Featured
-
-
Letter |
 New cofactor supports α,β-unsaturated acid decarboxylation via 1,3-dipolar cycloaddition
New cofactor supports α,β-unsaturated acid decarboxylation via 1,3-dipolar cycloaddition
The Fdc1 protein from Aspergillus niger (which is homologous to the UbiD enzyme) uses a new prenylated flavin cofactor to achieve 1,3-dipolar cycloaddition chemistry and catalyse the reversible decarboxylation of aromatic carboxylic acids.
- Karl A. P. Payne
- , Mark D. White
- & David Leys
-
Letter |
 UbiX is a flavin prenyltransferase required for bacterial ubiquinone biosynthesis
UbiX is a flavin prenyltransferase required for bacterial ubiquinone biosynthesis
Ubiquinone is an essential component of electron transfer chains found both in bacteria and in mitochondria; the bacterial enzyme UbiX involved in ubiquinone biosynthesis is a flavin prenyltransferase, and the flavin-derived cofactor synthesized by UbiX is used by the UbiD decarboxylase in the ubiquinone biosynthetic pathway.
- Mark D. White
- , Karl A. P. Payne
- & David Leys
-
Letter |
 Structures of human phosphofructokinase-1 and atomic basis of cancer-associated mutations
Structures of human phosphofructokinase-1 and atomic basis of cancer-associated mutations
The first structures of the mammalian phosphofructokinase-1 tetramer are reported, for the human platelet isoform, in complex with ATP–Mg2+ and ADP.
- Bradley A. Webb
- , Farhad Forouhar
- & Diane L. Barber
-
Article |
 Integrase-mediated spacer acquisition during CRISPR–Cas adaptive immunity
Integrase-mediated spacer acquisition during CRISPR–Cas adaptive immunity
The bacterial CRISPR/Cas system acquires short phage sequences known as spacers that integrate between CRISPR repeats and constitute a record of phage infection; this study shows that the Cas1–Cas2 complex is the minimal machinery required for spacer acquisition and the complex integrates oligonucleotide DNA substrates into acceptor DNA in a manner similar to retroviral integrases and DNA transposases with Cas 1 as the catalytic subunit and Cas2 acting to increase integration activity.
- James K. Nuñez
- , Amy S. Y. Lee
- & Jennifer A. Doudna
-
Letter |
 X-domain of peptide synthetases recruits oxygenases crucial for glycopeptide biosynthesis
X-domain of peptide synthetases recruits oxygenases crucial for glycopeptide biosynthesis
Glycopeptide antibiotics are biosynthesized by non-ribosomal peptide synthetases, which contain a previously uncharacterized ‘X-domain’ now shown to recruit three cytochrome P450 oxygenases that are necessary for the antibiotics to achieve their final, active conformation.
- Kristina Haslinger
- , Madeleine Peschke
- & Max J. Cryle
-
Letter |
 β-Lactam formation by a non-ribosomal peptide synthetase during antibiotic biosynthesis
β-Lactam formation by a non-ribosomal peptide synthetase during antibiotic biosynthesis
The monocyclic β-lactam rings of the nocardicin family of antibiotics are biosynthesized by an unprecedented activity of non-ribosomal peptide synthetases, a mechanism that is distinct from the pathways to the other classes of β-lactam antibiotics.
- Nicole M. Gaudelli
- , Darcie H. Long
- & Craig A. Townsend
-
Letter |
 Structure and mechanism of the tRNA-dependent lantibiotic dehydratase NisB
Structure and mechanism of the tRNA-dependent lantibiotic dehydratase NisB
Structural and biochemical studies show that the biosynthesis of the food preservative nisin involves the tRNA-dependent glutamylation of serine and threonine.
- Manuel A. Ortega
- , Yue Hao
- & Satish K. Nair
-
Letter |
 Reductive dehalogenase structure suggests a mechanism for B12-dependent dehalogenation
Reductive dehalogenase structure suggests a mechanism for B12-dependent dehalogenation
X-ray crystallography and EPR spectroscopy are used to characterize a soluble, oxygen-tolerant reductive dehalogenase from Nitratireductor pacificus pht-3B; the data suggest that the cobalt in the cobalamin cofactor ligates the halogen atom of the substrate, directly abstracting the halogen atom via an oxidative addition.
- Karl A. P. Payne
- , Carolina P. Quezada
- & David Leys
-
Letter |
 Transcript-RNA-templated DNA recombination and repair
Transcript-RNA-templated DNA recombination and repair
Endogenous RNA transcripts are shown to mediate recombination with yeast chromosomal DNA; as the level of RNAs in the nucleus is quite high, these results may open up new understanding of the plasticity of repair and genome instability mechanisms.
- Havva Keskin
- , Ying Shen
- & Francesca Storici
-
Letter |
 Histone H4 tail mediates allosteric regulation of nucleosome remodelling by linker DNA
Histone H4 tail mediates allosteric regulation of nucleosome remodelling by linker DNA
A nucleosome-spacing mechanism for human ATP-dependent chromatin assembly and remodelling factor (ACF).
- William L. Hwang
- , Sebastian Deindl
- & Xiaowei Zhuang
-
Letter |
 Coupled GTPase and remodelling ATPase activities form a checkpoint for ribosome export
Coupled GTPase and remodelling ATPase activities form a checkpoint for ribosome export
Two proteins are identified in yeast that regulate the timing of pre-ribosome export from the nucleus; Nug2 binds pre-60S particles until they are ready for export, at which time Nug2 is replaced by the export adaptor Nmd3, enabling the export machinery to recognise the pre-ribosome that is ready to be transferred to the cytoplasm.
- Yoshitaka Matsuo
- , Sander Granneman
- & Ed Hurt
-
Article |
 RNA catalyses nuclear pre-mRNA splicing
RNA catalyses nuclear pre-mRNA splicing
The spliceosome is shown to catalyse splicing through the RNA and not the protein components of the spliceosome; pre-messenger RNA splicing requires U6 snRNA acting by a mechanism similar to that used by group II self-splicing introns.
- Sebastian M. Fica
- , Nicole Tuttle
- & Joseph A. Piccirilli
-
Letter |
 Flavin-mediated dual oxidation controls an enzymatic Favorskii-type rearrangement
Flavin-mediated dual oxidation controls an enzymatic Favorskii-type rearrangement
Structural and functional studies reveal how the bacterial flavoenzyme EncM catalyses the oxygenation–dehydrogenation dual oxidation of a highly reactive substrate, and show that EncM maintains a stable flavin oxygenating species that promotes substrate oxidation and triggers a rarely seen Favorskii-type rearrangement.
- Robin Teufel
- , Akimasa Miyanaga
- & Bradley S. Moore
-
Letter |
 Precision is essential for efficient catalysis in an evolved Kemp eliminase
Precision is essential for efficient catalysis in an evolved Kemp eliminase
A computationally designed enzyme that was evolved to accelerate a chemical reaction 6 × 108-fold approaches the exceptional efficiency of highly optimized natural enzymes and provides valuable lessons for the creation of more sophisticated artificial catalysts.
- Rebecca Blomberg
- , Hajo Kries
- & Donald Hilvert
-
Letter |
 Hidden specificity in an apparently nonspecific RNA-binding protein
Hidden specificity in an apparently nonspecific RNA-binding protein
A novel high-throughput sequencing kinetics approach is used to measure functional binding of the apparently nonspecific RNA-binding protein C5 to all possible sequence variants in its substrate binding site; C5 binds different substrate variants with affinities varying widely, and with a similar affinity distribution to that of highly specific nucleic-acid-binding proteins, but it does not bind its physiological RNA targets with the highest affinity.
- Ulf-Peter Guenther
- , Lindsay E. Yandek
- & Eckhard Jankowsky
-
Letter |
 Vinylogous chain branching catalysed by a dedicated polyketide synthase module
Vinylogous chain branching catalysed by a dedicated polyketide synthase module
This study shows the structural and biochemical characterization of a new type of polyketide synthase module that catalyses the vinylogous addition of a malonyl unit to an unsaturated thioester, generating a branch in the growing polyketide chain; this characterization provides a mechanism by which the structural diversity of polyketide natural products can be increased.
- Tom Bretschneider
- , Joel B. Heim
- & Christian Hertweck
-
Letter |
 Pyrimidine homeostasis is accomplished by directed overflow metabolism
Pyrimidine homeostasis is accomplished by directed overflow metabolism
Here, the authors identify a previously unknown regulatory strategy used by Escherichia coli to control end-product levels of the pyrimidine biosynthetic pathway: this involves feedback regulation of the near-terminal pathway enzyme UMP kinase, with accumulation of UMP prevented by its degradation to uridine through UmpH, a phosphatase with a previously unknown function.
- Marshall Louis Reaves
- , Brian D. Young
- & Joshua D. Rabinowitz
-
Article |
 The linear ubiquitin-specific deubiquitinase gumby regulates angiogenesis
The linear ubiquitin-specific deubiquitinase gumby regulates angiogenesis
This study identifies a deubiquitinase (DUB) that specifically recognises and cleaves linear ubiquitin chains, implicating linear (de)ubiquitination in Wnt signalling and angiogenesis; mutations in gumby cause defects in angiogenesis in mice, and structural and biochemical analysis shows that gumby encodes a linear-ubiquitin-specific DUB.
- Elena Rivkin
- , Stephanie M. Almeida
- & Sabine P. Cordes
-
Letter |
 The catalytic mechanism for aerobic formation of methane by bacteria
The catalytic mechanism for aerobic formation of methane by bacteria
A mechanism is proposed for the formation of methane by bacteria, through the cleavage of a highly unreactive carbon–phosphorus bond in methyl phosphonate by PhnJ in the bacterial C–P lyase complex.
- Siddhesh S. Kamat
- , Howard J. Williams
- & Frank M. Raushel
-
Letter |
 Mechanistic studies of an unprecedented enzyme-catalysed 1,2-phosphono-migration reaction
Mechanistic studies of an unprecedented enzyme-catalysed 1,2-phosphono-migration reaction
The non-haem-iron-dependent enzyme HppE catalyses the final step in the biosynthesis of fosfomycin, a broad-spectrum, clinically useful antibiotic; here it is shown that HppE can also catalyse a 1,2-phosphono-migration reaction, previously undocumented for any enzyme.
- Wei-chen Chang
- , Mishtu Dey
- & Hung-wen Liu
-
Article |
 Crystal structure of the entire respiratory complex I
Crystal structure of the entire respiratory complex I
The atomic-resolution structure of the entire respiratory complex I is reported, with the resolution high enough to map out the locations and orientations of nearly all amino-acid side chains—some of which link to human neurodegenerative diseases—and reveals which amino-acid interactions take place at the hydrophilic domain–membrane domain interface.
- Rozbeh Baradaran
- , John M. Berrisford
- & Leonid A. Sazanov
-
Letter |
 Control of substrate access to the active site in methane monooxygenase
Control of substrate access to the active site in methane monooxygenase
The crystal structure of the complex between the hydroxylase and regulatory component of soluble methane monooxygenase is presented, revealing how the latter component controls substrate access to the hydroxylase active site.
- Seung Jae Lee
- , Michael S. McCormick
- & Uhn-Soo Cho
-
Article |
 Crystal structure of an RNA-bound 11-subunit eukaryotic exosome complex
Crystal structure of an RNA-bound 11-subunit eukaryotic exosome complex
The crystal structure of a complete yeast exosome (Exo-10) bound to a region of the Rrp6 nuclease and an RNA substrate is determined, demonstrating that the exosome binds and degrades RNA molecules with a channelling mechanism that is largely conserved in all kingdoms of life and is similar to the mechanism used by the proteasome to degrade polypeptides.
- Debora Lika Makino
- , Marc Baumgärtner
- & Elena Conti
-
Letter |
 Activated GTPase movement on an RNA scaffold drives co-translational protein targeting
Activated GTPase movement on an RNA scaffold drives co-translational protein targeting
Single-molecule fluorescence microscopy techniques are used to elucidate features of the highly conserved protein-targeting machinery known as the signal recognition particle (SRP); the long SRP RNA is shown to be crucial for correct timing and precision of cargo handover to the protein-translocation machinery, a finding that could help to explain how other ribonucleosome complexes function during complex cellular processes.
- Kuang Shen
- , Sinan Arslan
- & Shu-ou Shan
-
Article |
 The NAD-dependent deacetylase SIRT2 is required for programmed necrosis
The NAD-dependent deacetylase SIRT2 is required for programmed necrosis
Here it is shown that the NAD-dependent deacetylase SIRT2 is an essential component of necrosis, and that mouse hearts that do not contain SIRT2 or that are treated with a pharmacological inhibitor of SIRT2 are largely protected from ischaemic injury.
- Nisha Narayan
- , In Hye Lee
- & Toren Finkel
-
Article |
 Bypass of a protein barrier by a replicative DNA helicase
Bypass of a protein barrier by a replicative DNA helicase
Single-molecule and ensemble assays are used to show that large T antigen, the replicative DNA helicase of the simian virus 40 (SV40), unwinds DNA as a single hexamer by steric exclusion and is able to bypass covalent DNA–protein crosslinks.
- Hasan Yardimci
- , Xindan Wang
- & Johannes C. Walter
-
Letter |
 Structure and function of the initially transcribing RNA polymerase II–TFIIB complex
Structure and function of the initially transcribing RNA polymerase II–TFIIB complex
Crystal structures of the Pol II–TFIIB complex in free form and bound by the DNA template and a short RNA product are reported; the latter complex represents an initially transcribing complex, a critical transient state in the pathway from transcription initiation to elongation.
- Sarah Sainsbury
- , Jürgen Niesser
- & Patrick Cramer
-
Letter |
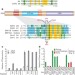 Regulation of ISWI involves inhibitory modules antagonized by nucleosomal epitopes
Regulation of ISWI involves inhibitory modules antagonized by nucleosomal epitopes
Two separate regulatory regions on the Drosophila chromatin remodeller ISWI are defined, AutoN and NegC, which negatively regulate ATP hydrolysis and the coupling of ATP hydrolysis to productive DNA translocation, respectively; epitopes on nucleosomes activate ISWI by inhibiting these negative regulatory domains, ensuring that remodelling occurs only in the appropriate chromatin context.
- Cedric R. Clapier
- & Bradley R. Cairns
-
Letter |
 Structural insight into the type-II mitochondrial NADH dehydrogenases
Structural insight into the type-II mitochondrial NADH dehydrogenases
Analysis of the respective crystal structures of the yeast single-component type-II NADH dehydrogenase Ndi1 in its substrate-free form and when bound to NADH, ubiquinone and NADH–ubiquinone shows that Ndi1 homodimerization through its carboxy-terminal domain is critical for its catalytic activity and membrane targeting.
- Yue Feng
- , Wenfei Li
- & Maojun Yang
-
Letter |
 A selective jumonji H3K27 demethylase inhibitor modulates the proinflammatory macrophage response
A selective jumonji H3K27 demethylase inhibitor modulates the proinflammatory macrophage response
A structure-guided small-molecule and chemoproteomics approach uncovers a catalytic site inhibitor selective for the jumonji subfamily of H3K27me3 demethylases; the inhibitor decreases lipopolysaccharide-induced proinflammatory cytokine production by human primary macrophages.
- Laurens Kruidenier
- , Chun-wa Chung
- & David M. Wilson
-
Article |
 Structure of a RING E3 ligase and ubiquitin-loaded E2 primed for catalysis
Structure of a RING E3 ligase and ubiquitin-loaded E2 primed for catalysis
This study presents the crystal structure of a RING-type E3 ligase bound to ubiquitin-loaded E2; the structure reveals how ubiquitin binding to E2 leads to changes in the catalytic site, priming it for catalysis by the E3 enzyme.
- Anna Plechanovová
- , Ellis G. Jaffray
- & Ronald T. Hay
-
Letter |
 Bacterial virulence proteins as tools to rewire kinase pathways in yeast and immune cells
Bacterial virulence proteins as tools to rewire kinase pathways in yeast and immune cells
Virulence factors from two bacteria are used to reprogram intracellular signalling in yeast and immune T cells, illustrating how pathogens can provide a toolkit to engineer cells for biotechnological or therapeutic applications.
- Ping Wei
- , Wilson W. Wong
- & Wendell A. Lim
-
Letter |
 The dynamic disulphide relay of quiescin sulphydryl oxidase
The dynamic disulphide relay of quiescin sulphydryl oxidase
The X-ray crystal structures of trypanosome and mammalian quiescin sulphydryl oxidase are determined; these structures and follow-up biochemical studies show that large conformational changes occur as the enzyme relays disulphide bonds through its redox-active sites.
- Assaf Alon
- , Iris Grossman
- & Deborah Fass
-
Review Article |
 Engineering the third wave of biocatalysis
Engineering the third wave of biocatalysis
Over the past ten years, protein engineering has established biocatalysis as a practical and environmentally friendly alternative to traditional forms of catalysis both in the laboratory and in industry.
- U. T. Bornscheuer
- , G. W. Huisman
- & K. Robins
-
Letter |
 Functional dissection of lysine deacetylases reveals that HDAC1 and p300 regulate AMPK
Functional dissection of lysine deacetylases reveals that HDAC1 and p300 regulate AMPK
Genetic interaction profiles of human lysine deacetylases are generated by RNA interference knockdown to reveal the involvement of deacetylases in many critical biological processes, including metabolism, the cell cycle and development.
- Yu-yi Lin
- , Samara Kiihl
- & Jef D. Boeke
-
Article |
 Structure and function of the AAA+ protein CbbX, a red-type Rubisco activase
Structure and function of the AAA+ protein CbbX, a red-type Rubisco activase
- Oliver Mueller-Cajar
- , Mathias Stotz
- & Manajit Hayer-Hartl
-
News & Views |
 More than a bystander
More than a bystander
The tendency of hydrophobic surfaces to aggregate in water is often invoked to explain how biomolecules recognize and bind to each other. Water seems to have a much more active role in these processes than had been thought.
- Philip Ball
-
Letter |
 Structural basis for the bifunctionality of fructose-1,6-bisphosphate aldolase/phosphatase
Structural basis for the bifunctionality of fructose-1,6-bisphosphate aldolase/phosphatase
- Shinya Fushinobu
- , Hiroshi Nishimasu
- & Takayoshi Wakagi
-
Letter |
 The structure and catalytic mechanism of a poly(ADP-ribose) glycohydrolase
The structure and catalytic mechanism of a poly(ADP-ribose) glycohydrolase
- Dea Slade
- , Mark S. Dunstan
- & Ivan Ahel
-
Letter |
 Different substrate-dependent transition states in the active site of the ribosome
Different substrate-dependent transition states in the active site of the ribosome
- Stephan Kuhlenkoetter
- , Wolfgang Wintermeyer
- & Marina V. Rodnina
-
Letter |
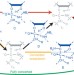 A two-step chemical mechanism for ribosome-catalysed peptide bond formation
A two-step chemical mechanism for ribosome-catalysed peptide bond formation
- David A. Hiller
- , Vipender Singh
- & Scott A. Strobel
-
Letter |
 UBCH7 reactivity profile reveals parkin and HHARI to be RING/HECT hybrids
UBCH7 reactivity profile reveals parkin and HHARI to be RING/HECT hybrids
- Dawn M. Wenzel
- , Alexei Lissounov
- & Rachel E. Klevit
-
Letter |
 Catalytic activity of the caspase-8–FLIPL complex inhibits RIPK3-dependent necrosis
Catalytic activity of the caspase-8–FLIPL complex inhibits RIPK3-dependent necrosis
Caspase-8 mediates apoptosis induced by death receptors. At the same time, this protease is able to prevent RIP-dependent necrosis. Without caspase-8 mice die during their embryonic development. Two papers now show that lethality is not caused by the absence of apoptosis, but by RIP3-dependent necrosis that is unleashed without caspase-8. Mice that lack both caspase-8 and RIP3 develop into viable, immunocompetent, fertile adult mice, but suffer from a progressive lymphoaccumulative disease similar to mice that lack the death receptor CD95. This paper further shows that caspase-8 forms a proteolytically active complex with FLIPL, and that this complex is required for protection against RIP3-dependent necrosis.
- Andrew Oberst
- , Christopher P. Dillon
- & Douglas R. Green
-
Letter |
 Dynamics and mechanism of repair of ultraviolet-induced (6–4) photoproduct by photolyase
Dynamics and mechanism of repair of ultraviolet-induced (6–4) photoproduct by photolyase
The repair enzyme (6–4) photolyase uses light energy to cleave the ultraviolet-induced bond between pyrimidine dimers. These authors use ultrafast spectroscopy to examine the detailed electron and proton movements during the catalytic photocycle. Histidine 364 is identified as the crucial residue involved in the rate-limiting step.
- Jiang Li
- , Zheyun Liu
- & Dongping Zhong
-
News |
 Retraction recommended for enzyme-chip paper
Retraction recommended for enzyme-chip paper
Reactome array study should not have been published, says ethics committee.
- Alison Abbott
-
Article |
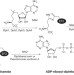 Diphthamide biosynthesis requires an organic radical generated by an iron–sulphur enzyme
Diphthamide biosynthesis requires an organic radical generated by an iron–sulphur enzyme
Translation elongation factor 2 (EF2) from archaea and eukaryotes contains a unique, post-translationally modified histidine residue called diphthamide, which is the target of diphtheria toxin. The biosynthesis of diphthamide involves three steps; here it is shown that the first step in the archaeon Pyrococcus horikoshii requires an unusual iron–sulphur-cluster enzyme, Dph2. It catalyses unprecedented chemistry.
- Yang Zhang
- , Xuling Zhu
- & Hening Lin
-
News & Views |
 Getting the metal right
Getting the metal right
Controversy has raged over the identity of the metal cofactor of membrane-bound methane monooxygenase, a methane-oxidizing enzyme. A study suggests that the answer is a cluster of two copper ions.
- J. Martin Bollinger Jr
-
Letter |
 Oxidation of methane by a biological dicopper centre
Oxidation of methane by a biological dicopper centre
Particulate methane monooxygenase (pMMO) is an integral membrane protein, found in methanotropic bacteria, that can selectively oxidize methane to produce methanol. This metalloenzyme contains three subunits, and the metal composition and exact location of its active site has been the subject of much speculation. Here it is found that the enzyme's activity is dependent on copper, and that the active site is located in the soluble domains of the pmoB subunit.
- Ramakrishnan Balasubramanian
- , Stephen M. Smith
- & Amy C. Rosenzweig
-
Letter |
 Transfer of carbohydrate-active enzymes from marine bacteria to Japanese gut microbiota
Transfer of carbohydrate-active enzymes from marine bacteria to Japanese gut microbiota
One of the roles of the human gut microbiota is to break down nutrients using bacterial enzymes that are lacking from the human genome. It is now shown that the gut microbiota of Japanese, but not American, individuals contains porphyranases, enzymes that digest sulphated polysaccharides which are present in the marine environment only. These findings indicate that diet can select for gene content of the human microbiota.
- Jan-Hendrik Hehemann
- , Gaëlle Correc
- & Gurvan Michel

 Mechanism of phospho-ubiquitin-induced PARKIN activation
Mechanism of phospho-ubiquitin-induced PARKIN activation