Abstract
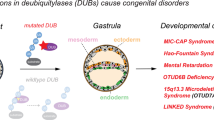
A complex interaction of signalling events, including the Wnt pathway, regulates sprouting of blood vessels from pre-existing vasculature during angiogenesis. Here we show that two distinct mutations in the (uro)chordate-specific gumby (also called Fam105b) gene cause an embryonic angiogenic phenotype in gumby mice. Gumby interacts with disheveled 2 (DVL2), is expressed in canonical Wnt-responsive endothelial cells and encodes an ovarian tumour domain class of deubiquitinase that specifically cleaves linear ubiquitin linkages. A crystal structure of gumby in complex with linear diubiquitin reveals how the identified mutations adversely affect substrate binding and catalytic function in line with the severity of their angiogenic phenotypes. Gumby interacts with HOIP (also called RNF31), a key component of the linear ubiquitin assembly complex, and decreases linear ubiquitination and activation of NF-κB-dependent transcription. This work provides support for the biological importance of linear (de)ubiquitination in angiogenesis, craniofacial and neural development and in modulating Wnt signalling.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Risau, W. Mechanisms of angiogenesis. Nature 386, 671–674 (1997)
Logan, C. Y. & Nusse, R. The Wnt signaling pathway in development and disease. Annu. Rev. Cell Dev. Biol. 20, 781–810 (2004)
Liebner, S. & Plate, K. H. Differentiation of the brain vasculature: the answer came blowing by the Wnt. J. Angio. Res. 2, 1 (2010)
Zerlin, M., Julius, M. A. & Kitajewski, J. Wnt/Frizzled signaling in angiogenesis. Angiogenesis 11, 63–69 (2008)
Daneman, R. et al. Wnt/β-catenin signaling is required for CNS, but not non-CNS, angiogenesis. Proc. Natl Acad. Sci. USA 106, 641–646 (2009)
Stenman, J. M. et al. Canonical Wnt signaling regulates organ-specific assembly and differentiation of CNS vasculature. Science 322, 1247–1250 (2008)
Cirone, P. et al. A role for planar cell polarity signaling in angiogenesis. Angiogenesis 11, 347–360 (2008)
Wharton, K. A., Jr Runnin’ with the Dvl: proteins that associate with Dsh/Dvl and their significance to Wnt signal transduction. Dev. Biol. 253, 1–17 (2003)
Kirisako, T. et al. A ubiquitin ligase complex assembles linear polyubiquitin chains. EMBO J. 25, 4877–4887 (2006)
Behrends, C. & Harper, J. W. Constructing and decoding unconventional ubiquitin chains. Nature Struct. Mol. Biol. 18, 520–528 (2011)
Walczak, H., Iwai, K. & Dikic, I. Generation and physiological roles of linear ubiquitin chains. BMC Biol. 10, 23 (2012)
Iwai, K. Linear polyubiquitin chains: a new modifier involved in NFκB activation and chronic inflammation, including dermatitis. Cell Cycle 10, 3095–3104 (2011)
Gerlach, B. et al. Linear ubiquitination prevents inflammation and regulates immune signalling. Nature 471, 591–596 (2011)
Ikeda, F. et al. SHARPIN forms a linear ubiquitin ligase complex regulating NF-κB activity and apoptosis. Nature 471, 637–641 (2011)
Tokunaga, F. et al. SHARPIN is a component of the NF-κB-activating linear ubiquitin chain assembly complex. Nature 471, 633–636 (2011)
Niu, J., Shi, Y., Iwai, K. & Wu, Z. H. LUBAC regulates NF-κB activation upon genotoxic stress by promoting linear ubiquitination of NEMO. EMBO J. 30, 3741–3753 (2011)
Nijman, S. M. et al. A genomic and functional inventory of deubiquitinating enzymes. Cell 123, 773–786 (2005)
Mar, L., Rivkin, E., Kim, D. Y., Yu, J. Y. & Cordes, S. P. A genetic screen for mutations that affect cranial nerve development in the mouse. J. Neurosci. 25, 11787–11795 (2005)
Hogan, K. A., Ambler, C. A., Chapman, D. L. & Bautch, V. L. The neural tube patterns vessels developmentally using the VEGF signaling pathway. Development 131, 1503–1513 (2004)
Gondo, Y. Trends in large-scale mouse mutagenesis: from genetics to functional genomics. Nature Rev. Genet. 9, 803–810 (2008)
Roca, C. & Adams, R. H. Regulation of vascular morphogenesis by Notch signaling. Genes Dev. 21, 2511–2524 (2007)
Edelmann, M. J. et al. Structural basis and specificity of human otubain 1-mediated deubiquitination. Biochem. J. 418, 379–390 (2009)
Juang, Y. C. et al. OTUB1 co-opts Lys48-linked ubiquitin recognition to suppress E2 enzyme function. Mol. Cell 45, 384–397 (2012)
Wiener, R., Zhang, X., Wang, T. & Wolberger, C. The mechanism of OTUB1-mediated inhibition of ubiquitination. Nature 483, 618–622 (2012)
Wang, T. et al. Evidence for bidentate substrate binding as the basis for the K48 linkage specificity of otubain 1. J. Mol. Biol. 386, 1011–1023 (2009)
Matsumoto, M. L. et al. Engineering and structural characterization of a linear polyubiquitin-specific antibody. J. Mol. Biol. 418, 134–144 (2012)
Rual, J. F. et al. Towards a proteome-scale map of the human protein–protein interaction network. Nature 437, 1173–1178 (2005)
Maretto, S. et al. Mapping Wnt/β-catenin signaling during mouse development and in colorectal tumors. Proc. Natl Acad. Sci. USA 100, 3299–3304 (2003)
HogenEsch, H. et al. A spontaneous mutation characterized by chronic proliferative dermatitis in C57BL mice. Am. J. Pathol. 143, 972–982 (1993)
Seymour, R. E. et al. Spontaneous mutations in the mouse Sharpin gene result in multiorgan inflammation, immune system dysregulation and dermatitis. Genes Immun. 8, 416–421 (2007)
Rahighi, S. et al. Specific recognition of linear ubiquitin chains by NEMO is important for NF-κB activation. Cell 136, 1098–1109 (2009)
Mainardi, P. C. et al. The natural history of Cri du Chat Syndrome. A report from the Italian Register. Eur. J. Med. Genet. 49, 363–383 (2006)
Mainardi, P. C. et al. Clinical and molecular characterisation of 80 patients with 5p deletion: genotype-phenotype correlation. J. Med. Genet. 38, 151–158 (2001)
Zhang, X. et al. High-resolution mapping of genotype-phenotype relationships in cri du chat syndrome using array comparative genomic hybridization. Am. J. Hum. Genet. 76, 312–326 (2005)
Ye, S., Dhillon, S., Ke, X., Collins, A. R. & Day, I. N. An efficient procedure for genotyping single nucleotide polymorphisms. Nucleic Acids Res. 29, e88 (2001)
Adams, D. J. et al. A genome-wide, end-sequenced 129Sv BAC library resource for targeting vector construction. Genomics 86, 753–758 (2005)
DasGupta, R. & Fuchs, E. Multiple roles for activated LEF/TCF transcription complexes during hair follicle development and differentiation. Development 126, 4557–4568 (1999)
Ambulos, N. P., Jr, Duvall, E. J. & Lovett, P. S. Method for blot-hybridization analysis of mRNA molecules from Bacillus subtilis. Gene 51, 281–286 (1987)
Kim, F. A. et al. The vHNF1 homeodomain protein establishes early rhombomere identity by direct regulation of Kreisler expression. Mech. Dev. 122, 1300–1309 (2005)
Vecchi, A. et al. Monoclonal antibodies specific for endothelial cells of mouse blood vessels. Their application in the identification of adult and embryonic endothelium. Eur. J. Cell Biol. 63, 247–254 (1994)
Abramoff, M. D., Magalhaes, P. J. & Ram, S. J. Image Processing with ImageJ. Biophotonics Intl 11, 36–42 (2004)
Torban, E., Wang, H. J., Groulx, N. & Gros, P. Independent mutations in mouse Vangl2 that cause neural tube defects in looptail mice impair interaction with members of the Dishevelled family. J. Biol. Chem. 279, 52703–52713 (2004)
Veeman, M. T., Axelrod, J. D. & Moon, R. T. A second canon. Functions and mechanisms of β-catenin-independent Wnt signaling. Dev. Cell 5, 367–377 (2003)
Ernst, A. et al. A strategy for modulation of enzymes in the ubiquitin system. Science 339, 590–595 (2013)
Kean, M. J., Couzens, A. L. & Gingras, A. C. Mass spectrometry approaches to study mammalian kinase and phosphatase associated proteins. Methods 57, 400–408 (2012)
Dunham, W. H. et al. A cost–benefit analysis of multidimensional fractionation of affinity purification-mass spectrometry samples. Proteomics 11, 2603–2612 (2011)
Liu, G. et al. ProHits: integrated software for mass spectrometry-based interaction proteomics. Nature Biotechnol. 28, 1015–1017 (2010)
Choi, H. et al. Analyzing protein-protein interactions from affinity purification-mass spectrometry data with SAINT. Curr. Protoc. Bioinform. Ch. 8, Unit 8.15. (2012)
Choi, H. et al. SAINT: probabilistic scoring of affinity purification-mass spectrometry data. Nature Methods 8, 70–73 (2011)
Acknowledgements
We thank T. Saunders and the Michigan University Transgenic Core facility for generating gumby BAC transgenic mice; K. Iwai for HOIL and HOIP constructs; K. Nakajima for NF-κB and AP2 reporter constructs; B. Alman and C. C. Hui for TOPGAL reporter mice; J. Woodgett for TOPFLASH, FOPFLASH and PRL vectors; T. Pawson for WNT3A expressing L-cells; M. Barrios-Ramos for help with fluorometry; J. Culotti and C. C. Hui for critical feedback; and I. Kourinov and staff at NE-CAT for collecting diffraction data set. B.R. holds the Canada research chair in proteomics and molecular medicine. A.-C.G. holds the Canada research chair in Functional Proteomics. F.S. holds the Canada research chair in Structural Principles of Signal Transduction. S.M.A. and T.S. were supported by CIHR predoctoral student fellowships. This work was supported by operating funds from the JSPS KAKENHI (grant numbers 15200032 to Y.G. and 21240043 to Y.G. and R.F.), and Canadian Institutes for Health Research to B.R. (MOP119289), A.-C.G. (MOP123433), F.S. (MOP57795) and S.P.C. (MOP 97966, IHO 94384 and MOP 111199).
Author information
Authors and Affiliations
Contributions
E.R. designed genetic experiments with S.P.C., identified and confirmed the gumby causative mutation, performed expression analyses and characterized the GumW96R angiogenic phenotype; S.M.A. performed protein interaction assays and analysed the gumby–LUBAC connection; D.F.C. and Y.-C.J. performed X-ray crystallographic analyses and DUB assays; T.A.M. analysed angiogenesis and Wnt signalling; T.S. performed DUB chain profiling assay; H.H. and Y.-C.J. performed ITC analyses; Y.G. supervised R.F.; R.F. identified the GumD336E mutant; W.H.D. ran the mass spectrometry; A.-C.G. supervised W.H.D.; G.X. helped with SNP mapping; B.R. supervised T.S. and F.S. supervised D.C., H.H., Y-C.J. and together they designed experiments and wrote the manuscript; S.P.C. conceived and coordinated the project, designed and performed experiments, supervised E.R., S.M.A. and T.A.M., and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1–11 and Supplementary Tables 1–3. (PDF 8693 kb)
Supplementary Data
This file contains tabulated data for the enzymatic Gumby assays. (XLSX 43 kb)
Rights and permissions
About this article
Cite this article
Rivkin, E., Almeida, S., Ceccarelli, D. et al. The linear ubiquitin-specific deubiquitinase gumby regulates angiogenesis. Nature 498, 318–324 (2013). https://doi.org/10.1038/nature12296
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12296
This article is cited by
-
LUBAC promotes angiogenesis and lung tumorigenesis by ubiquitinating and antagonizing autophagic degradation of HIF1α
Oncogenesis (2024)
-
OTULIN Haploinsufficiency-Related Fasciitis and Skin Necrosis Treated by TNF Inhibition
Journal of Clinical Immunology (2024)
-
Mechanisms underlying linear ubiquitination and implications in tumorigenesis and drug discovery
Cell Communication and Signaling (2023)
-
Deubiquitinases in cancer
Nature Reviews Cancer (2023)
-
OTULIN protects the intestinal epithelium from apoptosis during inflammation and infection
Cell Death & Disease (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.