Abstract
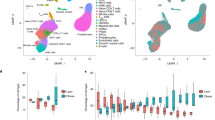
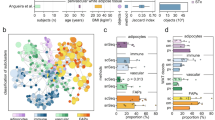
The complex relationship between metabolic disease risk and body fat distribution in humans involves cellular characteristics that are specific to body fat compartments. Here we show depot-specific differences in the stromal vascular fraction of visceral and subcutaneous adipose tissue by performing single-cell RNA sequencing of tissue specimens from obese individuals. We characterize multiple immune cells, endothelial cells, fibroblasts, adipose and haematopoietic stem cell progenitors. Subpopulations of adipose-resident immune cells are metabolically active and associated with metabolic disease status, including a population of potential dysfunctional CD8+ T cells that express metallothioneins. We identify multiple types of adipocyte progenitors that are common across depots, including a subtype enriched in individuals with type 2 diabetes. Depot-specific analysis reveals a class of adipocyte progenitors unique to visceral adipose tissue, which shares common features with beige pre-adipocytes. Our human single-cell transcriptome atlas across fat depots provides a resource to dissect the functional genomics of metabolic disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Raw data files from single-cell and bulk RNA-seq used in this study are uploaded in the Gene Expression Omnibus (GEO) in a SuperSeries with accession number: GSE136230. The expression data from the MuTHER cohort have been deposited in the ArrayExpress (https://www.ebi.ac.uk/arrayexpress/) with accession number E-TABM-1140. Source data are available online for Figs. 3, 5 and 6, and Supplementary Fig. 6.
References
Tchernof, A. & Despres, J. P. Pathophysiology of human visceral obesity: an update. Physiol. Rev 93, 359–404 (2013).
Porter, S. A. et al. Abdominal subcutaneous adipose tissue: a protective fat depot? Diabetes Care 32, 1068–1075 (2009).
Laforest, S., Labrecque, J., Michaud, A., Cianflone, K. & Tchernof, A. Adipocyte size as a determinant of metabolic disease and adipose tissue dysfunction. Crit. Rev. Clin. Lab. Sci. 52, 301–313 (2015).
Denis, G. V. & Obin, M. S. ‘Metabolically healthy obesity’: origins and implications. Mol. Aspects Med. 34, 59–70 (2013).
Michaud, A. et al. Relevance of omental pericellular adipose tissue collagen in the pathophysiology of human abdominal obesity and related cardiometabolic risk. Int. J. Obes. (Lond) 40, 1823–1831 (2016).
Schipper, H. S., Prakken, B., Kalkhoven, E. & Boes, M. Adipose tissue-resident immune cells: key players in immunometabolism. Trends Endocrinol. Metab. 23, 407–415 (2012).
Nawaz, A. et al. CD206(+) M2-like macrophages regulate systemic glucose metabolism by inhibiting proliferation of adipocyte progenitors. Nat. Commun. 8, 286 (2017).
Olsen, T. K. & Baryawno, N. Introduction to Single-Cell RNA Sequencing. Curr. Protoc. Mol. Biol. 122, e57 (2018).
Schwalie, P. C. et al. A stromal cell population that inhibits adipogenesis in mammalian fat depots. Nature 559, 103–108 (2018).
Helper, C. et al. Identification of functionally distinct fibro-inflammatory and adipogenic stromal subpopulations in visceral adipose tissue of adult mice. eLife 7, e39636 (2018).
Ehrlund, A. et al. The cell-type specific transcriptome in human adipose tissue and influence of obesity on adipocyte progenitors. Sci. Data 4, 170164 (2017).
Briot, A. et al. Senescence alters PPARγ (peroxisome proliferator-activated receptor gamma)-dependent fatty acid handling in human adipose tissue microvascular endothelial cells and favors inflammation. Arterioscler. Thromb. Vasc. Biol. 38, 1134–1146 (2018).
Banerji, S. et al. LYVE-1, a new homologue of the CD44 glycoprotein, is a lymph-specific receptor for hyaluronan. J. Cell Biol. 144, 789–801 (1999).
Schluns, K. S., Kieper, W. C., Jameson, S. C. & Lefrancois, L. Interleukin-7 mediates the homeostasis of naive and memory CD8 T cells in vivo. Nat. Immunol. 1, 426–432 (2000).
Michelet, X. et al. Metabolic reprogramming of natural killer cells in obesity limits antitumor responses. Nat. Immunol. 19, 1330–1340 (2018).
Schall, T. J. et al. A human T cell-specific molecule is a member of a new gene family. J. Immunol. 141, 1018–1025 (1988).
Small, K. S. et al. Identification of an imprinted master trans regulator at the KLF14 locus related to multiple metabolic phenotypes. Nat. Genet. 43, 561–564 (2011).
Civelek, M. et al. Genetic regulation of adipose gene expression and cardio-metabolic traits. Am. J. Hum. Genet. 100, 428–443 (2017).
Kratz, M. et al. Metabolic dysfunction drives a mechanistically distinct proinflammatory phenotype in adipose tissue macrophages. Cell Metab. 20, 614–625 (2014).
Jager, N. A. et al. Folate receptor-β imaging using 99mTc-folate to explore distribution of polarized macrophage populations in human atherosclerotic plaque. J. Nucl. Med. 55, 1945–1951 (2014).
Liao, X. et al. Kruppel-like factor 4 regulates macrophage polarization. J. Clin. Invest. 121, 2736–2749 (2011).
Villani, A. C. et al. Single-cell RNA-seq reveals new types of human blood dendritic cells, monocytes, and progenitors. Science 356, eaah4573 (2017).
Acosta, J. R. et al. Increased fat cell size: a major phenotype of subcutaneous white adipose tissue in non-obese individuals with type 2 diabetes. Diabetologia 59, 560–570 (2016).
Ribeiro, R. et al. Human periprostatic white adipose tissue is rich in stromal progenitor cells and a potential source of prostate tumor stroma. Exp. Biol. Med. (Maywood) 237, 1155–1162 (2012).
Yang, R. Z. et al. Identification of omentin as a novel depot-specific adipokine in human adipose tissue: possible role in modulating insulin action. Am. J. Physiol. Endocrinol. Metab. 290, E1253–E1261 (2006).
de Souza Batista, C. M. et al. Omentin plasma levels and gene expression are decreased in obesity. Diabetes 56, 1655–1661 (2007).
Watanabe, T., Watanabe-Kominato, K., Takahashi, Y., Kojima, M. & Watanabe, R. Adipose tissue-derived omentin-1 function and regulation. Compr. Physiol. 7, 765–781 (2017).
Chau, Y. Y. et al. Visceral and subcutaneous fat have different origins and evidence supports a mesothelial source. Nat. Cell Biol. 16, 367–375 (2014).
Winnier, D. A. et al. Transcriptomic identification of ADH1B as a novel candidate gene for obesity and insulin resistance in human adipose tissue in Mexican Americans from the Veterans Administration Genetic Epidemiology Study (VAGES). PLoS One 10, e0119941 (2015).
Vaittinen, M. et al. MFAP5 is related to obesity-associated adipose tissue and extracellular matrix remodeling and inflammation. Obesity (Silver Spring) 23, 1371–1378 (2015).
Hou, S. et al. S100A4 protects mice from high-fat diet-induced obesity and inflammation. Lab. Invest. 98, 1025–1038 (2018).
Kuefner, M. S. et al. Secretory phospholipase A2 group IIA modulates insulin sensitivity and metabolism. J. Lipid. Res. 58, 1822–1833 (2017).
Perdikari, A. et al. BATLAS: deconvoluting brown adipose tissue. Cell Rep. 25, 784–797 e784 (2018).
Wankhade, U. D. et al. TGF-β receptor 1 regulates progenitors that promote browning of white fat. Mol. Metab. 16, 160–171 (2018).
Zou, Y. et al. IRX3 promotes the browning of white adipocytes and its rare variants are associated with human obesity risk. EBioMedicine 24, 64–75 (2017).
Roberts, A. C. & Porter, K. E. Cellular and molecular mechanisms of endothelial dysfunction in diabetes. Diab. Vasc. Dis. Res. 10, 472–482 (2013).
Escobedo, N. & Oliver, G. The lymphatic vasculature: Its role in adipose metabolism and obesity. Cell Metab. 26, 598–609 (2017).
Nishimura, S. et al. CD8+ effector T cells contribute to macrophage recruitment and adipose tissue inflammation in obesity. Nat. Med. 15, 914–920 (2009).
Wu, H. et al. T-cell accumulation and regulated on activation, normal T cell expressed and secreted upregulation in adipose tissue in obesity. Circulation 115, 1029–1038 (2007).
Singer, M. et al. A distinct gene module for dysfunction uncoupled from activation in tumor-infiltrating T cells. Cell 166, 1500–1511 e1509 (2016).
Russo, L. & Lumeng, C. N. Properties and functions of adipose tissue macrophages in obesity. Immunology 155, 407–417 (2018).
Coats, B. R. et al. Metabolically activated adipose tissue macrophages perform detrimental and beneficial functions during diet-induced obesity. Cell Rep. 20, 3149–3161 (2017).
Song, N. J. et al. Small molecule-induced complement factor D (Adipsin) promotes lipid accumulation and adipocyte differentiation. PLoS One 11, e0162228 (2016).
Li, C. Y. et al. Comparative analysis of human mesenchymal stem cells from bone marrow and adipose tissue under xeno-free conditions for cell therapy. Stem Cell Res. Ther. 6, 55 (2015).
Acosta, J. R. et al. Single cell transcriptomics suggest that human adipocyte progenitor cells constitute a homogeneous cell population. Stem Cell Res. Ther. 8, 250 (2017).
Chung, S. S. et al. Glutathione peroxidase 3 mediates the antioxidant effect of peroxisome proliferator-activated receptor γ in human skeletal muscle cells. Mol. Cell Biol. 29, 20–30 (2009).
Hammarstedt, A. et al. WISP2 regulates preadipocyte commitment and PPARγ activation by BMP4. Proc. Natl Acad. Sci. USA 110, 2563–2568 (2013).
Jang, M. K. & Jung, M. H. ATF3 inhibits PPARγ-stimulated transactivation in adipocyte cells. Biochem. Biophys. Res. Commun. 456, 80–85 (2015).
Kim, J. Y. et al. Activating transcription factor 3 is a target molecule linking hepatic steatosis to impaired glucose homeostasis. J. Hepatol. 67, 349–359 (2017).
Wu, J. et al. Beige adipocytes are a distinct type of thermogenic fat cell in mouse and human. Cell 150, 366–376 (2012).
Tchernof, A. et al. Regional differences in adipose tissue metabolism in women: minor effect of obesity and body fat distribution. Diabetes 55, 1353–1360 (2006).
Laitinen, A. et al. A robust and reproducible animal serum-free culture method for clinical-grade bone marrow-derived mesenchymal stromal cells. Cytotechnology 68, 891–906 (2016).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172 (2019).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13 (2009).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
This work was supported by a Canadian Institute of Health Research (CIHR) team grant awarded to E.G. (EGM141898) and the CIHR-funded Epigenome Mapping Centre at McGill University (EP1-120608) awarded to T.P. E.G. holds the Roberta D. Harding & William F. Bradley, Jr. Endowed Chair in Genomic Research and T.P. holds the Dee Lyons/Missouri Endowed Chair in Pediatric Genomic Medicine. A.T. is the director of a Research Chair in Bariatric and Metabolic Surgery. M.C.V. is the recipient of the Canada Research Chair in Genomics Applied to Nutrition and Metabolic Health (tier 1). The MuTHER study was funded by a program grant from the Wellcome Trust (081917/Z/07/Z) and core funding for the Wellcome Trust Centre for Human Genetics (090532). The TwinsUK study was funded by the Wellcome Trust and European Community’s Seventh Framework Programme (FP7/2007-2013). The TwinsUK study also receives support from the National Institute for Health Research (NIHR)-funded BioResource, Clinical Research Facility and Biomedical Research Centre based at Guy’s and St Thomas’ NHS Foundation Trust in partnership with King’s College London. The authors thank M. Korhonen and M. Ahti at the Finnish Red Cross Blood Service, Helsinki, Finland, for valuable help in this study. The GTEx Project was supported by the Common Fund of the Office of the Director of the NIH, and by the National Cancer Institute (NCI), National Human Genome Research Institute (NHGRI), National Heart, Lung and Blood Institute (NHLBI), National Institute of Drug Abuse (NIDA), National Institute of Mental Health (NIMH) and National Institue of Neurological Disorders and Stroke (NINDS). The data used for the analyses described in this manuscript were obtained from the dbGaP accession number phs000424.v7.p2 on 7 October 2019.
Author information
Authors and Affiliations
Contributions
E.G. conceived the study. E.G., A.T., T.P. and M.-C.V. designed experiments. A.T., J.D.F. and M.-F.G prepared or provided the clinical samples. L.B. and B.B. managed the clinical aspects of the study. A.P., R.L.B., M.G., D.A.L. and M.-M.S performed the experiments. W.A.C, J.J.J., H.D., A.S. and G.B. provided bioinformatics support. A.L and J.N. provided the MSC cell lines and culture protocols. J.V. and E.G. analysed the data and drafted the manuscript. All authors reviewed and contributed feedback on the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
A.T. receives funding from Johnson & Johnson Medical Companies and Medtronic for research unrelated to the present manuscript. The other authors declare no competing interests.
Additional information
Editor recognition Primary handling editor: Elena Bellafante.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Multiple macrophage clusters were identified in SVF from both SAT and VAT depots.
Multiple macrophage clusters were identified in SVF from both SAT and VAT depots (a) 4 distinct macrophage clusters showing varying expression of CD68 (19–52% of cells), CD9 (10–51% of cells) and CD36 (25–72% of cells). The y axis of the violin plot indicate log transformed expression values and the width indicate number of cells expressing the particular gene. (b)Genes involves in lipid metabolism is found expressed in macrophage cluster – IS2, whereas IS3 is rich in inflammatory markers.
Extended Data Fig. 2 Gene expression of marker genes in 6 visceral specific progenitor clusters.
Gene expression of marker genes in 6 visceral specific progenitor clusters. The y axis of the violin plot indicate log transformed expression values and the width indicate number of cells expressing the particular gene.
Supplementary information
Supplementary Information
Supplementary Figures 1–7
Supplementary Tables
Supplementary Tables 1–23
Supplementary Data
Statistical source data for Supplementary Figure 6
Source data
Source Data Fig. 3
Statistical source data for Figure 3c
Source Data Fig. 5
Statistical source data for Figure 5d and 5f
Source Data Fig. 6
Statistical source data for Figure 6d
Rights and permissions
About this article
Cite this article
Vijay, J., Gauthier, MF., Biswell, R.L. et al. Single-cell analysis of human adipose tissue identifies depot- and disease-specific cell types. Nat Metab 2, 97–109 (2020). https://doi.org/10.1038/s42255-019-0152-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-019-0152-6
This article is cited by
-
Single-cell RNA sequencing in donor and end-stage heart failure patients identifies NLRP3 as a therapeutic target for arrhythmogenic right ventricular cardiomyopathy
BMC Medicine (2024)
-
PDGFRβ + cell HIF2α is dispensable for white adipose tissue metabolic remodeling and hepatic lipid accumulation in obese mice
Lipids in Health and Disease (2024)
-
Macrophage and T cell networks in adipose tissue
Nature Reviews Endocrinology (2024)
-
Clusters of Body Fat and Nutritional Parameters are Strongly Associated with Diabetic Kidney Disease in Adults with Type 2 Diabetes
Diabetes Therapy (2024)
-
Unraveling the complex roles of macrophages in obese adipose tissue: an overview
Frontiers of Medicine (2024)