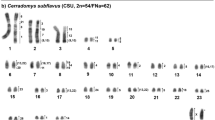
Abstract
West African gerbils of the genus Taterillus constitute a complex of seven sibling species distributed from sudano-guinean to saharo-sahelian regions. They display radically rearranged karyotypes despite low genic divergence and a very recent differentiation, that is, within the last 0.4 Myr for the six most derived species. We here provide a comparison of the seven specific karyotypes and perform a cladistic analysis using chromosomal rearrangements character states. When a posteriori polarized mutations were mapped onto the phylogenetic tree, 38 rearrangements were identified as fixed during the evolution of these rodents. This makes Taterillus one of the most striking examples of accelerated chromosomal evolution within placental mammals. Taking into account the types of chromosomal changes involved, divergence times between lineages, genetic distances, as well as reassessed geographic distributions, we suggest that (1) speciation in West African Taterillus was driven by chromosomal changes, and (2) the paleoclimatic oscillations of the Sahara desert have played a major role in their evolution. In particular, elevated plasticity of the Taterillus genome, as suggested by the patterns observed for some repetitive elements, would have led to a higher probability of mutation. We hypothesize that the process underpinning cladogenesis most probably involved highly underdominant genomic rearrangements that were fixed following pronounced populational bottlenecks resulting from drastic climatic and subsequent environmental changes. Major African rivers formed significant barriers to dispersal, limiting expansion during the more moist and so favorable periods. This scenario would explain the current parapatric species distributions and their relationship to the West African hydrographic features.
Similar content being viewed by others
Introduction
There has been extensive debate on the role of chromosomal changes in evolution (reviews in Sites and Moritz, 1987; King, 1993; Rieseberg, 2001). In particular, opposite views have persisted about the ability of chromosomal changes to initiate reproductive isolation and thus drive speciation. Arguments against chromosomal speciation are two fold. The first one relies on some examples of important polymorphism in natural populations (Nachman and Myers, 1989), which implies that chromosomal changes are probably not strongly underdominant, and consequently cannot insure reproductive isolation by themselves (Patton and Sherwood, 1983). Although acceptable in some cases, this argument is not generally applicable since many species display very stable karyotypes, high polymorphism being more the exception than the rule. On the contrary, numerous studies have clearly demonstrated that few or even a single chromosomal rearrangements can have dramatic effects on meiosis in heterozygous carreers (King, 1993).
A second argument against chromosomal speciation is that chromosomal structural rearrangements were too deleterious and required too drastic- and hence rare-population conditions to be fixed (see Patton and Sherwood, 1983; Chesser and Baker, 1986). This argument looses credence however since most species have species-specific karyotypes, meaning that the differences, that is, the karyotypic changes, have necessarily been fixed either during or subsequently to the speciation event.
As with other speciation processes, the difficulty with identifying promoting chromosomal change as a driver of speciation is one usually investigates groups that already display morphologic, eco-ethologic and/or genetic divergences (see Zhang et al, 2004 and references therein, for the example of the highly debated case of human and chimpanzee). It is thus often difficult to assess the exact sequence of events leading to reproductive isolation and to identify the first step of differentiation process. However, several studies have recently suggested that karyotype repatterning plays a significant role in speciation in some organisms. For instance, in Drosophila, inversions have clearly been shown to reduce gene flow between inverted segments, thus allowing differentiation of coadapted genes complexes (‘suppressed recombination model’ sensu Noor et al, 2001; Ayala and Coluzzi, 2005; reviewed in Rieseberg, 2001; Hoffman et al, 2004; Ayala and Coluzzi, 2005). Similarly, translocations have been proved to lead to a strong decrease in reproductive fitness in engineered yeast hybrids with a controlled genomic background (Delneri et al, 2003; ‘hybrid dysfuntion model’ sensu Ayala and Coluzzi, 2005).
However, the identification of additional examples, particularly in natural settings is required to further evaluate the importance of chromosomal speciation as an evolutionary process. Unfortunately, experiments involving controlled manipulations in the laboratory, such as the one performed by Delneri et al (2003), are hardly realistic when dealing with natural populations. Nevertheless, sibling species complexes offer an opportunity to control some of the myriad of variables associated with natural populations allowing testable hypotheses based on reasonable assumptions. Indeed, sibling species are reproductively isolated but morphologically similar. Therefore, this phenotypic similarity may reflect a recent differentiation, so that not enough time has elapsed for any significant genetical and eco-ethological divergence. Although some examples of sibling but genetically well-differentiated species have been provided (eg, Arvicanthis spp, Ducroz et al, 1998), many cryptic taxa have been karyotypically differentiated and have proved to be recently diverged (Elder, 1980; Taylor, 2000; Volobouev et al, 2002). As such, once these prerequisites are confirmed, sibling species complex constitute valuable biological models for investigating the effect of chromosomal changes in evolution.
Such a complex of sibling species of recent origin is represented by the African genus Taterillus (Rodentia, Gerbillinae), and more specially by the West African species. The latter are a monophyletic relative to the Eastern African species, a conclusion which is supported by (1) two synapomorphic autosome–gonosome translocations, one of which results in a sex trivalent XY1Y2 in males (reviewed in Dobigny, 2002), and (2) mitochondrial DNA sequence data (Dobigny, 2002). These murid rodents have previously been shown to exhibit morphological conservatism in both size and shape due to adaptative constraints and/or very recent differentiation (Dobigny et al, 2002b). Previous cytotaxonomic surveys have allowed the identification of several sibling taxa within the West African lineage: T. gracilis and T. pygargus (2n=36/37 and 2n=22/23, respectively; Matthey and Jotterand, 1972), T. arenarius (2n=30/31; Robbins, 1974), T. petteri (2n=18/19, Sicard et al, 1988), Taterillus sp2 (2n=24/25; Dobigny et al, 2002c), T. tranieri (2n=14/15; Dobigny et al, 2003a) and Taterillus sp1 (2n=22/23; Granjon and Dobigny, 2003). Pairwise comparisons between species have shown that these karyotypic differences were sufficient to insure postzygotic reproductive isolation (Dobigny et al, 2002a, 2003a). Molecular analyses using mitochondrial gene sequences showed that the West African Taterillus display very low interspecific genetic distances. In particular, they never exceed 15% between T. gracilis and the six most derived West African species, whereas variations within the latter range between 7 and 1.9% (Kimura two parameter model, applied to partial cytochrome b sequences of 796 bp; Dobigny, 2002). Therefore, the genetic diversity of these six species lies within the intraspecific range usually observed in rodents (Bradley and Baker, 2001). This low mitochondrial nucleotide divergence was confirmed by nuclear analyses involving nine enzymatic systems for which no variation was observed between T. pygargus and T. petteri (Benazzou, 1984). Taken together, the species-specific karyotypes, the very low genetic distance and the high morphological similarity make the West African Taterillus complex a useful model taxon for testing hypotheses of a causal role for chromosomal rearrangements in speciation.
Finally, admittedly on a limited amount of data, Petter (1974) observed that most of the West African species show geographic patterns shaped by the Senegal and Niger rivers. On this basis, he proposed that these rivers may have been responsible for population isolations and subsequent allopatric speciation, and that these events took place during wet episodes during which rivers would have fragmented populations thus allowing allopatric speciation. Alternatively, Sicard et al (1988) favored the hypothesis of differentiation within small isolated demes during dry periods, and proposed that the parapatric distributions currently observed are the result of rivers acting as barriers to subsequent dispersal during wet episodes. To further investigate these two hypotheses, and to provide insights on the tempo and mode of evolution in Taterillus, we conducted wide-scale and nonambiguous cytotaxonomic surveys all over West Africa. We then use chromosomal rearrangements identified by comparative banding patterns in a cladistic analysis of the West African species. By taking the type of karyotypical changes that occurred into account, the extant species distributions, the timeframe provided by our previous molecular analyses, as well as eco-ethological and paleobiogeographical data, we propose an integrated hypothesis for the radiation of this group. This scenario implicates both a predisposition to genome repatterning and drastic populational fluctuations, due to paleoclimatic changes, as drivers of speciation in Taterillus.
Materials and methods
In total, 136 specimens were sampled from 41 localities in Mauritania, Senegal, Mali, Burkina-Faso, Niger, and Chad during several rodent surveys (Benazzou, 1984; Volobouev and Granjon, 1996; Dobigny et al, 2002a, 2002b, 2003a; Granjon and Dobigny, 2003; Granjon et al, 2004; Granjon, unpublished) and used for this study (see Table 1). Fibroblast cell cultures were set up from biopsies for 32 individuals, and metaphases were obtained following standard procedures. Metaphases for the 104 remaining specimens were obtained using direct karyotyping of bone-marrow cells (Lee and Elder, 1980). The diploid and fundamental numbers, as well as the morphology of all chromosomes were assessed for each individual. GTG banding (Seabright, 1971) and CBG banding (Sumner, 1972) were performed on 30 and 27 of the test animal, respectively.
GTG-banding patterns were compared between several representatives of all identified cytotypes (corresponding to eight species; see below), and chromosomal homologies were assessed (Table 2). A phylogenetic analysis was performed according to cladistics concepts and methods as reviewed in Dobigny et al (2004a): 46 nonpolarized structural chromosome rearrangements, excluding heterochromatin changes, were used as binary characters, and their presence/absence was coded as character states (Table 3). The East African species T. congicus, which displays XX/XY sex chromosomes, was used as an outgroup. The most parsimonious topologies were retrieved using an exhaustive search in PAUP 4.0b8 (Swofford, 1998). Robustness was evaluated using both decay index and bootstrap support (using 1000 replicates).
Species-specific distribution maps were redefined on the basis of data from literature (see above), this study, and the collections of the Muséum National d'Histoire Naturelle (France) and the Institut de Recherche pour le Développement (France). Only nonambiguous, that is, chromosomally validated sites were considered.
Results
All the karyotypes obtained could be assigned to one of the suite of previously described cytotypes (see above); GTG- and CBG-banding patterns were obtained from representatives of each. Importantly, G-banded karyotypes are presented for the first time for T. petteri (Figure 1) T. gracilis (Figure 2) and T. congicus (Figure 3). Polymorphism was detected in several species, confirming and extending earlier observations: Robertsonian changes in T. gracilis (2n=36/37 to 2n=38/39; Matthey and Jotterand, 1972; Dobigny et al, 2002a), pericentric inversions in T. pygargus (Dobigny et al, 2002b) and T. petteri (Tranier, 1974) and heterochromatin amplification in Taterillus sp1 (Dobigny et al, 2002b). Note that two cytotypes were considered for the cladistic analysis involving T. gracilis (2n=36/37 and 2n=38/29) in order to polarize this particular mutational event, although they are clearly conspecific (Dobigny et al, 2002b). Diploid (2n), autosomal fundamental (NFa) numbers, and the observed variation within each species are summarized in Table 1. C-banding patterns (see Figures 1, 2 and 3 for examples in T. petteri, T. gracilis and T. congicus; see Volobouev and Granjon, 1996; Dobigny et al, 2002a, 2003a, for the others) are markedly different between West and East African species. T. congicus possesses well-delineated C-positive heterochromatic blocks on all 18 acrocentric and subtelocentric pairs of its karyotype, whereas the C-positive regions in the other species were generally small and indistinct. Only T. gracilis has 10 acrocentric pairs possessing slightly C-positive pericentromeric regions.
G-banding patterns comparison allowed us to assess all the chromosomal homologies between the eight species (Table 2; examples of pairwise comparisons are provided in Dobigny et al, 2002a, 2003a) and to identify the structural differences among karyotypes. A cladistic analysis using these chromosomal rearrangements as characters led to two equally parsimonious (L=48 steps) trees (Figure 4) in which T. gracilis was the first taxon to diverge, followed by Taterillus sp2 and Taterillus sp1, and then T. arenarius, T. pygargus, T. petteri and T. tranieri. The only difference between the two topologies lies in the relative position of Taterillus sp1 and Taterillus sp2, whose relationships are not resolved in one of the topologies whereas Taterillus sp2 is more basal in the other one (Figure 4). No differences could be found between the Acctran and Deltran options (data not shown). The chromosomal changes were polarized a posteriori using the outgroup comparison criterion and then reported along the cladogram branches (Figure 4). This showed that Taterillus karyotype evolution involved at least 11 tandem fusions, six pericentric inversions, 18 or 19 Robertsonian fusions (tree 2 and 1, respectively) and three or four centric fissions (tree 1 and 2, respectively) within the ingroup. Furthermore, two mutations (n°35 and 21, Table 3) were found homoplasic, which can be explained as two convergent events, or one convergence and one reversal, depending on the tree considered (Figure 4). The consistency, retention and rescaled consistency indexes are CI=0.958, RI=0.929 and RC=0.89, respectively. The skewness value is g1=−1.025. Bootstrap support and decay index values range from 71 to 100% and from 1 to 4, respectively (Figure 4).
(a, b) The two most parsimonious trees (L=48) retrieved from our cladistic analysis of chromosomal characters in West African Taterillus, and (c) their consensus tree. The polarization of each character changes were assessed a posteriori using outgroup comparison. Gray circles, balck losanges, white blocks and gray triangles represent Robertsonian fusions, centric fissions, tandem fusions and pericentric inversions, repsectively. Close and open circles above these symbols correspond to convergent and reversal events, respectively. Bootstrap and decay index values are indicated above and below the branches of the consensus tree, respectively.
Discussion
Phylogenetic relationships between West African Taterillus
The consensus topology obtained for the eight cytotypes was well resolved with only one trifurcation. It is well supported by the CI, RI, RC and g1 statistics and by bootstrap and decay index values. In addition, the majority of the chromosome changes inferred correspond to Robertsonian changes, something which is commonly observed in mammals including gerbilline rodents (Qumsiyeh et al, 1987). More importantly, the cladistic method used here implies that character changes are not polarized a priori, thus allowing both fusions and fissions to be inferred (reviewed in Dobigny et al, 2004a). Yet, 19 (tree 1) or 18 (tree 2) of the rearrangements observed in this study were polarized a posteriori using the outgroup comparison criterion as fusions, with only three (tree 1) or four (tree 2) fissions inferred. In particular, none of the 11 tandem translocations coded here were found to be fission by the analysis. The occurrence of tandem fissions, even if theoretically possible, remains highly unprobable due to the requirement of both a neocentromere and neotelomeres for the fission products to be cytologically viable. As far as we know, this type of mutation has been very rarely documented in mammals (see Table 1 in Modi, 1987 for a possible example in arvicolid rodents). The fact that all of the 11 tandem mutations of our analysis were classified as fusions is, in that sense, reassuring. Finally, the only resolved and supported node previoiusly obtained using molecular phylogenetics, that is, T. gracilis as the most basal West African species (Dobigny, 2002), was also retrieved and strongly supported using our chromosomal dataset.
Cladistic analysis of karyotypic rearrangements has been proved to be a highly valuable mean for resolving phylogenetic relationships within several African murid sibling species complexes (Volobouev et al, 2002). The well-supported Taterillus phylogeny retrieved here, in the absence of sufficient morphological and genetic differences (at least according to current knowledge), provides another example of how the use of chromosomal characters can sometimes provide good insights into phylogenetics.
Accelerated rate of chromosomal evolution
Using the a posteriori polarized karyotypical rearrangements mapped along the branches of the phylogram (Figure 4), it is possible to evaluate the number of each type of chromosomal changes characterizing specific karyotypes. It should be emphasized that this includes the convergent changes, which remain cryptic when only two karyotypes are compared independently of a phylogenetic framework. This number ranges from 6 between T. pygargus (2n=22/23) and T. tranieri (2n=14/15) or T. petteri (2n=18/19), to 21 between T. gracilis (2n=36/37) and T. petteri or T. tranieri (see Figure 4). In total, at least 38 mutations have been fixed during the course of the West African Taterillus investigated here (Figure 4). Divergence times between Taterillus-specific lineages were estimated on molecular grounds, suggesting that East and West African species split ∼3.5 Myr ago, T. gracilis and the other West African species ∼2 Myr, whereas the six most derived West African species have radiated during the last 0.4 Myr (Dobigny, 2002; see also Chevret and Dobigny, 2005). This is one of the most recent speciation events documented in mammals. As such, it is of interest to compare the tempo of chromosomal change among these species to those observed in eutherian lineages usually considered as the most chromosomally active ones.
Dog-like carnivores provide several examples of quite extensive karyotypical evolution and are thought to display the highest genomic rate of evolution documented so far in placentals (see Table 1 in Murphy et al, 2003). For instance, the red and arctic foxes diverged only 2 Myr ago (Bininda-Emonds et al, 1999) but their karyotypes already differ by more than 30 karyotypical rearrangements (Nash et al, 2001). Extant gibbons constitute another well-documented case of genome repatterning since Hylobates hoolock and H. leucogenys differ by 62 (Müller et al, 2003) to 83 (Nie et al, 2001) rearrangements. Dates of divergence between gibbons lineages are still controversial but they probably diverged from each other soon after the hylobatids/great apes split, that is, approximately 5 Myr ago (Roos and Geissmann, 2001). Explosive chromosomal evolution is also found in equids, where the E. caballus karyotype differs from those of E. burchelli and E. zebra hartmannae by 23 and 32 rearrangements, respectively (Yang et al, 2003), whereas the four extant equid lineages are thought to have differentiated only recently (∼2.4 myr bp; Oakenfull and Clegg, 1998). Yet, the most striking example is provided by muntjak deer, with more than 30 karyotypic changes which have been fixed between the Indian and the Chinese muntjaks (see Figure 4 in Yang et al, 1997) since their last common ancestor between 0.9 and 3.7 Myr ago (Wang and Lan, 2000).
Although we are aware that other factors (such as generation time) should be taken into account for comparing evolutionary rates, we can approximate the tempo of chromosomal evolution by a simple ratio between the number of changes and times of divergence. Considering only the highest values available for the previous examples, the average rate of chromosomal change per myr reaches 13.3, 15, 16.6 and 33.3 in these equids, carnivores, primates and cervids, respectively. It was suggested that genomic rearrangements between mouse and rat proceed 10 times faster than between human and cat (Stanyon et al, 1999) suggesting high rates of chromosomal changes in some rodents. Indeed, several investigations in other rodent groups showed that they sometimes undergo extensive genome repatterning leading and/or accompanying rapid and successive speciation events (Bell et al, 2001; Volobouev et al, 2002). Although at the intraspecific level, the Robertsonian populations of house mice provide several striking examples of extremely rapid karyotypic evolution in rodents (eg, Britton-Davidian et al, 2000; reviewed in Pialek et al, 2005). Our analysis of West African Taterillus largely support this view as the most recently diverged karyospecies (T. tranieri or T. pygargus and Taterillus sp1 or Taterillus sp2; Figure 4) differ by 18 changes, leading to an average rate of 45 changes per myr.
The speciation process in Taterillus
The high interspecific chromosomal divergence is in sharp contrast with the low levels of morphological and molecular differences and are hardly compatible with the speciation models supporting a leading role of genetic divergence (Orr, 2001). Mutations on major genes, which could not have been detected by the molecular analyses (mainly on neutral markers) performed to date in Taterillus, may be responsible for these extremely rapid series of speciation events, or at least some of them. However, this type of process has never been demonstrated in mammals and appears quite unlikely to have occurred six times during the last 0.4 Myr (but see Forejt, 1996). As a consequence, although finer-scale genetic investigations would be needed to confirm such a conclusion, it seems probable that genic differences are not a valuable explanation for all speciation events in Taterillus.
Ecological and sexual selection speciation models have recently received renewed interest (Panhuis et al, 2001; Greenberg et al, 2003). These authors propose a role for disruptive selection, which could lead to eco-ethological differentiation and prezygotic isolation. Some evidence regarding these processes is available from eco-ethological and physiological studies were conducted in West African Taterillus species (Poulet, 1981; Hubert, 1982; Sicard, 1992). Unfortunately, all of them concentrate on one or two species, and only allow one to compare T. gracilis (the most basal and anciently diverged West African species) with T. pygargus or T. petteri (which both belong to the most recently derived clade). Experiments in captivity showed that F1 hybrids between T. gracilis and T. pygargus could be produced, but that these hybrids are sterile (Baron et al, 1974; Petter, unpublished). Although reproductive features are sometimes different between captivity and nature (eg, Forejt, 1996), this suggests a possible absence of prezygotic isolation and confirms the existence of a post-zygotic barrier. Furthermore, and probably more important, no sister species (as inferred from the topology provided on Figure 4) are known to live in sympatry (Figure 5). On the contrary, species distributions are clearly allopatric. Taken together, these data suggest that ecological and sexual speciation models may not be suitable for Taterillus biology and evolution.
Specific geographic distributions based only on nonambiguous cytotaxonomic identification. Data were obtained from this study as well as unpublished results from the Muséum National d'Histoire Naturelle and the Institut de Recherche pour le Développement, France. Main permanent and seasonal river courses which may have played a role in shaping Taterillus-specific distributions are indicated as solid and dash lines, respectively. Waves symbolize current surface of Lake Chad.
We recently presented pairwise comparisons between Taterillus sp1, Taterillus sp2, T. petteri and T. tranieri by reconstructing the very complex theoretical meiotic configurations in potential hybrids (see Dobigny et al, 2002a, 2003a). The latter strongly suggested postzygotic isolation due to important karyotypic differences. Here, we can extend these results to all West African species. Indeed, mapping of chromosomal changes on the tree demonstrate that all pairwise differences between species include at least one tandem fusion (and up to 8) and/or Robertsonian fusions with monobrachial homologies (Figure 4) which are both known to be highly deleterious (Baker and Bickham, 1986; Elder and Hsu, 1988; King, 1993).
Monobrachial homologies can be fixed very quickly, as observed in domestic mice on Madeira Island where 20 different fusions have been fixed within the last 500 years (Britton-Davidian et al, 2000). Tandem translocations have also been fixed in several mammalian lineages (eg, Rodentia: Elder, 1980; Fagundes et al, 1997; Artiodactyla: Yang et al, 1997; Carnivora: Nash et al, 2001; reviewed in King, 1993) although no case of polymorphism has been documented, with the possible exception of the bat Uroderma bilobatum. The latter species has been claimed to display cases of double heterozygotes for tandem translocations (see Table 1 in Owen and Baker, 2001). However, no banding data were provided by the authors that would have allowed for a nonambiguous identification of the rearrangements involved. Although prezygotic factors cannot be excluded, crossing experiments between Otomys cytotypes (Rodentia, Muridae) differing by a single tandem translocation were only 4% successful (Taylor, 2000). Together these data strongly suggest that (1) tandem translocations are highly deleterious for an heterozygous carrier, and that (2) when they occur and reach the homozygous state, they probably insure an almost instantly and complete reproductive isolation. If this is indeed the case, their presence in the homozygous state within a population would lead either to nearly instantaneous extinction or to the independant evolution (ie speciation) of the nascent cytotype. In Taterillus, tandem translocations were particularly numerous, mainly in the most recently speciated groups (Figure 4). The occurrence of this particular class of rearrangements was confirmed by an in situ hybridization survey showing the presence of Interstitial Telomeric Signals, which were correlated with the breakpoints of the tandem translocations inferred by the comparison of the GTG-banding patterns (Dobigny et al, 2003b). In conclusion, the successive fixation of monobrachial homology and tandem translocations during Taterillus genome repatterning must have been rapid events, maybe following an ‘all-or-none’ rule, and which may not have required other changes than chromosomal ones. This makes it reasonable to assume that the West African Taterillus have differentiated following karyotype repatterning, thus illustrating a new case of chromosomal speciation.
The probable role of paleoclimatic changes in West Africa
Considering the type of chromosome changes that led to Taterillus differentiation, it is also highly probable that the karyotypic changes characterizing the extant species were fixed in very small, isolated populations as predicted by the population cytogenetic models of Lande (1979, 1985) and Chesser and Baker (1986).
Thanks to a growing amount of nonambiguous cytotaxonomic data (see references above, this study, and unpublished data from the Museum National d'Histoire Naturelle and Institut de Recherche pour le Développement), it is now possible to precisely delimit the West African Taterillus species distributions (Figure 5). It is apparent that several of the species' ranges are parapatric and often demarcated by main river courses. In addition, the most derived species display rather narrow geographic ranges compared to the widely distributed T. gracilis, which occupies the Sahel and Sudan savannah from Senegal to Chad and Nigeria (Figure 5). Moreover, the six derived species are restricted to the northernmost arid Sahel regions up to the border of the Sahara desert (Figure 5). Interestingly, the latter areas were also the most influenced by the paleoclimatic variations of the Sahara (reviewed in Rognon, 1993). Indeed, during the last 4 Myr, the desert has undergone important changes, rendering its boundaries highly variable. On the one hand, many regions which are now dry where once much more humid, sometimes harboring steppe, savannah or even forest conditions. On the other hand, the desert has displayed periods of expansion, when the extant Sahelian region was a wide desert band of sand dunes (Rognon, 1993). The frequency of the Saharan cycles of desert spread and retraction are estimated to have occured at cycles of approximately 100 000–20 000 years during the last 3 Myr (Rognon, 1993). Taterillus species are not adapted to strictly desert environments and prefer subdesert to more moist conditions (Nowak, 1999). Assuming that their ecological preferences have not significantly changed during the last 4 Myr, it is very likely that these rapid and important paleoclimatic shifts have affected their geographic distribution and evolution.
Petter's (1974) hypothesis of speciation during wet periods following population fragmentation by great rivers does not fit the population cytogenetics patterns observed here, and thus appears highly improbable. Indeed, during such periods, environmental conditions must have been suitable for the expansion of Taterillus, allowing wide geographic distributions and consequently preventing cytogenetic drift. On the contrary, their phylogenetic relationships, their redefined geographic ranges and the chromosomal changes that characterize their evolution largely support Sicard et al's (1988) hypothesis. This proposes that during particularly dry periods, Taterillus individuals may have been trapped in refuge habitats that still displayed subarid conditions. These small and isolated demes would have provided the necessary population bottlenecks for underdominant karyotypic rearrangements to be stochastically fixed (Lande, 1979, 1985), thus leading to allopatric speciation. It is interesting to note that such refuges have already been identified, including the neighborhood of Lake Chad (cf. Granjon and Dobigny, 2003, and references therein), the coastal and the inner deltas of the Senegal and Niger rivers (see Rognon, 1993). This hypothesis is further supported by the more southern and wide distribution of the basal species, T. gracilis, suggesting this species has never benefited from conditions favoring the fixation of highly deleterious changes. If our conclusions are correct, the current pattern of parapatric-specific distributions result from expansion from discrete refugial areas during more favorable periods. These dispersal phases have subsequently been constrained by the main river courses as Taterillus cannot swim (Petter, personal communication).
Conclusion: population and genomic features required for allopatric chromosomal speciation
Since some organisms have fixed no or only a few karyotypical changes during very long periods of time (eg, Marsupialia: Rens et al, 2001; Perissodactyla, Rhinocerotidae: Trifonov et al, 2003), rapid genome repatterning, like that described here, must imply special conditions. In particular, it requires a high rate of both occurrence and fixation of chromosomal mutations.
As far as the Taterillus species are concerned, both the populations' demography and the genomes' lability may be appropriate. As suggested hereabove, the Sahara Desert's variation in both space (leading to the formation of different potential refuge habitats throughout the extant Sahel and Sahara regions) and time (the successive cycles of spreading/contraction of the desert) may have provided the population bottlenecks required for the recurrent fixation of chromosomal rearrangements. Furthermore, the Taterillus genome has been shown to display several atypical characteristics, which may reflect a high genomic plasticity. For instance, it displays 28S and 5S rDNA gene clusters that are variable between and within species both in number and location (Dobigny et al, 2003b), probably reflecting extensive fine-scale genomic reorganization. In addition, the explosive karyotype repatterning has been accompanied by amplification of LINE-1 retrotransposons (Dobigny et al, 2004b). Finally, T. congicus possesses C-positive pericentromeric blocks which are not observed in the West African species (Figure 6). Such highly repeated sequences may have favored the occurence of chromosome mutations such as Robertsonian fusions in the ancestral genome, and have been subsequently lost, as in case of the European house mouse (Garagna et al, 2001). All these properties suggest that the Taterillus genome is (or has recently been) is particularly plastic and may consequently have a high probability of chromosomal mutation.
This model of simultaneous and favorable conditions at both the populational and genomic levels would explain the very rapid fixation of strongly underdominant mutations. It has been proposed to explain extensive karyotypic evolution in muntjak deer (Wang and Lan, 2000). In the case of Taterillus, all genetic, cytogenetic and biogeographic data support such a view. It would be worth testing this hypothesis using population genetics surveys of microstatellite data, in order to detect and date populational bottlenecks in relation to paleoclimatic changes and the temporal phylogenetic framework now available.
References
Ayala FJ, Coluzzi M (2005). Chromosome speciation: humans, Dorosphila and mosquitoes. Proc Natl Acad Sci USA 102: 6535–6542.
Baker RJ, Bickham JW (1986). Speciation by monobrachial centric fusions. Proc Natl Acad Sci USA 83: 8245–8249.
Baron JC, Hubert B, Lambin P, Fine JM (1974). Serological differentiation of two species of Taterillus (Rodentia, Gerbillidae) from Senegal: T. gracilis (Thomas, 1892) and T. pygargus (Cuvier, 1832). Comp Biochem Physiol 47A: 441–446.
Bell DN, Hamilton NJ, Edwards CW, Wiggins LE, Martinez RM, Strauss RE et al (2001). Patterns of karyotypic megaevolution in Reithrodontomys: evidence from a cytochrome-b phylogenetic hypothesis. J Mammalogy 82: 81–91.
Benazzou T (1984). Contribution à l'étude chromosomique et de la diversification biochimique des Gerbillidés (Rongeurs). PhD Thesis, Université de Paris XI.
Bininda-Emonds OR, Gittleman JL, Purvis A (1999). Building large trees by combining phylogenetic information: a complete phylogeny of the extant Carnivora (Mammalia). Biol Rev 74: 143–175.
Bradley RD, Baker RJ (2001). A test of the genetic species concept: cytochrome-b sequences and mammals. J Mammalogy 82: 960–973.
Britton-Davidian J, Catalan J, Ramalhino DG, Ganem G, Auffray JC, Capela R et al (2000). Rapid chromosomal evolution in island mice. Nature 403: 158.
Chesser RK, Baker RJ (1986). On factors affecting the fixation of chromosomal rearrangements and neutral genes: computer simulations. Evolution 40: 625–632.
Chevret P, Dobigny G (2005). Systematics and evolution of the subfamily Gerbillinae (Mammalia, Rodentia, Muridae). Mol Phyl Evol 35: 674–688.
Delneri D, Colson I, Grammenoudi S, Roberts IN, Louis EJ, Oliver SG (2003). Engineering evolution to study speciation in yeasts. Nature 422: 68–72.
Dobigny G (2002). Speciation chromosomique chez les espèces jumelles ouest-africaines du genre Taterillus (Rodentia, Gerbillinae): Implications systématiques et biogéographiques, Hypothèses génomiques. PhD thesis, Muséum National d'Histoire Naturelle, France.
Dobigny G, Aniskin V, Volobouev V (2002a). Explosive chromosome evolution and speciation in the gerbil genus Taterillus (Rodentia, Gerbillinae): a case of two new cryptic species. Cytogenet Genome Res 96: 117–124.
Dobigny G, Baylac M, Denys C (2002b). Geometric morphometrics, neural networks and diagnosis of sibling Taterillus species (Rodentia, Gerbillinae). Biol J Linnean Soc 77: 319–327.
Dobigny G, Ducroz JF, Robinson TJ, Volobouev V (2004a). Cytogenetics and cladistics. Syst Biol 53: 470–484.
Dobigny G, Granjon L, Aniskin V, Ba K, Volobouev V (2003a). A new sibling species of Taterillus (Rodentia, Gerbillinae) from West Africa. Mammal Biol 68: 299–316.
Dobigny G, Nomao A, Gautun JC (2002c). A cytotaxonomic survey of rodents from Niger: implications for systematics, biodiversity and biogeography. Mammalia 66: 495–523.
Dobigny G, Ozouf-Costaz C, Bonillo C, Volobouev V (2003b). Evolution of rDNA clusters and telomeric repeats during explosive genome repatterning in Taterillus (Rodentia, Gerbillinae). Cytogenet Genome Res 103: 94–103.
Dobigny G, Ozouf-Costaz C, Waters PD, Bonillo C, Coutanceau JP, Vitaly V (2004b). LINE-1 amplification accompanies explosive genome repatterning in rodents. Chromosome Res 12: 787–793.
Ducroz JF, Volobouev V, Granjon L (1998). A molecular perspective on the systematics and evolution of the genus Arvicanthis (Rodentia, Muridae): inferences from complete cytochrome b gene sequences. Mol Phyl Evol 10: 104–117.
Elder FFB (1980). Tandem fusion, centric fusion and chromosomal evolution in the cotton rats, genus Sigmodon. Cytogenet Cell Genet 26: 199–210.
Elder FFB, Hsu TC (1988). Tandem fusion in the evolution of mammalian chromosomes. In: Daniel, Alan A, Liss R (eds) The Cytogenetics of Mammalian Autosomal Rearrangement. Sandberg, AA: New York, pp 481–506.
Fagundes V, Scalzi-Martin JM, Sims K, Hozier J, Yonenaga-Yassuda Y (1997). ZOO-FISH of a microdissection DNA library and G-banding patterns reveals the homeology between the Brazilian rodents Akodon cursior and A montensis. Cytogenet Cell Genet 78: 224–228.
Forejt J (1996). Hybrid sterility in the house mouse. Trends Genet 12: 412–417.
Garagna S, Marziliano N, Zuccotti M, Searle JB, Capanna E, Redi CA (2001). Pericentromeric organization at the fusion point of mouse Robertsonian translocation chromosomes. Proc Nat Acad Sci USA 98: 171–175.
Granjon L, Dobigny G (2003). Chromosomally characterised murid rodents from the edges of Lake Chad: evidence for an African biogeographical crossroads, or a centre of endemism? Mammal Rev 33: 77–91.
Granjon L, Houssin C, Lecompte E, Angaya M, César J, Cornette R et al (2004). Community ecology of the terrestrial small mammals of Zakouma National Park, Chad. Acta Theriol 49: 215–234.
Greenberg AJ, Moran JR, Coyne JA, Wu CI (2003). Ecological adaptation during incipient speciation revealed by precise gene replacement. Science 302: 1754–1757.
Hoffman AA, Sgro CM, Weeks AR (2004). Chromosomal inversion polymorphisms and adaptation. Trends Ecol Evol 19: 483–488.
Hubert B (1982). Ecologie des populations de rongeurs sahélo-soudaniens à Bandia, Sénégal. PhD Thesis, Université de Paris-Sud, France.
King M (1993). Species Evolution: The Role of Chromosomal Change. Cambridge University Press: Cambridge.
Lande R (1979). Effective deme sizes during long-term evolution estimated from rates of chromosomal rearrangement. Evolution 33: 234–251.
Lande R (1985). The fixation of chromosomal rearrangements in a subdivided population with local extinction and colonization. Heredity 54: 323–332.
Lee MR, Elder FFB (1980). Yeast stimulation of bone marrow mitosis for cytogenetic investigations. Cytogenet Cell Genet 26: 36–40.
Matthey R, Jotterand M (1972). L'analyse du caryotype permet de reconnaître deux espèces cryptiques confondues sous le nom de Taterillus gracilis Th. (Rongeurs, Gerbillidae). Mammalia 36: 193–209.
Modi WS (1987). Phylogenetic analyses of chromosomal banding patterns among the Nearctic Arvicolidae. Syst Zool 36: 109–136.
Müller S, Hollatz M, Wienberg J (2003). Chromosomal phylogeny and evolution of gibbons (Hylobatidae). Hum Genet 113: 493–501.
Murphy WJ, Fronicke L, O'Brien SJ, Stanyon R (2003). The origin of human chromosome 1 and its homologs in placental mammals. Genome Res 13: 1880–1888.
Nachman MY, Myers P (1989). Exceptional chromosomal mutations in a rodent population are not strongly underdominant. Proc Natl Acad Sci USA 86: 6666–6670.
Nash WG, Menninger JC, Wienberg J, Padilla-Nash HM, O'Brien SJ (2001). The pattern of phylogenomic evolution of the Canidae. Cytogenet Cell Genet 95: 210–224.
Nie W, Rens W, Wang J, Yang F (2001). Conserved chromosome segments in Hylobates hoolock revealed by human and H. leucogenys paint probes. Cytogenet Cell Genet 92: 248–253.
Noor MAF, Grams KL, Bertucci LA, Reiland J (2001). Chromosomal inversions and the reproductive isolation of species. Proc Natl Acad Sci USA 98: 12084–12088.
Nowak RM (1999). Mammals of the World. The Johns Hopkins University Press: Baltimore and London.
Oakenfull EA, Clegg JB (1998). Phylogenetic relationships within the genus Equus and the evolution of alpha and beta globin genes. J Mol Evol 47: 772–783.
Orr HA (2001). The genetics of species differences. Trends Ecol Evol 16: 342–350.
Owen JG, Baker RJ (2001). The Uroderma bilobatum (Chiroptera, Phyllostomidae) cline revisited. J Mammalogy 82: 1102–1113.
Panhuis TM, Butlin R, Zuk M, Tregenza T (2001). Sexual selection and speciation. Trends Ecol Evol 16: 364–371.
Patton JL, Sherwood SW (1983). Chromosome evolution and speciation in rodents. Ann Rev Ecol Syst 14: 139–158.
Petter F (1974). Facteurs de répartition des rongeurs sahariens et périsahariens. In: Symposium Israël-France: recherches écologiques relatives au développement des zones arides (déserts méditerranéens à précipitations hivernales). Organ. Rech. Agron.: Bet-Dagan, Israel, pp 91–97.
Pialek J, Hauffe HC, Searle JB (2005). Chromosomal variation in the house mouse. Biol J Linnean Soc 85: 535–563.
Poulet AR (1981). Pullulation de rongeurs dans le sahel: mécanismes et déterminismes du cycle d'abondance de Taterillus pygargus et d'Arvicanthis niloticus (Rongeurs, Gerbillidés et Muridés) dans le sahel du Sénégal de 1975 à 1977. PhD Thesis, ORSTOM, France.
Qumsiyeh MB, Hamilton MJ, Schlitter DA (1987). Problems in using Robertsonian rearrangements in determining monophyly: example from the genera Tatera and Gerbillurus. Cytogenet Cell Genet 44: 198–208.
Rens W, O'Brien PCM, Yang F, Solanky N, Perelman P, Graphodatsky AS et al (2001). Karyotype relationships between distantly related marsupials from South America and Australia. Cytogenet Genome Res 9: 301–308.
Rieseberg LH (2001). Chromosomal rearrangements and speciation. Trends Ecol Evol 16: 351–358.
Robbins CB (1974). Comments on the taxonomy of the West African Taterillus (Rodentia, Cricetidae) with the description of a new species. Proc Biol Soc Wash 87: 395–404.
Rognon P (1993). Biographie d'un désert: le Sahara. L'Harmattan. Paris, France.
Roos C, Geissmann T (2001). Molecular phylogeny of the major hylobatid divisions. Mol Phyl Evol 19: 486–494.
Seabright M (1971). A rapid banding technique for human chromosome. Lancet 2: 971–972.
Sicard B (1992). Influence de l'aridité sur la biologie des rongeurs soudano-sahéliens. In: L'aridité: une contrainte au développement. Actiques/ORSTOM: Paris, pp 311–333.
Sicard B, Tranier M, Gautun JC (1988). Un rongeur nouveau du Burkina-Faso (ex Haute-Volta): Taterillus petteri, sp. nov. (Rodentia, Gerbillidae). Mammalia 52: 187–198.
Sites JW, Moritz C (1987). Chromosomal evolution and speciation revisited. Syst Zool 36: 153–214.
Stanyon R, Yang F, Cavagna P, O'Brien PCM, Bagga M, Ferguson-Smith MA et al (1999). Reciprocal chromosome painting shows that genomic rearrangement between rat and mouse proceeds then times faster than between human and cats. Cytogenet Cell Genet 84: 150–155.
Sumner AT (1972). A simple technique for demonstrating centromeric heterochromatin. Exp Cell Res 7: 304–306.
Swofford DL (1998). PAUP: Phylogenetic Analysis Using Parcimony. Version 4.0. Sinauer Associates: Sunderland, MA.
Taylor P (2000). Patterns of chromosomal variation in Southern African rodents. J Mammalogy 81: 317–331.
Tranier M (1974). Polymorphisme chromosomique multiple chez des Taterillus du Niger (Rongeurs, Gerbillidés). C R Acad Sci Paris 278: 3347–3350.
Trifonov V, Yang F, Ferguson-Smith MA, Robinson TJ (2003). Cross-species chromosome painting in the Perissodactyla: delimitation of homologous regions in Burchell's zebra (Equus burchellii) and the white (Ceratotherium simum) and black rhinoceros (Diceros bicornis). Cytogenet Chromosome Res 103: 104–110.
Volobouev V, Aniskin VM, Lecompte E, Ducroz JF (2002). Patterns of karyotype evolution in complexes of sibling species within three genera of African murid rodents inferred from the comparison of cytogenetic and molecular data. Cytogenet Genome Res 96: 261–275.
Volobouev V, Granjon L (1996). A finding of the XX/XY1Y2 sex-chromosome system in Taterillus arenarius (Gerbillinae, Rodentia) and its phylogenetic implications. Cytogenet Cell Genet 75: 45–48.
Wang W, Lan H (2000). Rapid and parallel chromosomal number reductions in Muntjac deer inferred from mitochrondrial DNA phylogeny. Mol Biol Evol 17: 1326–1333.
Yang F, Fu B, O'Brien PCM, Robinson TJ, Ryder OA, Ferguson-Smith MA (2003). Karyotypic relationships of horses and zebras: results of cross-species chromosome painting. Cytogenet Genome Res 102: 235–243.
Yang F, O'Brien PCM, Wienberg J, Neitzel H, Lin CC, Ferguson-Smith MA (1997). Chromosomal evolution of the Chinese muntjac (Muntiacus reevesi). Chromosoma 106: 37–43.
Zhang J, Wang X, Podlaha O (2004). Testing the chromosomal speciation hypothesis for humans and chimpanzees. Genome Res 14: 845–851.
Acknowledgements
We are grateful to J Britton-Davidian, TJ Robinson and F Veyrunes for helpful comments and suggestions on the manuscript. S Ag Atteynine, M Angaya, K Bâ, C Denys, A Doukary, J-C Gautun, C Houssin, E Lecompte, A Nomao, A Oumarou, B Sicard, I Sidibé and B Sidiki are gratefully acknowledged for their help in the field. The data from Ibadan, Nigeria, were kindly provided by M Tranier.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Dobigny, G., Aniskin, V., Granjon, L. et al. Recent radiation in West African Taterillus (Rodentia, Gerbillinae): the concerted role of chromosome and climatic changes. Heredity 95, 358–368 (2005). https://doi.org/10.1038/sj.hdy.6800730
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800730
Keywords
This article is cited by
-
Comprehensive cytogenetic analysis of the most chromosomally variable mammalian genus from South America: Ctenomys (Rodentia: Caviomorpha: Ctenomyidae)
Mammalian Biology (2022)
-
The saprotrophic Pleurotus ostreatus species complex: late Eocene origin in East Asia, multiple dispersal, and complex speciation
IMA Fungus (2020)
-
Phylogeography and evolutionary history of the Crocidura olivieri complex (Mammalia, Soricomorpha): from a forest origin to broad ecological expansion across Africa
BMC Evolutionary Biology (2015)
-
Chromosomal evolution in Rattini (Muridae, Rodentia)
Chromosome Research (2011)
-
Robertsonian fusions, pericentromeric repeat organization and evolution: a case study within a highly polymorphic rodent species, Gerbillus nigeriae
Chromosome Research (2010)