Abstract
We assessed predictive models (PMs) for diagnosing Pneumocystis jirovecii pneumonia (PCP) in AIDS patients seen in the emergency room (ER), aiming to guide empirical treatment decisions. Data from suspected PCP cases among AIDS patients were gathered prospectively at a reference hospital's ER, with diagnoses later confirmed through sputum PCR analysis. We compared clinical, laboratory, and radiological data between PCP and non-PCP groups, using the Boruta algorithm to confirm significant differences. We evaluated ten PMs tailored for various ERs resource levels to diagnose PCP. Four scenarios were created, two based on X-ray findings (diffuse interstitial infiltrate) and two on CT scans (“ground-glass”), incorporating mandatory variables: lactate dehydrogenase, O2sat, C-reactive protein, respiratory rate (> 24 bpm), and dry cough. We also assessed HIV viral load and CD4 cell count. Among the 86 patients in the study, each model considered either 6 or 8 parameters, depending on the scenario. Many models performed well, with accuracy, precision, recall, and AUC scores > 0.8. Notably, nearest neighbor and naïve Bayes excelled (scores > 0.9) in specific scenarios. Surprisingly, HIV viral load and CD4 cell count did not improve model performance. In conclusion, ER-based PMs using readily available data can significantly aid PCP treatment decisions in AIDS patients.
Similar content being viewed by others
Introduction
Global HIV data show that in 2021, 38,4 million people were living with HIV (PLHIV) worldwide, with 650,000 associated deaths. Most of these deaths occurred in Sub-Saharan Africa, followed by East Asia and Latin America1,2. In Brazil, recent data reported 50,000 new annual infections, a 5% increase since 2010, and almost 13,000 associated deaths3. Unfortunately, late presentation to care and initiation of antiretroviral therapy (ART) with advanced HIV disease are still common in Latin America, with almost 56% of the new diagnoses having T CD4 lymphocytes (CD4) counts below 200 cells/mm3 at the time of diagnosis2,4. Consequently, opportunistic infections remain a major cause of HIV-associated deaths in this region5,6.
Although Pneumocystis jirovecii pneumonia (PCP) incidence has continuously decreased after the introduction of ART and prophylaxis7,8, it remains among the leading pulmonary opportunistic infections in several developing and developed countries5,6. The estimated incidence in Brazilian AIDS patients varies widely, ranging from 5.6 to 36%, owing to the variability in the methods and source of samples used to reach the diagnosis9,10. PCP accounts for almost 400,000 cases/year, with 200,000 deaths/year, mainly in developing countries11.
Diagnosing PCP continues to pose challenges due to various factors, including the lack of conventional culture systems for P. jirovecii12, the limited specificity of clinical symptoms, the reduced sensitivity of the usual diagnostic methods, and the complexities associated with sample collection. Numerous studies have highlighted the polymerase chain reaction (PCR) assay as a more sensitive method for diagnosing PCP. However, no standard technique has been widely incorporated in routine laboratories, nor are molecular biology and biomarkers assays easily accessible13. As a result, the lack of a PCP diagnosis leads to the implementation of empirical treatment in almost all cases, particularly in resource-limited settings.
More recently, the expanded use of machine learning (ML) has increased the possibilities of using health care data, enabling the creation of systems that assist human decision14. ML has already been tested in different areas of health care, showing promising clinical applications15. Several reports of ML application in infectious diseases improved the diagnosis, especially in settings lacking specific laboratory or radiology tests16.
Our research aimed to identify and evaluate predictors associated with PCP in AIDS patients among different types of supervised ML algorithms. We constructed predictive models based on clinical, laboratory, and radiology aspects easily accessible at most emergency rooms (ERs), including those from low-income countries. Some of the predictive models achieved high accuracy in different ERs’ scenarios. They can constitute valuable tools to improve the physicians' decision-making process of treating AIDS patients with suspected PCP.
Material and methods
Study design and patients
This was a prospective study that enrolled AIDS patients admitted between December 2016 and February 2020 at the ER of the Instituto de Infectologia Emílio Ribas (IIER), who were initially suspected of having PCP according to the following criteria: the presence of subacute cough and dyspnea (≥ 7 days), a current CD4 cell count < 250 cells/mm3, and poor compliance to or not on ART. Induced sputum was collected in a room with negative pressure before starting treatment for PCP (or with up to one dose) through inhalation of hypersaline solution (3–5% of NaCl), for 15–20 min, collected in a sterile container and stored at 4ºC until DNA extraction up to the next day, as previously described17. We performed an “in-house” quantitative PCR (qPCR) assay after DNA extraction of induced sputum, and serum samples collected simultaneously to the induced sputum were tested with the Fungitell® assay18 (Associates of Cape Cod, East Falmouth, MA, USA) for (1,3)-β-d-glucan (BDG) measurement according to the manufacturer's instructions.
We used this qPCR as standard diagnoses and considered patients with PCP when the threshold (Cq) of the qPCR was less or equal to 31 and colonized or without PCP when Ct was greater than 31, as previously described17. We collected demographic, clinical, laboratory, and radiological data of all patients. To predict PCP, we opted to include data usually associated with PCP in AIDS patients, which could be quickly accessed at ERs with different levels of resources (Table 1).
Statistical analysis
All categorical variables were compared using Fisher's exact test, and continuous variables were tested for normal distribution using the Shapiro–Wilk test before statistical analysis. The Shapiro–Wilk test showed a non-normal distribution of all variables. The continuous variables were expressed as the median and interquartile range (IQR) and compared using the Student t test.
The patients' variables that were gathered were first tested by classical statistical models comparing the patients with qPCR-confirmed PCP with those in whom the qPCR ruled out PCP. The variables that presented statistical difference were additionally evaluated through Boruta algorithm (Fig. 1—Supplementary information)19. The validated variables were further analyzed using univariable and multivariable logistic regression to calculate the odds ratio (OR) and corresponding 95% confidence interval (CI) to confirm whether the selected variables are risk factors for PCP before being considered for use in the predictive models. All statistical analyses were performed using R Statistical Software v4.2.2 (R Core Team, 2022: A language and environment for statistical computing, R Foundation for Statistical Computing, Vienna, Austria)20. For all analyses, differences with p < 0.05 were deemed statistically significant.
Data preprocessing
Before model fitting, categorical variables were transformed into binary dummy variables, as most predictive models are affected by the difference in the variables' scales. As data contained various scales for various quantities (e.g., C-reactive protein (CRP), lactate dehydrogenase (LDH), CD4 cell count, HIV viral load), data normalization was necessary to rescale all numeric values with a standard deviation of one and a mean of zero. This makes the various predictive models more effective. All values were normalized to reduce the dimension-introduced bias using Z-score standardization21. The dataset was randomly divided into a 70% training set to construct the predictive model and a 30% testing set for performance assessment, stratifying by the PCP outcome22.
Missing values
For physical parameters, radiological and laboratory data, which were associated with observed variables based on the clinical decision practice, we identified missing, not at random. The overall dataset exhibited a missing data rate of 3%. For each variable requiring imputation, a bagged tree was created where the outcome is the PCP variable, and the predictors are all other variables. One advantage of the bagged tree is that it can accept predictors with missing values23. The matrix layout of all intersections is demonstrated in the supplementary material (Fig. 2—Supplementary information).
Imbalanced data
This dataset was unbalanced. In this study, an unbalanced ratio showed that the minority class was 51.2%, less than the majority class when analyzing the number of observations. Therefore, to reduce data bias, we opted for the synthetic minority over-sampling technique (SMOTE)24, which manages overfitting induced by a limited decision interval and controls the generation and distribution of manual samples using the minority class sample.
Predictive models
Predictive models training may overfit algorithms to the nuances of a specific dataset, resulting in a model that does not generalize well to new data22. We compared ten predictive models to evaluate their effectiveness in predicting PCP in patients with AIDS. For the linear model, we opted for simple probabilistic classifiers, such as Naïve Bayes (NB)25, elastic network model (EN)26, and linear support vector machines (LSVM)27. For the kernel-based model, we utilized a multilayer perceptron (MLP)28. For the decision tree approach, the random forest (RF) model29, decision tree, bagged trees (BT), boosted trees light GBM (LightGBM), and the extreme gradient boosting (XGBoost) model30 have been used. Finally, multi-class algorithms as nearest neighbor (NN) were built31. We aimed to include different classes of ML methods.
Evaluation metrics
In the training set, the k-fold cross-validation with three folds and ten resamples was used to mitigate the potential bias or variance issues stemming from a single train-test split. An ANOVA-based racing tuning method was employed to optimize the hyperparameters for each candidate model, focusing on accuracy enhancement32.
Finally, after completing adjustments and training with the training set, the models were evaluated against the test set to ensure an accurate estimation of the performance of the model candidates without overfitting. The accuracy, precision, recall, F1-Score, and the area under the ROC curve (AUC) of each model were evaluated to establish a model ranking. Generally, these metrics indicate good performance when scores exceed 0.8 and poor performance below 0.733.
Ethical approval
The Comitê de Ética em Pesquisa from the Instituto de Infectologia Emílio Ribas approved the study (protocol 06/2016). All study was conducted in accordance with relevant institutional guidelines, and all patients consented to participate by signing an informed consent form.
Results
Ninety-seven PLHIV admitted to the emergency unit of the IIER with respiratory manifestations suggestive of PCP were enrolled. Eight patients were excluded for being transferred to another health service within the first 24 h of admission (n = 6) or for failing to provide induced sputum (n = 2). Therefore, 86 patients underwent the radiology and laboratory workouts prescribed by the attending physician. Variables statistically different between the two groups, with and without qPCR-proven PCP, are shown in Table 1. Additional sociodemographic and clinical data are shown in Supplementary Table 1. Patients with PCR results suggestive of colonization were grouped with the PCR negative patients, since the purpose of the study was to support the treatment decision.
As previously described, the two groups did not significantly differ regarding sociodemographic aspects or other clinical, radiology, and laboratory variables17.
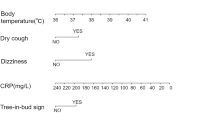
In our study, the clinical, laboratory, and radiological variables commonly associated with PCP that showed statistical differences were as follows: dry cough, increased respiratory frequency, decreased O2 saturation (O2sat) in arterial blood gas, elevated LDH levels, lower CRP values, low CD4 cell count, higher HIV viral load, chest X-ray showing diffuse interstitial infiltrate (DII), CT scan indicating a “ground-glass” image, presence of associated cytomegalovirus disease (CMV), and higher BDG values. BDG value was excluded since it is not available in most Brazilian ERs. These variables were then submitted to Boruta's analyses to determine the weight of each to the diagnosis of PCP. Boruta's analysis validated all variables except CMV co-infection. Ground-glass opacity on the CT scan was most strongly associated with PCP prediction, followed by LDH, arterial O2sat, CRP, and HIV viral load. Less but still significantly associated with PCP prediction were chest X-ray with DII, CD4 cell count, a respiratory rate greater than 24 bpm, and dry cough (Fig. 1—Supplementary information).
In parallel, we also designed four possible scenarios aiming at encompassing the variable range of facilities provided at ERs in Brazil, as depicted in Table 2. We used six variables in two scenarios and eight variables in the other two. The scenarios were headed depending on whether the ER has X-ray equipment or a CT scan (which presents greater sensitivity for diagnosing interstitial pulmonary diseases34), associated with the following set of variables: LDH (U/L), O2sat on arterial blood (%), CRP (mg/dL), respiratory rate > 24 bpm and dry cough. As CD4 cell and HIV viral load are carried out only in a few Brazilian Ministry of Health's reference laboratories, their results are not promptly accessible, so they were included for analyses only in secondary scenarios as additional variables.
We applied ten predictive models, as described in the methods section, to the four scenarios and used five metrics to evaluate the designed models' performance, as presented in Tables 3, 4, 5, and 6. Recall is relevant in settings where no patient should miss specific treatment because, e.g., the disease may be life-threatening (as is the case in PCP). However, it can otherwise lead to the treatment of false positive cases. Precision informs the capacity of the model to indicate the correct treatment for true positive PCP cases. Accuracy corresponds to both the ability to implement treatment for truly positive PCP cases and not implementing treatment for negative patients. AUC indicates the utility of the predictor in giving the best points of balance between true positive and false positive rates and summarizing the performance across all operating point tradeoffs.
All ten models performed satisfactorily in the four scenarios, suggesting that selecting the variables based on prior knowledge of statistical and Boruta analyses was appropriate. Four notably performed particularly well: NB, NN, RF, and XGBoost. They in general yielded indices greater than 0.8 for most scenarios and all five metrics, which is the usual recommendation for diagnostic tests33. One of the most familiar criteria used in the literature to evaluate the performance of a predictive model is the AUC, whose overall performance allows us to compare the performance of the predictive models graphically. Figure 1 depicts the AUC for these four models in the four scenarios, showing frequent indices above 0.9. However, as our primary goal is to provide treatment only for true PCP cases, avoiding unnecessary treatment of non-PCP cases, we opted for accuracy as the major criterion. Accuracy measures the overall correctness for true positive and true negative patients, informing the ability to implement treatment for PCP and not for non-PCP patients. Furthermore, accuracy, precision, and negative predictive value are prevalence-dependent metrics, whereas AUC, recall, and specificity are prevalence-independent.
Area under the curve (AUC) of the predictive models with best performance calculated for each of the A, B, C and D scenarios: extreme gradient boosting (XGboost), Naïve Bayes, nearest neighbor, and random forest. Figure 1 shows AUC from predictive models that presented a greater performance for each scenario. Scenario A: NN, NB and RF. Scenario B: NB, RF, and XGBoost. Scenario C: NB, RF and NN. Scenario D: RF, NB and NN.
Concerning the scenario A (Table 3), which mimics the usual common ERs’ setting (i.e., an X-ray is available, but not a CT scan), the NN model yielded the highest accuracy score (0.923), followed closely by both RF and NB with 0.885. All three also showed an AUC > 0.9. NN and NB presented precision and recall indices > 0.8. Although the RF model reached the highest precision (1.0), it presented a low recall (0.7), negatively impacting its F1-score. In addition, a fourth model, EN, also showed high accuracy (> 0.8) but somewhat weaker precision (0.78) and recall (0.7) scores. The remaining six models performed modestly only compared to those above three yielded accuracy indices between 0.7 and 0.8 and three below 0.7, with variable performances below 0.8 in the other criteria.
In scenario B (Table 4), the models using CT scan instead of X-ray showed overall better performances than in scenario A, considering the remarkable (n = 8) number of predictive models that reached accuracy values > 0.8. This is likely because the thoracic CT scan has greater sensitivity than chest X-rays in detecting pulmonary interstitial lesions35. Differently from scenario A, in scenario B it was the NB that reached the highest accuracy (0.923) as well as ≥ 0.9 scores in the other metrics, especially the AUC, with a score of 0.981. Additional seven predictive models presented high accuracy scores (≥ 0.8), such as RF and XGBoost (0.885), with high scores (≥ 0.8) also in the other metrics. Although BT and NN showed good accuracy (0.846), NN yielded a modest precision (0.75), and BG a modest recall score (0.7). The remaining five models, decision tree, LightGBM, MP, EN, and LSVM, performed somewhat more modestly than those mentioned above.
The analyses of scenarios, including thorax CT scan, raised the issue of how important this variable for the models' performance is. Even though its recognized better performance for diagnosing interstitial diseases, in scenarios B and D the models reached scores like those with chest X-ray, except for the highest AUC of 0.981 with the NB in scenario B. The presence of “ground-glass” opacity in the thorax CT scan of PLHIV presenting pulmonary symptoms is well-established as highly associated with PCP or viral infections35. However, it is not a specific signal and should not be taken alone for diagnosing PCP, especially in AIDS patients who not uncommonly develop concomitant pulmonary opportunistic infections35. For this reason, we still recommend its utilization in settings where a CT scan is available.
In scenario C (Table 5), unexpectedly, adding CD4 cell count and HIV viral load to the variables of scenario A did not result in higher performances, with the highest accuracy score being 0.885 (NB). Four models reached an accuracy greater than 0.8, with recalls of 0.9. Still, three of them had precision values < 0.8, which can lead to the undesired outcome of implementing empirical treatment in non-PCP patients. Overall, the models' performance in this scenario was slightly weaker than in scenarios A and B.
Scenario D (Table 6), with the addition of CD4 cell count and HIV viral load to the set of variables, also did not further improve the model's accuracy. The highest accuracy score was reached with RF (0.923), which also yielded scores greater than 0.9 regarding precision, recall, and AUC, a performance much like that observed with the NB in scenario B. In scenario B, the other seven models presented accuracy scores > 0.8. NB reached the second-highest best accuracy (0.885), followed closely by decision tree, BT, NN, and LSVM (0.846). These four models also performed well in the other metrics, reaching values ≥ 0.8.
Discussion
Predictive models for diagnostic purposes have already been tested in different areas of health care36. Although many specialties were covered36, there has been special interest in evaluating predictive models to improve decision-making processes in infectious diseases, from diagnosis to the risk of developing symptomatic infection and from predicting severity/mortality or complications to treatment response. These studies applied a wide range of models, the most commonly used being support vector machine (SVM), XGBoost, decision tree, RF, and NB, and several metrics used in the present study36. Of the ten models we have tested, NB, RF, and NN presented the overall best performance, with NB being increasingly studied and generally yielding good accuracy results37.
The use of predictive models in infectious diseases can be exemplified by the numerous models tested as alternative methods to diagnosing SARS-CoV-2 infection in a period when laboratory diagnosis was a challenge due to the high volume of patients, among other issues38. For example, Mei et al. 2020, evaluated a data set acquired from Chinese patients for whom there was a clinical concern of COVID-19 between January and March 2020. SVM, RF, and MLP were applied using pulmonary CT scan data associated with easily accessible demographic, clinical, and laboratory variables similar to our study. Confirmatory diagnosis of COVID-19 infection was achieved by real-time PCR (RT-PCR), being positive in 46.9% of the cohort. In this study, MLP performed better than the other two models, reaching a sensitivity of 0.843, a specificity of 0.828, and an AUC of 0.92. However, contrary to our study, where imaging evaluation was based on the presence/absence of interstitial infiltrate/ground grass images according to the ER clinicians' interpretation, they used a convolutional neural network model for CT scan analyses, which limits its applicability to limited-resource ERs39. In addition, our slightly better results could be accounted for, at least in part, by using Boruta's analysis of selected PCP-associated variables. This step seems important to increase the performance and can bring more confidence and adhesion by the clinicians than using random variables. We also designed our study to test a larger number of models to find the one that provided the best fit.
Predictive models were also used to investigate other viral diseases with some diagnostic challenges40. Dengue diagnosis was retrospectively studied in a cohort of Paraguayan patients with fever and initial clinical dengue suspicion, subsequently confirmed either by IgM serology, virologic isolation, or RT-PCR. Moreover, the authors used the SVM, MLP, and radial basis function as predictive models throughout 37 clinical-epidemiological and demographic variables that can be associated with dengue. SVM performed better, reaching an accuracy of 0.92 as well as a sensitivity of 0.93 and specificity of 0.92, providing an apparently helpful tool for the viral infection diagnosis40.
Studies comparable to ours were also done in acute bacterial diseases but with less successful results. A study investigated several models in diagnosing Clostridioides difficile infection (CDI) in a cohort of inpatients undergoing C. difficile testing. This study used clinical-demographic and laboratory data and, as our study, ten different predictive models. However, all 10 presented weak performances, with AUC up to 0.60 (the single metric used). In addition, classical CDI-associated parameters were chosen, such as high white blood cells and creatinine value, which did not improve the performance. One possible concern is the eventual gastrointestinal tract colonization with C. difficile, which can confound the diagnosis: in this study, from 3514 possible CDI records, only 136 were confirmed41.
The use of predictive models to study invasive fungal infections is still rare despite the fact that diagnosis of such infections still poses a challenge: usual diagnostic methods (e.g., blood culture) exhibit low sensitivity (compared with other types of infectious agents), some fungi lack or have slow growing properties in culture media, and in several instances, differentiation between colonization and invasion is difficult42. A review of ML methods applied to clinical microbiology found 97 valid articles; only three dealt with fungal infections16. Ripoli et al. 2020, evaluated a model to predict candidemia bloodstream infection (CBI) in at-risk patients using the records of a cohort of 157 patients with confirmed candidemia (positive blood culture) compared to 138 patients with bacteremia. The RF was applied to 17 clinic-demographic variables associated with an increased risk of developing candidemia. This model reached an AUC of 0.87, a sensitivity of 0.84, and a specificity of 0.9143. As in the present study, the model's good performance was likely linked to the appropriate selection of variables. However, using blood culture as a gold standard may misdiagnose some patients, especially those with low fungal burden. These promising results warrant that validation studies or other prospective real-world studies are undertaken. Another recently published study applied predictive models similar to ours in the context of PCP in kidney transplant recipients, with good results. However, the focus was not on the diagnosis of PCP but on the design of a prognostic model to predict the development of severe disease following PCP in these patients44.
In fact, one major concern in ML studies aiming to improve medical processes is that there is little evidence that these models have entered into clinical practice. External validation is a mandatory step since assessing the model's reproducibility and generalization is fundamental. Predictive models should not be addressed before extensive evaluation since mistakes and patient harm can occur, which enhances the importance of clinical knowledge and judgment. However, a survey of PubMed using "prediction models" retrieved almost 90,000 related articles in the year 2019, but when searched allied with "external validation," only 7% of the studies remained45.
Although we are just beginning to understand the wealth of opportunities afforded by ML methods, there is a growing concern in the academic community that, because the products of these methods are not perceived in the same way as other medical interventions, they do not have well-defined guidelines for development and use, and rarely undergo the same degree of scrutiny as others new technologies. The kind of evidence necessary to adequately recommend the widespread use of ML methods is still debated46. Some steps should be followed to build confidence in the prediction model, such as adequate reporting of data source, study design, modeling processes, number of predictors, etc., which facilitates the interpretation and increases the clinician's confidence. Predictive models are not meant to replace a clinician's judgment, and they should be tested through application within existing workflows to convince clinicians of the test's applicability since they tend to resist processes that interfere with their routine or challenge their autonomy47,48.
Our study was conducted at the emergence room of a teaching reference center for infectious diseases, where the clinicians are highly skilled in diagnosing and treating AIDS-associated OIs. Empiric treatment was prescribed to 90% of the cohort's patients who subsequently confirmed the diagnosis of PCP, but also to 30% of the patients in whom PCP was later ruled out (data not shown). On the other hand, the NN (scenario A) and NB (scenario B) predictive models would also indicate treatment for 90% of the confirmed PCP patients while treating only 1 out of 16 (6.25%) non-PCP patients, even if used by inexperienced clinicians. Unexpectedly, including CD4 cell count and HIV viral load did not improve overall predictive models' performances (Table C and D), suggesting that, in our setting, they functioned only as marginal predictors. A likely explanation relies on the patients' inclusion criterium of absence or irregular use of ART. Almost all (95%) of the patients had comparable high HIV viral load, and all had comparable low CD4 cell count (< 250 CD4 cells/mm).
Conversely, we estimate that implementing our tested model in non-specialized infectious diseases ERs may bring even more substantial improvement in the empirical treatment of patients with presumed PCP. We plan to proceed with validation studies at our reference hospital and other ER settings where patients with PCP are less prevalent and the medical staff is not specially trained in PCP diagnosis. Other limitations of our study are the relatively small sample size of the cohort and the fact that the data source arose from a single, reference hospital for infectious diseases with a high burden of AIDS patients, making it important cross-validation studies with larger cohorts.
Conclusion
In conclusion, after testing scenarios mimicking different ER settings, representative of either low/middle or wealthy countries, we strongly recommend that validation studies to be conducted with NN in X-ray-equipped ERs and with NB for CT scan-equipped ERs. Our models could be easily implemented in ER routine protocols to aid clinicians, particularly those not skilled in HIV/AIDS opportunistic infections, in the decision of introducing (or not) empirical treatment for suspected PCP patients.
Data availability
The data used in this study are available from the corresponding author upon reasonable request.
Abbreviations
- ART:
-
Antiretroviral therapy
- AUC:
-
Area under the receiver operating characteristic (ROC) curve
- BDG:
-
(1-3)-β-B-glucan
- CD4:
-
Lymphocytes T CD4
- CMV:
-
Cytomegalovirus disease
- CRP:
-
C-reactive protein
- CT:
-
Computerized tomography
- DII:
-
Diffuse interstitial infiltrate
- ER:
-
Emergency room
- LDH:
-
Lactate dehydrogenase
- ML:
-
Machine learning
- PCP:
-
Pneumocystis jirovecii Pneumonia
- PCR:
-
Polymerase chain reaction
- PLHIV:
-
People living with HIV
References
HIV data and statistics. https://www.who.int/teams/global-hiv-hepatitis-and-stis-programmes/hiv/strategic-information/hiv-data-and-statistics.
Global HIV & AIDS statistics—Fact sheet. https://www.unaids.org/en/resources/fact-sheet.
HIV Country Profiles. https://cfs.hivci.org/index.html.
Belaunzarán-Zamudio, P. F. et al. The population impact of late presentation with advanced HIV disease and delayed antiretroviral therapy in adults receiving HIV Care in Latin America. Am. J. Epidemiol. 189, 564–572 (2020).
Núñez, I. et al. Late-onset opportunistic infections while receiving anti-retroviral therapy in Latin America: Burden and risk factors. Int. J. Infect. Dis. IJID Off. Publ. Int. Soc. Infect. Dis. 122, 469–475 (2022).
Ford, N. et al. Causes of hospital admission among people living with HIV worldwide: A systematic review and meta-analysis. Lancet HIV 2, e438–e444 (2015).
Mocroft, A. et al. Decline in the AIDS and death rates in the EuroSIDA study: An observational study. The Lancet 362, 22–29 (2003).
Elango, K. et al. Trends in the epidemiology and outcomes of pneumocystis pneumonia among Human Immunodeficiency Virus (HIV) Hospitalizations. Int. J. Environ. Res. Public. Health 19, 2768 (2022).
Focaccia, R. et al. Clinical and epidemiology evaluation of Aids-infected patients hospitalized between 2011 and 2016 in the Santos region of Brazil. Rev. Soc. Bras. Med. Trop. 52, e20180126 (2019).
Figueiredo-Mello, C., Naucler, P., Negra, M. D. & Levin, A. S. Prospective etiological investigation of community-acquired pulmonary infections in hospitalized people living with HIV. Medicine 96, e5778 (2017).
Fungal Disease Frequency - Gaffi | Gaffi - Global Action For Fungal Infections. https://gaffi.org/why/fungal-disease-frequency/.
Riebold, D. et al. Axenic long-term cultivation of Pneumocystis jirovecii. J. Fungi 9, 903 (2023).
Gits-Muselli, M. et al. The Fungal PCR Initiative’s evaluation of in-house and commercial Pneumocystis jirovecii qPCR assays: Toward a standard for a diagnostics assay. Med. Mycol. 58, 779–788 (2020).
Beam, A. L. & Kohane, I. S. Big data and machine learning in health care. JAMA 319, 1317–1318 (2018).
Topol, E. J. High-performance medicine: The convergence of human and artificial intelligence. Nat. Med. 25, 44–56 (2019).
Peiffer-Smadja, N. et al. Machine learning in the clinical microbiology laboratory: Has the time come for routine practice?. Clin. Microbiol. Infect. 26, 1300–1309 (2020).
Chagas, O. J. et al. Performance of a real time PCR for Pneumocystis jirovecii identification in induced sputum of aids patients: Differentiation between pneumonia and colonization. J. Fungi 8, 222 (2022).
Tran, T. & Beal, S. G. Application of the 1,3-β-D-glucan (fungitell) assay in the diagnosis of invasive fungal infections. Arch. Pathol. Lab. Med. 140, 181–185 (2016).
Kursa, M. B. Robustness of random forest-based gene selection methods. BMC Bioinformatics 15, 8 (2014).
R: The R Project for Statistical Computing. https://www.r-project.org/.
Aho, K. A. Foundational and Applied Statistics for Biologists Using R. (Chapman and Hall/CRC, 2016). doi:https://doi.org/10.1201/b16126.
Sidey-Gibbons, J. A. M. & Sidey-Gibbons, C. J. Machine learning in medicine: A practical introduction. BMC Med. Res. Methodol. 19, 64 (2019).
Breiman: Classification and regressiontrees, wadsworth... - Google Acadêmico. https://scholar.google.com/scholar_lookup?journal=Biometrics&title=Classification+and+regression+trees+(CART)&author=L.+Breiman&author=J.+H.+Friedman&author=R.+A.+Olshen&author=C.+J.+Stone&volume=40&issue=3&publication_year=1984&.
Chawla, N. V., Bowyer, K. W., Hall, L. O. & Kegelmeyer, W. P. SMOTE: Synthetic minority over-sampling technique. J. Artif. Intell. Res. 16, 321–357 (2002).
Efron, B. Mathematics. Bayes’ theorem in the 21st century. Science 340, 1177–1178 (2013).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Cortes, C. & Vapnik, V. Support-vector networks. Mach. Learn. 20, 273–297 (1995).
Menzies, T., Kocagüneli, E., Minku, L., Peters, F. & Turhan, B. Using goals in model-based reasoning. In: Menzies, T., Kocagüneli, E., Minku, L., Peters, F. & Turhan, B.) (eds.) Sharing Data and Models in Software Engineering 321–353 (Morgan Kaufmann, Boston, 2015). doi:https://doi.org/10.1016/B978-0-12-417295-1.00024-2.
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Chen, T. et al. xgboost: Extreme Gradient Boosting. (2023).
Altman, N. S. An introduction to kernel and nearest-neighbor nonparametric regression. Am. Stat. 46, 175–185 (1992).
Kuhn, M. Futility Analysis in the Cross-Validation of Machine Learning Models. Preprint at https://doi.org/10.48550/arXiv.1405.6974 (2014).
Royston, P., Moons, K. G. M., Altman, D. G. & Vergouwe, Y. Prognosis and prognostic research: Developing a prognostic model. BMJ 338, b604 (2009).
Gruden, J. F. et al. High-resolution CT in the evaluation of clinically suspected Pneumocystis carinii pneumonia in AIDS patients with normal, equivocal, or nonspecific radiographic findings. Am. J. Roentgenol. 169, 967–975 (1997).
Hartman, T. E., Primack, S. L., Müller, N. L. & Staples, C. A. Diagnosis of thoracic complications in AIDS: Accuracy of CT. Am. J. Roentgenol. 162, 547–553 (1994).
Saberi-Karimian, M. et al. Potential value and impact of data mining and machine learning in clinical diagnostics. Crit. Rev. Clin. Lab. Sci. 58, 275–296 (2021).
Jiang, X., Xue, D., Brufsky, A., Khan, S. & Neapolitan, R. A new method for predicting patient survivorship using efficient bayesian network learning. Cancer Inform. 13, 47–57 (2014).
Adamidi, E. S., Mitsis, K. & Nikita, K. S. Artificial intelligence in clinical care amidst COVID-19 pandemic: A systematic review. Comput. Struct. Biotechnol. J. 19, 2833–2850 (2021).
Mei, X. et al. Artificial intelligence–enabled rapid diagnosis of patients with COVID-19. Nat. Med. 26, 1224–1228 (2020).
Mello-Román, J. D., Mello-Román, J. C., Gómez-Guerrero, S. & García-Torres, M. Predictive models for the medical diagnosis of dengue: A case study in Paraguay. Comput. Math. Methods Med. 2019, e7307803 (2019).
Marra, A. R. et al. Modest Clostridiodes difficile infection prediction using machine learning models in a tertiary care hospital. Diagn. Microbiol. Infect. Dis. 98, 115104 (2020).
Pasqualotto, A. & Denning, D. Diagnosis of invasive fungal infections: Current limitations of classical and new diagnostic methods. Eur. Oncol. Haematol. https://doi.org/10.17925/EOH.2005.0.0.1p (2005).
Ripoli, A. et al. Personalized machine learning approach to predict candidemia in medical wards. Infection 48, 749–759 (2020).
Liu, Y. et al. Machine learning models for prediction of severe Pneumocystis carinii pneumonia after kidney transplantation: A single-center retrospective study. Diagnostics 13, 2735 (2023).
Ramspek, C. L., Jager, K. J., Dekker, F. W., Zoccali, C. & van Diepen, M. External validation of prognostic models: What, why, how, when and where?. Clin. Kidney J. 14, 49–58 (2021).
Dhindsa, K., Bhandari, M. & Sonnadara, R. R. What’s holding up the big data revolution in healthcare?. BMJ 363, k5357 (2018).
Benjamens, S., Dhunnoo, P. & Meskó, B. The state of artificial intelligence-based FDA-approved medical devices and algorithms: An online database. NPJ Digit. Med. 3, 118 (2020).
Lovejoy, C. A., Buch, V. & Maruthappu, M. Artificial intelligence in the intensive care unit. Crit. Care Lond. Engl. 23, 7 (2019).
Funding
No funding was received for conducting this study.
Author information
Authors and Affiliations
Contributions
O.J.C. participated in the construction of the research, inclusion of patients, collection of clinical samples, creation of the database, analyses of the database, construction, analysis, and review of the paper. F.A.R.G. contributed with the statistical analyses of the database and analysis of the paper. P.P.N. performed the analyses of clinical samples (RT-PCR in induced sputum and 1,3-Beta D Glucan in serum) and analysis of the paper. R.B. participated in the construction of the research, inclusion of patients, and analyses of the paper. V.L.F. participated in the analyses of RT-PCR in sputum and analysis of the paper. G.M.B.D.N. participated in the analyses of clinical samples (RT-PCR in induced sputum and 1,3-Beta D Glucan in serum), construction, analysis, and review of the paper, and overall supervision. G.B. participated in the construction of the research, analyses of the database, construction, analysis, and review of the paper, and overall supervision. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chagas, O.J., Gonçalves, F.A.R., Nagatomo, P.P. et al. Predictive models-assisted diagnosis of AIDS-associated Pneumocystis jirovecii pneumonia in the emergency room, based on clinical, laboratory, and radiological data. Sci Rep 14, 11247 (2024). https://doi.org/10.1038/s41598-024-61174-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-61174-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.