulation. PLoS Biol 4: e129.
Hartl DL, Clark AG (1997). Principles of Population Genetics, 3rd edn. Sinauer Associates: Sunderland, MA.
Hernandez OM, Szczesna-Cordary D, Knollmann BC, Miller T, Bell M, Zhao J et al. (2005). F110I and R278C troponin T mutations that cause familial hypertrophic cardiomyopathy affect muscle contraction in transgenic mice and reconstituted human cardiac fibers. J Biol Chem 280: 37183–37194.
Herranz R, Mateos J, Mas JA, Garcia-Zaragoza E, Cervera M, Marco R (2005). The coevolution of insect muscle TpnT and TpnI gene isoforms. Mol Biol Evol 22: 2231–2242.
Ho TH, Charlet BN, Poulos MG, Singh G, Swanson MS, Cooper TA (2004). Muscleblind proteins regulate alternative splicing. EMBO J 23: 3103–3112.
Izem R, Kingsolver J (2005). Variation in continuous reaction norms: Quantifying directions of biological interest. Am Nat 166: 277–289.
Johnson JM, Castle J, Garrett-Engele P, Kan Z, Loerch PM, Armour CD et al. (2003). Genome-wide survey of human alternative pre-mRNA splicing with exon junction microarrays. Science 302: 2141–2144.
Knies JL, Izem R, Supler KL, Kingsolver JG, Burch CL (2006). The genetic basis of thermal reaction norm evolution in lab and natural phage populations. PLoS Biol 4: e201.
Komamura K, Iwai N, Kokame K, Yasumara Y, Kim J, Yamagishi M et al. (2004). The role of a common TNNT2 polymorphism in cardiac hypertrophy. J Hum Genet 49: 129–133.
Ladd AN, Stenberg MG, Swanson MS, Cooper TA (2005). Dynamic balance between activation and repression regulates pre-mRNA alternative splicing during heart development. Dev Dyn 233: 783–793.
Lazaridis KN, Tietz P, Wu T, Kip S, Dawson PA, LaRusso NF (2000). Alternative splicing of the rat sodium/bile acid transporter changes its cellular localization and transport properties. Proc Natl Acad Sci USA 97: 11092–11097.
Le K, Mitsouras K, Roy M, Wang Q, Xu Q, Nelson SF et al. (2004). Detecting tissue-specific regulation of alternative splicing as a qualitative change in microarray data. Nucleic Acids Res 32: e180.
Leimar O (2005). The evolution of phenotypic polymorphism: randomized strategies versus evolutionary branching. Am Nat 165: 669–681.
Lipscombe D (2005). Neuronal proteins custom designed by alternative splicing. Curr Opin Neurobiol 15: 358–363.
Lopez AJ (1998). Alternative splicing of pre-mRNA: developmental consequences and mechanisms of regulation. Annu Rev Genet 32: 279–305.
MacFarland SM, Jin JP, Brozovich FV (2002). Troponin T isoforms modulate calcium dependence of the kinetics of the cross-bridge cycle: studies using a transgenic mouse line. Arch Biochem Biophys 405: 241–246.
Mackay TF, Lyman RF (2005). Drosophila bristles and the nature of quantitative genetic variation. Philos Trans R Soc Lond B Biol Sci 360: 1513–1527.
Malko DB, Makeev VJ, Mironov AA, Gelfand MS (2006). Evolution of exon-intron structure and alternative splicing in fruit flies and malarial mosquito genomes. Genome Res 16: 505–509.
Marden JH, Cobb JR (2004). Territorial and mating success of dragonflies that vary in muscle power output and presence of gregarine gut parasites. Anim Behav 68: 857–865.
Marden JH, Fitzhugh GH, Wolf MR, Arnold KD, Rowan B (1999). Alternative splicing, muscle calcium sensitivity, and the modulation of dragonfly flight performance. Proc Natl Acad Sci USA 96: 15304–15309.
Marden JH, Fitzhugh GH, Girgenrath M, Wolf MR, Girgenrath S (2001). Alternative splicing, muscle contraction and intraspecific variation: associations between troponin T transcripts, Ca(2+) sensitivity and the force and power output of dragonfly flight muscles during oscillatory contraction. J Exp Biol 204: 3457–3470.
Meshorer E, Erb C, Gazit R, Pavlovsky L, Kaufer D, Friedman A et al. (2002). Alternative splicing and neuritic mRNA translocation under long-term neuronal hypersensitivity. Science 295: 508–512.
Meshorer E, Soreq H (2006). Virtues and woes of AChE alternative splicing in stress-related neuropathologies. Trends Neurosci 29: 216–224.
Meshorer E, Bryk B, Toiber D, Cohen J, Dori A, Soreq H (2005). SC35 promotes sustainable stress-induced alternative splicing of neuronal acetylcholinesterase mRNA. Mol Psych 10: 985–997.
Nassar R, Malouf NN, Mao L, Rockman HA, Oakeley AE, Frye JR et al. (2005). cTnT1, a cardiac troponin T isoform, decreases myofilament tension and affects the left ventricular pressure waveform. Am J Physiol Heart Circ Physiol 288: H1147–H1156.
Nussey DH, Postma E, Gienapp P, Visser ME (2005). Selection on heritable phenotypic plasticity in a wild bird population. Science 310: 304–306.
Ogut O, Granzier H, Jin JP (1999). Acidic and basic troponin T isoforms in mature fast-twitch skeletal muscle and effect on contractility. Am J Physiol 276: C1162–C1170.
Pigliucci M, Pollard H, Cruzan MB (2003). Comparative studies of evolutionary responses to light environments in Arabidopsis. Am Nat 161: 68–82.
Pollard VW, Malim MH (1998). The HIV-1 Rev protein. Annu Rev Microbiol 52: 491–532.
Rossi EL (2004). Stress-induced alternative splicing in mind body medicine. Adv Mind Body Med 20: 12–19.
Savaldi-Goldstein S, Aviv D, Davydov O, Fluhr R (2003). Alternative splicing modulation by a LAMMER kinase impinges on developmental and transcriptome expression. Plant Cell 15: 926–938.
Savaldi-Goldstein S, Sessa G, Fluhr R (1996). The ethylene-inducible PK12 kinase mediates the phosphorylation of SR splicing factors. Plant J 21: 91–96.
Shai O, Morris QD, Blencowe BJ, Frey BJ (2006). Inferring global levels of alternative splicing isoforms using a generative model of microarray data. Bioinformatics 22: 606–613.
Shemesh R, Novik A, Edelheit S, Sorek R (2006). Genomic fossils as a snapshot of the human transcriptome. Proc Natl Acad Sci USA 103: 1364–1369.
Sklan EH, Lowenthal A, Korner M, Ritov Y, Landers DM, Rankinen T et al. (2004). Acetylcholinesterase/paraoxonase genotype and expression predict anxiety scores in Health, Risk Factors, Exercise Training, and Genetics study. Proc Natl Acad Sci USA 101: 5512–5517.
Srinivasan K, Shiue L, Hayes JD, Centers R, Fitzwater S, Loewen R et al. (2005). Detection and measurement of alternative splicing using splicing-sensitive microarrays. Methods 37: 345–359.
Stamm S (2002). Signals and their transduction pathways regulating alternative splicing: a new dimension of the human genome. Hum Mol Genet 11: 2409–2416.
Stamm S, Ben-Ari S, Rafalska I, Tang Y, Zhang Z, Toiber D et al. (2005). Function of alternative splicing. Gene 344: 1–20.
Sternfeld M, Shoham S, Klein O, Flores-Flores C, Evron T, Idelson GH et al. (2000). Excess ‘read-through’ acetylcholinesterase attenuates but the ‘synaptic’ variant intensifies neurodeterioration correlates. Proc Natl Acad Sci USA 97: 8647–8652.
Stetefeld J, Ruegg MA (2005). Structural and functional diversity generated by alternative mRNA splicing. Trends Biochem Sci 30: 515–521.
Stoltzfus CM, Madsen JM (2006). Role of viral splicing elements and cellular RNA binding proteins in regulation of HIV-1 alternative RNA splicing. Curr HIV Res 4: 43–55.
Su Z, Wang J, Yu J, Huang X, Gu X (2006). Evolution of alternative splicing after gene duplication. Genome Res 16: 182–189.
Terashima J, Bownes M (2004). Translating available food into the number of eggs laid in Drosophila melanogaster. Genetics 167: 1711–1719.
Wada K, Sakaguchi H, Jarvis ED, Hagiwara M (2004). Differential expression of glutamate receptors in avian neural pathways for learned vocalization. J Comp Neurol 476: 44–64.
Wang BB, Brendel V (2006). Genomewide comparison of alternative splicing in plants. Proc Natl Acad Sci USA 103: 7175–7180.
Watson FL, Puttmann-Holgado R, Thomas F, Lamar DL, Hughes M, Kondo M et al. (2005). Extensive diversity of Ig-superfamily proteins in the immune system of insects. Science 309: 1874–1878.
Wayne ML, Korol A, Mackay TF (2005). Microclinal variation for ovariole number and body size in Drosophila melanogaster in ‘Evolution Canyon’. Genetica 123: 263–270.
West-Eberhard MJ (2003). Developmental Plasticity and Evolution. Oxford University Press: Oxford.
West-Eberhard MJ (2005). Developmental plasticity and the origin of species differences. Proc Natl Acad Sci USA 102: 6543–6549.
Xing Y, Lee C (2005). Evidence of functional selection pressure for alternative splicing events that accelerate evolution of protein subsequences. Proc Natl Acad Sci USA 102: 13526–13531.
Zhang XS (2005). Evolution and Maintenance of the Environmental Component of the Phenotypic Variance: Benefit of Plastic Traits under Changing Environments. Am Nat 166: 569–580.
Acknowledgements
I thank H Fescemyer and two reviewers for constructive comments on the manuscript. This work was supported by NSF EF-0412651.
Author information
Authors and Affiliations
Corresponding author
Additional information
Note added in proof
A new paper addressing immune function in Anopholes mosquitos has shown that alternative splicing of the Dscam gene produces pathogen-specific pattern recognition receptors that mediate phagocytosis of bacteria and the malarial parasite Plasmodium. Silencing of specific splice forms compromises resistance in a pathogen-specific manner. This study reveals that AS may be the key feature allowing plasticity and broad effectiveness of invertebrate immune systems.
Dong Y, Taylor HE, Dimopoulos G (2006). AgDscam, a hypervariable immunoglobulin domain-containing receptor of the Anopheles gambiae innate immune system. PLoS Biol 4: e229.
Rights and permissions
About this article
Cite this article
Marden, J. Quantitative and evolutionary biology of alternative splicing: how changing the mix of alternative transcripts affects phenotypic plasticity and reaction norms. Heredity 100, 111–120 (2008). https://doi.org/10.1038/sj.hdy.6800904
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800904
Keywords
This article is cited by
-
The evolutionary significance of post-transcriptional gene regulation
Heredity (2024)
-
Genetic constraints in genes exhibiting splicing plasticity in facultative diapause
Heredity (2024)
-
Alternative splicing is highly variable among Daphnia pulex lineages in response to acute copper exposure
BMC Genomics (2020)
-
Identification of genes underlying phenotypic plasticity of wing size via insulin signaling pathway by network-based analysis in Sogatella furcifera
BMC Genomics (2019)
-
Adaptive Transcriptome Profiling of Subterranean Zokor, Myospalax baileyi, to High- Altitude Stresses in Tibet
Scientific Reports (2018)