Abstract
Chronic inflammation is associated with increased risk and poor prognosis of heart failure; however, the precise mechanism that provokes sustained inflammation in the failing heart remains elusive. Here we report that depletion of carnitine acetyltransferase (CRAT) promotes cholesterol catabolism through bile acid synthesis pathway in cardiomyocytes. Intracellular accumulation of bile acid or intermediate, 7α-hydroxyl-3-oxo-4-cholestenoic acid, induces mitochondrial DNA stress and triggers cGAS–STING-dependent type I interferon responses. Furthermore, type I interferon responses elicited by CRAT deficiency substantially increase AIM2 expression and AIM2-dependent inflammasome activation. Genetic deletion of cardiomyocyte CRAT in mice of both sexes results in myocardial inflammation and dilated cardiomyopathy, which can be reversed by combined depletion of caspase-1, cGAS or AIM2. Collectively, we identify a mechanism by which cardiac energy metabolism, cholesterol homeostasis and cardiomyocyte-intrinsic innate immune responses are interconnected via a CRAT-mediated bile acid synthesis pathway, which contributes to chronic myocardial inflammation and heart failure progression.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The data that support the findings of this study are available within the paper in Supplementary Information files. Detailed information of key reagents is provided in Supplementary Table 1. The raw sequence data reported in this paper have been deposited in GEO (accession no. GSE200057). This RNA-seq dataset, together with the other single-cell RNA-seq datasets GSE109816 and GSE121893 (ref. 6), is publicly accessible at GEO. Source data are provided with this paper.
References
Hunt, S. A. et al. ACC/AHA guidelines for the evaluation and management of chronic heart failure in the adult: executive summary a report of the American College of Cardiology/American Heart Association Task Force on practice guidelines (committee to revise the 1995 guidelines for the evaluation and management of heart failure): developed in collaboration with the International Society for Heart and Lung Transplantation; endorsed by the Heart Failure Society of America. Circulation 104, 2996–3007 (2001).
McDonagh, T. A. et al. 2021 ESC Guidelines for the diagnosis and treatment of acute and chronic heart failure. Eur. Heart J. 42, 3599–3726 (2021).
Doenst, T., Nguyen, T. D. & Abel, E. D. Cardiac metabolism in heart failure: implications beyond ATP production. Circ. Res. 113, 709–724 (2013).
Muoio, D. M. et al. Muscle-specific deletion of carnitine acetyltransferase compromises glucose tolerance and metabolic flexibility. Cell Metab. 15, 764–777 (2012).
Davidson, M. T. et al. Extreme acetylation of the cardiac mitochondrial proteome does not promote heart failure. Circ. Res. 127, 1094–1108 (2020).
Wang, L. et al. Single-cell reconstruction of the adult human heart during heart failure and recovery reveals the cellular landscape underlying cardiac function. Nat. Cell Biol. 22, 108–119 (2020).
Bersell, K. et al. Moderate and high amounts of tamoxifen in αMHC-MerCreMer mice induce a DNA damage response, leading to heart failure and death. Dis. Model Mech. 6, 1459–1469 (2013).
Davies, M. N. et al. The acetyl group buffering action of carnitine acetyltransferase offsets macronutrient-induced lysine acetylation of mitochondrial proteins. Cell Rep. 14, 243–254 (2016).
Motwani, M., Pesiridis, S. & Fitzgerald, K. A. DNA sensing by the cGAS–STING pathway in health and disease. Nat. Rev. Genet. 20, 657–674 (2019).
Yu, C. H. et al. TDP-43 triggers mitochondrial DNA release via mPTP to activate cGAS/STING in ALS. Cell 183, 636–649 (2020).
West, A. P. et al. Mitochondrial DNA stress primes the antiviral innate immune response. Nature 520, 553–557 (2015).
Kim, J. et al. VDAC oligomers form mitochondrial pores to release mtDNA fragments and promote lupus-like disease. Science 366, 1531–1536 (2019).
Sprenger, H. G. et al. Cellular pyrimidine imbalance triggers mitochondrial DNA-dependent innate immunity. Nat. Metab. 3, 636–650 (2021).
Chapman, J., Ng, Y. S. & Nicholls, T. J. The maintenance of mitochondrial DNA integrity and dynamics by mitochondrial membranes. Life https://doi.org/10.3390/life10090164 (2020).
Fontana, G. A. & Gahlon, H. L. Mechanisms of replication and repair in mitochondrial DNA deletion formation. Nucleic Acids Res. 48, 11244–11258 (2020).
Siperstein, M. D. & Fagan, V. M. Feedback control of mevalonate synthesis by dietary cholesterol. J. Biol. Chem. 241, 602–609 (1966).
Russell, D. W. The enzymes, regulation, and genetics of bile acid synthesis. Annu. Rev. Biochem. 72, 137–174 (2003).
Saeed, A. et al. 7α-hydroxy-3-oxo-4-cholestenoic acid in cerebrospinal fluid reflects the integrity of the blood-brain barrier. J. Lipid Res. 55, 313–318 (2014).
Chiang, J. Y. Regulation of bile acid synthesis: pathways, nuclear receptors, and mechanisms. J. Hepatol. 40, 539–551 (2004).
Botla, R., Spivey, J. R., Aguilar, H., Bronk, S. F. & Gores, G. J. Ursodeoxycholate (UDCA) inhibits the mitochondrial membrane permeability transition induced by glycochenodeoxycholate: a mechanism of UDCA cytoprotection. J. Pharmacol. Exp. Ther. 272, 930–938 (1995).
Kandell, R. L. & Bernstein, C. Bile salt/acid induction of DNA damage in bacterial and mammalian cells: implications for colon cancer. Nutr. Cancer 16, 227–238 (1991).
Guo, G. L. & Chiang, J. Y. L. Is CYP2C70 the key to new mouse models to understand bile acids in humans? J. Lipid Res. 61, 269–271 (2020).
Quinn, C. M., Jessup, W., Wong, J., Kritharides, L. & Brown, A. J. Expression and regulation of sterol 27-hydroxylase (CYP27A1) in human macrophages: a role for RXR and PPARγ ligands. Biochem. J. 385, 823–830 (2005).
Leesnitzer, L. M. et al. Functional consequences of cysteine modification in the ligand binding sites of peroxisome proliferator activated receptors by GW9662. Biochemistry 41, 6640–6650 (2002).
Lugrin, J. & Martinon, F. The AIM2 inflammasome: sensor of pathogens and cellular perturbations. Immunol. Rev. 281, 99–114 (2018).
Brunette, R. L. et al. Extensive evolutionary and functional diversity among mammalian AIM2-like receptors. J. Exp. Med. 209, 1969–1983 (2012).
Schattgen, S. A. & Fitzgerald, K. A. The PYHIN protein family as mediators of host defenses. Immunol. Rev. 243, 109–118 (2011).
Hu, B. et al. The DNA-sensing AIM2 inflammasome controls radiation-induced cell death and tissue injury. Science 354, 765–768 (2016).
Shi, J. et al. Cleavage of GSDMD by inflammatory caspases determines pyroptotic cell death. Nature 526, 660–665 (2015).
Ding, J. et al. Pore-forming activity and structural autoinhibition of the gasdermin family. Nature 535, 111–116 (2016).
Edgeworth, J. D., Spencer, J., Phalipon, A., Griffin, G. E. & Sansonetti, P. J. Cytotd interleukinoxicity an-1β processing following Shigella flexneriinfection of human monocyte-derived dendritic cells.Eur. J. Immunol. 32, 1464–1471 (2002).
Bergsbaken, T., Fink, S. L. & Cookson, B. T. Pyroptosis: host cell death and inflammation. Nat. Rev. Microbiol. 7, 99–109 (2009).
Miao, E. A., Rajan, J. V. & Aderem, A. Caspase-1-induced pyroptotic cell death. Immunol. Rev. 243, 206–214 (2011).
Levine, B., Kalman, J., Mayer, L., Fillit, H. M. & Packer, M. Elevated circulating levels of tumor necrosis factor in severe chronic heart failure. N. Engl. J. Med. 323, 236–241 (1990).
Testa, M. et al. Circulating levels of cytokines and their endogenous modulators in patients with mild to severe congestive heart failure due to coronary artery disease or hypertension. J. Am. Coll. Cardiol. 28, 964–971 (1996).
Torre-Amione, G. et al. Proinflammatory cytokine levels in patients with depressed left ventricular ejection fraction: a report from the Studies of Left Ventricular Dysfunction (SOLVD). J. Am. Coll. Cardiol. 27, 1201–1206 (1996).
Chung, E. S. et al. Randomized, double-blind, placebo-controlled, pilot trial of infliximab, a chimeric monoclonal antibody to tumor necrosis factor-α, in patients with moderate-to-severe heart failure: results of the anti-TNF Therapy Against Congestive Heart Failure (ATTACH) trial. Circulation 107, 3133–3140 (2003).
Mann, D. L. et al. Targeted anticytokine therapy in patients with chronic heart failure: results of the Randomized Etanercept Worldwide Evaluation (RENEWAL). Circulation 109, 1594–1602 (2004).
Everett, B. M. et al. Anti-inflammatory therapy with canakinumab for the prevention of hospitalization for heart failure. Circulation 139, 1289–1299 (2019).
Bujak, M. et al. Interleukin-1 receptor type I signaling critically regulates infarct healing and cardiac remodeling. Am. J. Pathol. 173, 57–67 (2008).
Bageghni, S. A. et al. Fibroblast-specific deletion of interleukin-1 receptor-1 reduces adverse cardiac remodeling following myocardial infarction. JCI Insight https://doi.org/10.1172/jci.insight.125074 (2019).
Zeng, C. et al. NLRP3 inflammasome-mediated pyroptosis contributes to the pathogenesis of non-ischemic dilated cardiomyopathy. Redox Biol. 34, 101523 (2020).
Gulick, T., Chung, M. K., Pieper, S. J., Lange, L. G. & Schreiner, G. F. Interleukin 1 and tumor necrosis factor inhibit cardiac myocyte β-adrenergic responsiveness. Proc. Natl Acad. Sci. USA 86, 6753–6757 (1989).
Yokoyama, T. et al. Cellular basis for the negative inotropic effects of tumor necrosis factor-α in the adult mammalian heart. J. Clin. Invest. 92, 2303–2312 (1993).
Yu, X., Kennedy, R. H. & Liu, S. J. JAK2/STAT3, not ERK1/2, mediates interleukin-6-induced activation of inducible nitric-oxide synthase and decrease in contractility of adult ventricular myocytes. J. Biol. Chem. 278, 16304–16309 (2003).
Murphy, S. P., Kakkar, R., McCarthy, C. P. & Januzzi, J. L. Jr. Inflammation in heart failure: JACC state-of-the-art review. J. Am. Coll. Cardiol. 75, 1324–1340 (2020).
Noutsias, M., Pauschinger, M., Schultheiss, H. & Uwe, K. H. Phenotypic characterization of infiltrates in dilated cardiomyopathy—diagnostic significance of T-lymphocytes and macrophages in inflammatory cardiomyopathy.Med Sci. Monit. 8, CR478–CR487 (2002).
King, K. R. et al. IRF3 and type I interferons fuel a fatal response to myocardial infarction. Nat. Med. 23, 1481–1487 (2017).
Ridker, P. M. Hyperlipidemia as an instigator of inflammation: inaugurating new approaches to vascular prevention. J. Am. Heart Assoc. 1, 3–5 (2012).
Duewell, P. et al. NLRP3 inflammasomes are required for atherogenesis and activated by cholesterol crystals. Nature 464, 1357–1361 (2010).
Seiler, S. E. et al. Obesity and lipid stress inhibit carnitine acetyltransferase activity. J. Lipid Res. 55, 635–644 (2014).
Angelini, A. et al. PHDs/CPT1B/VDAC1 axis regulates long-chain fatty acid oxidation in cardiomyocytes. Cell Rep. 37, 109767 (2021).
Mao, H. et al. Endothelium-specific depletion of LRP1 improves glucose homeostasis through inducing osteocalcin. Nat. Commun. 12, 5296 (2021).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Shi, J. W. & Walker, M. G. Gene set enrichment analysis (GSEA) for interpreting gene expression profiles. Curr. Bioinform. https://doi.org/10.2174/157489307780618231 (2007).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. https://doi.org/10.1186/s13059-017-1382-0 (2018).
Virtanen, P. et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat. Methods 17, 261–272 (2020).
Amara, C. S. et al. Serum metabolic profiling identified a distinct metabolic signature in bladder cancer smokers: a key metabolic enzyme associated with patient survival. Cancer Epidemiol. Biomark. Prev. 28, 770–781 (2019).
Vantaku, V. et al. Large-scale profiling of serum metabolites in African American and European American patients with bladder cancer reveals metabolic pathways associated with patient survival. Cancer 125, 921–932 (2019).
Louch, W. E., Sheehan, K. A. & Wolska, B. M. Methods in cardiomyocyte isolation, culture, and gene transfer. J. Mol. Cell. Cardiol. 51, 288–298 (2011).
Acknowledgements
This work is supported by National Institutes of Health grants R01 HL122736 (to L.X.), R01 HL166549 (to L.X.) and R01 DK123186 (to X.P.).
Author information
Authors and Affiliations
Contributions
L.X. and X.P. conceived and designed the studies. H.M., A.A., L.L., C.P., X.P. and L.X. performed the experiments and interpreted data. G.W. and S.L. performed the RNA-seq analysis. L.X. and X.P. co-wrote the manuscript. L.X. carried out experimental design and interpretation and supervised the project. All authors approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Joseph Hill, A. Phillip West and the other, anonymous, reviewer for their contribution to the peer review of this work. Primary Handling Editor: Ashley Castellanos-Jankiewicz, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Activation of Myh6-MerCreMer with low doses of tamoxifen has no deleterious effects on cardiac function.
Adult Myh6-MerCreMer− or Myh6-MerCreMer + male and female mice were injected with tamoxifen for 4 consecutive days at a dose of 20 mg/kg/day. a, b, Echocardiographic analyses were performed to estimate cardiac contractile function after 2 months. c-f, RT-PCR analysis of the expression of the hypertrophic marker genes and the pro-fibrotic genes in the hearts. No significant changes were observed between these two groups. n = 5 mice per group. Data are shown as the mean ± SEM. Analysis was unpaired two-tailed Student’s t-test (a-f). NS, not significant.
Extended Data Fig. 2 Cardiomyocyte-specific deletion of CRAT increases acetyl-CoA level in the hearts.
a, Western-blots showed that CRAT was specifically depleted in CRAT-mKO male and female hearts but not in lungs or gastrocnemius muscles. b, CRAT activity was dramatically decreased in CRAT-mKO male and female hearts. c, Acetyl-CoA level was significantly increased in CRAT-mKO male and female hearts. n = 4 (a), 9 (b, WT), 7 (b, mKO), 12 (c, WT), 7 (c, mKO) mice per group. Data are shown as the mean ± SEM. Analysis was unpaired two-tailed Student’s t-test (a-c).
Extended Data Fig. 3 Knockdown of CRAT in NRVMs increases the expression of pro-inflammatory cytokines.
NRVMs were transduced with lentivirus expressing Crat shRNA or control shRNA for 5 days. RNAs were extracted from the cells and RT-PCRs were then performed to determine the expression of hypertrophy, pro-fibrotic or pro-inflammatory genes. a, knockdown of CRAT expression was confirmed by RT-PCR. b-f, the expression of hypertrophic and pro-fibrotic genes was not increased by CRAT knockdown. g-i, the expression of Il-1β, Il-6 and Tnf-α was substantially increased in CRAT-deficient NRVMs. n = 6-7 (a-i, Sh-Control), 6 (a-i, Sh-Crat) biologically independent samples per group. Data are shown as the mean ± SEM. Analysis was unpaired two-tailed Student’s t-test (a-i). NS, not significant.
Extended Data Fig. 4 CRAT deficiency induces type I interferon responses in NRVMs.
a-e, NRVMs were transduced with lentivirus expressing Crat shRNA, control shRNA or no virus for 3 or 5 days. RNAs were extracted from the cells and RT-PCRs were then performed to determine the expression of Aim2 (a), Ddx58 (b), Ifih1(c), Ifit3 (d) and Irf7 (e). n = 4 (a-e) biologically independent samples per group. Data are shown as the mean ± SEM. Analysis was two-way ANOVA followed by Bonferroni’s multiple comparison test (a-e).
Extended Data Fig. 5 CRAT deficiency induces type I interferon responses in cardiac fibroblasts.
a-d, Cardiac fibroblasts were transduced with lentivirus expressing Crat shRNA, control shRNA or no virus for 3 or 5 days. RNAs were extracted from the cells and RT-PCRs were then performed to determine the expression of Aim2 (a), Ddx58 (b), Irf7 (c) and Ifit3 (d). n = 4 (a-d, no virus, Sh-Control), 3-4 (a-d, Sh-Crat) biologically independent samples per group. Data are shown as the mean ± SEM. Analysis was two-way ANOVA followed by Bonferroni’s multiple comparison test (a-d).
Extended Data Fig. 6 Depletion of CRAT has no effect on total mitochondrial DNA level in NRVMs and adult CMs.
a, b, NRVMs were transduced with lentivirus expressing control shRNA or Crat shRNA. Cells were harvested and mitochondria and cytosol fractionation was then performed after 5 days. Western blots indicated that there was no contamination between these two fractions (a). RT-PCRs were then performed with mitochondrial fraction to quantify the total mitochondrial DNA level (b). c, Adult CRAT-WT and CRAT-mKO male and female mice were injected with tamoxifen (20 mg/kg/day) for four consecutive days. Three weeks later, CMs were isolated from hearts for mitochondria and cytosol fractionation. RT-PCRs were then performed with mitochondrial fraction to quantify the total mitochondria DNA level. n = 5 biologically independent samples (b), 6 mice (c) per group. Data are shown as the mean ± SEM. Analysis was unpaired two-tailed Student’s t-test (b-c). NS, not significant.
Extended Data Fig. 7 RNA-sequencing analysis for NRVMs transduced with lentivirus expressing control shRNA or Crat shRNA.
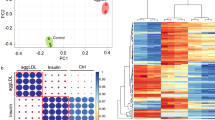
a, b, Heatmap and volcano plot of the significantly differential expressed genes. c, d, GSEA analysis of hallmark pathways shows 17 upregulated and 17 downregulated pathways in CRAT knockdown (KD) NRVMs. e. Heatmap analysis of the major enzymes involved in cholesterol biosynthesis pathway.
Extended Data Fig. 8 Bile acids and 7-HOCA induce type I interferon responses in NRVMs.
a, b, NRVMs were treated with different doses of CDCA for 1 day. Cells were then harvested for RT-PCR to determine the expression of the indicated ISGs. c, d, NRVMs were treated with different doses of CDCA, 7-HOCA or MCA for 1 day. Cells were then harvested for RT-PCR to determine the expression of the indicated ISGs. n = 6 (a-d) biologically independent samples per group. Data are shown as the mean ± SEM. Analysis was one-way ANOVA (a) or two-way ANOVA (c-d) followed by Bonferroni’s multiple comparison test and unpaired two-tailed Student’s t-test (b).
Extended Data Fig. 9 PPARs inhibitor, GW9662, inhibits the type I interferon responses in CRAT-deficient NRVMs.
NRVMs were transduced with lentivirus expressing control shRNA or Crat shRNA. Cells were then treated with GW9662 (10 μM) or T0070907 (10 μM) as indicated for two days. a, b, RT-PCR indicated that GW9662 inhibited the induction of Cyp27a1 (a) and Cyp7b1(b) in CRAT-deficient NRVMs. c, d, GW9662 but not T0070907 abolished the increase in ISG expression induced by CRAT depletion. n = 6 (a-c), 4 (d) biologically independent samples per group. Data are shown as the mean ± SEM. Analysis was two-way ANOVA (a-b) or one-way ANOVA (d) followed by Bonferroni’s multiple comparison test and unpaired two-tailed Student’s t-test (c). NS, not significant.
Extended Data Fig. 10 Mitochondrial DNA activates AIM2-inflammasome in vitro.
Purified GST–AIM2 (2 μg), ASC protein (1 μg), His-Caspase-1 (2 μg) were incubated in the presence or absence of mtDNA (1 μg) or poly (dA:dT) (1 μg) in TE buffer for 30 min at 37 °C. The reaction mixtures were then loaded on SDS–PAGE gel and Western-blots were performed with the indicated antibodies. Here is the representative result from three independent experiments.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2.
Source data
Source Data Figs. 1–6
Statistical source data for Figs. 1–6.
Source Data Fig. 1
Unprocessed western blots for Fig. 1d.
Source Data Fig. 3
Unprocessed western blots for Fig. 3i.
Source Data Fig. 5
Unprocessed western blots for Fig. 5b,g.
Source Data Fig. 6
Unprocessed western blots for Fig. 6i.
Source Data Extended Data Figs. 1–9
Statistical source data for Extended Data Figs. 1–9.
Source Data Extended Data Fig. 2
Unprocessed western blots for Extended Data Fig. 2a.
Source Data Extended Data Fig. 6
Unprocessed western blots for Extended Data Fig. 6a.
Source Data Extended Data Fig. 10
Unprocessed western blots for Extended Data Fig. 10.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mao, H., Angelini, A., Li, S. et al. CRAT links cholesterol metabolism to innate immune responses in the heart. Nat Metab 5, 1382–1394 (2023). https://doi.org/10.1038/s42255-023-00844-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-023-00844-5
This article is cited by
-
Cholesterol catabolism and bile acid synthesis in cardiomyocytes promote inflammation and heart failure
Nature Reviews Cardiology (2023)