Abstract
How information flow is coordinated for managing transit of 1/3 of the genome through endomembrane pathways by the coat complex II (COPII) system in response to human variation remains an enigma. By examining the interactome of the COPII cage-assembly component Sec13, we show that it is simultaneously associated with multiple protein complexes that facilitate different features of a continuous program of chromatin organization, transcription, translation, trafficking, and degradation steps that are differentially sensitive to Sec13 levels. For the trafficking step, and unlike other COPII components, reduction of Sec13 expression decreased the ubiquitination and degradation of wild-type (WT) and F508del variant cargo protein cystic fibrosis transmembrane conductance regulator (CFTR) leading to a striking increase in fold stability suggesting that the events differentiating export from degradation are critically dependent on COPII cage assembly at the ER Golgi intermediate compartment (ERGIC) associated recycling and degradation step linked to COPI exchange. Given Sec13’s multiple roles in protein complex assemblies that change in response to its expression, we suggest that Sec13 serves as an unanticipated master regulator coordinating information flow from the genome to the proteome to facilitate spatial covariant features initiating and maintaining design and function of membrane architecture in response to human variation.
Similar content being viewed by others
Introduction
An understanding of the spatial covariant properties of information flow linking genotype to phenotype through transcription, translation, degradation, and endomembrane trafficking pathways1,2,3,4,5,6,7 is required for addressing the critical features regulating the genome to proteome transformation driving cell development, differentiation, and its response to aging and environmental stress. As part of this information flow system, an understanding of the spatial and temporal design of the coat complex II (COPII) system that distinguishes amongst > 8000 cargo destined for export from the endoplasmic reticulum (ER) and its response to human variation impacting protein folding in health and disease remains largely an enigma8,9,10,11,12,13,14,15. We have demonstrated that these events are part of an encompassing endomembrane ‘quality system (QS)’ designed to manage sequence-to-function-to-structure relationships dictated by the protein fold that can be quantitatively interrogated by a Gaussian process (GP) based spatial covariance (SCV) machine learning framework1,2,3,4,5,6,7. SCV reveals the intrinsic design features of the protein fold on a residue-by-residue basis at atomic resolution for the entire polypeptide sequence. SCV relationships provide detailed insight into the diverse roles of individual residues in the cell in the context of the spatial–temporal dynamics dictating endomembrane flow and function in health and disease1,2,3,4,5,7.
Although considerable progress has been made in unraveling the molecular basis of the basic pathways directing cargo flow through exocytic and endocytic compartments, how genome sequence information is transformed into endomembrane biology will require a deep understanding of the individual roles of trafficking machinery components. The first step in the exocytic pathway involves the assembly of COPII10,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32. In vitro and in vivo studies have now shown that Sar1, Sec23, Sec24, Sec13 and Sec31 are cytosolic proteins that are required for COPII coat formation8,9,10,25,27,32,33,34,35,36,37. Together, they direct nascent cargo in the ER to all downstream exocytic, endocytic and ER-associated autophagic pathways21,31,38,39,40. Coat recruitment is initiated through the activation of Sar1 by the guanine nucleotide exchange factor (GEF) Sec12. The membrane recruitment and activation of Sar1 is followed by the binding of the cytosolic heterodimeric Sec23/24 complex. The Sec23 subunit is a Sar1-specific GTPase-activating protein (GAP) that has a catalytic role in GTP hydrolysis. By contrast, Sec24 serves as a multi-functional cargo-adaptor subunit that makes cargo choices for ER export. In general, the Sec23/24 complex functions as an kinetic adaptor platform for the selection of cargo for export that is sensitive to, for example, both di-acidic exit41 and basic retention codes42 as well higher order protein–protein interactions10,18,19,22,28,43,44,45,46,47. Subsequently, the Sec 13/31 heterotetramer complex is recruited to the nascent cargo-containing coat complex assembly8,9,33,37,48,49,50,51,52. Sec13/31 is currently thought of as a cage assembly factor to recruit cargo laden Sec23/24 complexes to form coated vesicles that bud from the ER for transit to the Golgi8,9,33,37,53,54,55,56. More recently, they are thought to contribute to the formation of collars at ER export sites (ERES) that are proposed to spatially regulate ‘cargo flow’ into tubules for direct trafficking to downstream ER Golgi intermediate compartments (ERGIC), Golgi and/or lysosomal compartments25.
While COPII dependent anterograde cargo export from the ERGIC provides a mechanism for export, it also initiates the recruitment of COPI coats where COPI directs retrograde recycling of misfolded cargo51,57, ER cargo receptors and ER-resident proteins such as the folding chaperone Hsp70 paralog BiP, by exchange of COPII for COPI51,57. COPI mediated recycling can lead to cargo degradation by the ERAD machinery13,20,25,44,58,59,60,61,62, an important step in the overall QS maintaining normal function of a cell1,2,3,4,5,6,7. Curiously, Sec13, a WD-repeat β-propeller family member, is also involved in nuclear pore complex (NPC) assembly directing flow of mRNA from the nucleus to the cytoplasm63,64, the GATOR2 complex65,66,67 that serves as a nutrient sensor through mTOR in the lysosomal pathway, as well as interacting with other WD-repeat containing proteins that regulate, for example, neuronal function leading to severe peripheral neuropathies when mutated47. The full-extent of Sec13 interactions in the cell remains unknown- nor is the coordination of Sec13 function between these different systems directing the composition and dynamics of endomembrane structures appreciated. These results raise the possibility that Sec13 may have a more fundamental role in controlling global information flow in the cell from the genome in response to diverse SCV relationships1,2,3,4,5,6,7.
As a model cargo protein engaged by COPII3,4,7,8,9,33,34,41,50,51,68,69,70, the cystic fibrosis (CF) transmembrane conductance regulator (CFTR) is a multi-membrane–spanning polypeptide belonging to the ATP binding cassette (ABC) transporter family71,72. It is composed of five functional domains: two nucleotide binding domains (NBD1 and NBD2), two trans-membrane domains (TMD1 and TMD2) and one regulatory domain73,74,75,76,77. The CFTR molecule is largely a cytosolic oriented protein with the regulatory ATP binding domains NBD1 and NBD2 required for export from the ER4,78,79,80.The biogenesis of CFTR requires trafficking from the ER, the first step in the exocytic pathway, through the Golgi to its final destination at the apical cell surface of epithelial cells in multiple tissues. CFTR functions as a cAMP-sensitive chloride channel at the apical plasma membrane (PM) of polarized epithelial cells76,77,78. It is charged with maintaining ion balance and hydration in, among others, sweat, intestinal, pancreatic, and pulmonary tissues81. Over 2000 variants contribute to human disease with the primary variant affecting 70–80% of the population having a deletion of residue Phe508 (F508del) leading to instability in the ER, degradation and loss of function at the cell surface3,4,7,34,68,74,75,77,82,83,84,85,86,87,88,89,90. The loss of a functional CFTR channel disrupts ion homeostasis, resulting in increased mucus viscosity in the airway of the lung and ductal systems of the pancreas and liver, and hydration of the intestinal tract. The increased mucus viscosity causes increased risk for inflammation and infection by Pseudomonas aeruginosa and other bacterial pathogens in the lung81. We have applied GP based SCV machine learning1,2,3,4,5,6,7 to achieve a deep understanding of the QS that dictates management of CFTR variation across the entire CF population to begin to understand as a collective the basis for precision management of disease in the individual3,4,7.
To address the global role of the COPII cage assembly component Sec13 in information flow through the endomembrane system via cargo selection into the COPII coat, we generated a Sec13 interactome using human bronchial epithelial (HBE) cells expressing wild-type CFTR, identifying 226 and 174 proteins significantly increased or decreased upon siRNA mediated Sec13 silencing, respectively. These changes potentially impact nucleopore complex (NPC) assembly, trafficking, the autophagy-lysosomal function through GATOR2 complex regulating mTORC165,66,67, along with a number of unanticipated protein complexes that contribute to functionality of endomembrane systems in each cell type. We now show that siRNA Sec13 (siSec13) silencing dramatically (~ ten-fold) improves the stability WT CFTR for trafficking from the ER, leading to a decrease of both WT and F508del CFTR degradation. This striking change in stability was not observed in response to the silencing of other COPII components including Sar1a, Sec31, Sec23 and 24 isoforms. In conjunction with its additional roles in chromatin organization, transcription, translation, and autophagy, we now suggest that Sec13 based complex assembly serves as an unanticipated master regulator impacting information flow through endomembrane compartments3,4. By providing a pivotal ‘set-point’7 for diversion of traffic at the ERES-ERGIC interface through COPII dependent COPI exchange9,34,49,50,51,57,70,91, a step dictated by SCV relationships revealed by human variation1,2,3,4,5,6,7, Sec13 could have a central role in balancing global endomembrane function impacting health, disease and aging in response to the environment and natural selection.
Results
Sec13 interactome profile in human bronchial epithelial cells
To improve our understanding of COPII coat complex function in cargo traffic from the ER3,4, we first applied mass spectrometry (MS) to determine the Sec13 interactome given the central role of the Sec13/31 cage assembly complex in coordinating the assembly of Sec23/24 coat components that select newly synthesized cargo for maturation through exocytic pathway for function in downstream endomembrane compartments and at the cell surface9,32,54,56,92. This is the key step in the biology of the cell that controls information flow from the genome for nearly 1/3 of the proteome. For this purpose, we applied co-purifying protein identification technology (CoPIT), an immunoprecipitation (IP)-based proteomic-profiling approach for capturing protein–protein interactions coupled with multidimensional chromatography and MS87. Using a wild-type (WT) human bronchial (lung) epithelial (HBE) 41o- (HBE41o-) cell line harboring the COPII dependent cargo protein CFTR3,4,7,34,68,87, we recovered proteins mapping to 413 genes classified as high-confidence interactors (Fig. 1A, Supplementary Table S1) given the strong correlation observed among three biological replicates (Supplementary Fig. S1, Pearson’s r = 0.82–0.93). These 413 proteins form the core of the Sec13 interactome and include both direct and indirect interactions amenable to CoPIT recovery using a Sec13 specific antibody. An additional 1219 interactors with an increased fold-change over background, but with lower confidence scores, were further assembled into an extended interactome (Supplementary Table S2). All known interactors of Sec13 such as Sec31A of the COPII complex29,33,37,52,93, Nup98 and other components of the nuclear pore complex (NPC)94,95,96,97,98, as well as Mios, WDR24, WDR59 and Seh1L of the GATOR2 complex65,66,67, were identified, validating the CoPIT approach for discovery of additional Sec13 interactors (Fig. 1A).
Sec13 core interactome. (A) Volcano plot of 3753 quantified proteins from HBE cells recovered by Sec13 IP compared to control IP displaying the relationship between statistical significance and fold-change of each protein. The fold-change is plotted on the x axis, and the − log2 p-value is plotted on the y axis. The blue circles represent the 413 proteins that had statistically significant differential interactions, log2 fold-change > 0 (p-value < 0.05). (B) UpSet plots showing the intersections of protein sets interacting with Sec13 present in the nucleus (Nuc), cytoplasm (Cyt), mitochondria (Mit), endoplasmic reticulum (ER), cell membrane (CM), Golgi apparatus (Gol), endosome (End), secreted (Sec), peroxisome (Per) and lysosome (Lys). The connected circles indicate which locations are included in the intersection. The horizontal bar chart displays the total number of proteins significantly interacting with Sec13 in a specific location. The vertical bar chart displays the number of proteins significantly interacting with Sec13 at that intersection. (C) The 413 proteins composing the Sec13 core interactome were mapped to the STRING database for cellular component analysis. 15 independent cellular components highly enriched (STRING enrichment strength > 1) were identified. (D) Networks of highly enriched complexes in the Sec13 interactome displaying the fold-change (yellow to purple) and the p-value (big to small circle) of the Sec13 interactome data set for each of the individual known components of these complexes. Gray circles represent complex subunits that were not identified in the Sec13 interactome.
In addition to the known interactions, we identified multiple interactors distributed along the exocytic and endocytic pathways (Fig. 1B, Supplementary Fig. S1). Interestingly, only Sec31A was found in the Sec13 interactome- not other components of the COPII coat complex, including any of the cargo selection Sec23/24 isoforms. These results highlight the expected strong interaction between Sec13 and Sec31A, but also emphasize the weak and likely transient interactions between the first layer of the COPII coat complex containing Sec23/24 cargo complexes and the second layer cage assembly Sec13/Sec31A complex leading to cargo collection and export from the ER, a state that is believe to be rapidly disassembled, coupled to COPI exchange51,57 to allow continued flow of cargo for recycling or to downstream endomembrane compartments8,9,37,99.
WD40 repeat domains, such as found in Sec13 are largely recognized to coordinate the assembly of multi-protein complexes100,101,102. To understand this role, we calculated the cellular enrichment of the Sec13 interactome components through STRING analysis tool103 (Fig. 1C, Supplementary Fig. S2). Here, we observed that all highly enriched components are multi-protein complexes (Fig. 1C, D) such as its known interactions facilitating NPC assembly94,95,96,97 to direct flow of mRNA from the nucleus to the cytoplasm63,64, and the GATOR2 complex65,66,67 that serves as a nutrient sensor in the mTORC1 lysosomal/autophagy pathway. In addition, we now show that it also includes the eukaryotic translation initiation factor 2B (EIf2B) complex104, the chromatin acetylating STAGA complex105,106,107 and the ubiquitous AMP-activated protein kinase complex (PKC), among many others (Fig. 1C, D; Supplementary Fig. S2). These results highlight a more general role of the steady state pool of Sec13 in managing multi-protein complex assemblies involved in the continuum of processes stemming from chromatin organization through transcription in the nucleus as well as co-translational insertion into the ER to generate a functional protein fold that can interact with trafficking components to initiate and maintain the composition and flow of secretory and membrane-bound cargo through the endomembrane system to the cell surface.
Proteomic profiling of human bronchial epithelial cells treated with siSec13
To understand in more detail the role of Sec13 in the cell, we investigated the relative expression level of proteins in HBE41o- cells expressing WT CFTR that were treated with either siSec13 or a control scrambled siRNA (siScr) using a TMT-based isobaric labeling method followed by data acquisition using MudPIT108. siSec13 gives a nearly complete reduction of Sec13, although residual levels Sec13 may insure cell survival given that it is an essential protein32,114,115. We identified a total of 3603 distinct proteins, with 226 and 174 proteins significantly increased and decreased upon Sec13 silencing, respectively (Fig. 2A, Supplementary Table S3) with a high correlation between the different biological replicates (Fig. 2B, Pearson’s r = 0.99). These results suggest that loss of Sec13 can simultaneously either negatively or positively influence the stability of multiple pathways. Among the more stabilized pool, CFTR is the 19th most increased protein upon Sec13 silencing (Fig. 2A, Supplementary Fig. S3, Supplementary Table S3) (see below).
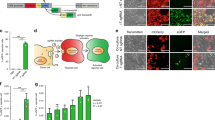
Proteome profiling following siRNA-mediated Sec13 silencing. (A) Volcano plot of 3603 quantified proteins from HBE41o- cells recovered following siRNA-mediated Sec13 silencing compared to siScr control displaying the relationship between statistical significance and fold-change of each protein. The fold-change is plotted on the x axis, and the − log2 p value is plotted on the y axis. The purple and orange circles represent the 184 and 241 proteins significantly decreased and increased upon Sec13 silencing, respectively (FC < 0.8 or FC > 1.2 with p-value < 0.05). (B) Scatterplot showing correlation of the NASF-scores among three replicates of quantified proteins recovered from HBE41o- cells following siRNA-mediated Sec13 silencing. (C) The 241 and 184 proteins significantly increased and decreased upon Sec13 silencing, respectively, were mapped to the STRING database for cellular component analysis. 5 and 11 cellular components are highly enriched among the proteins significantly increased and decreased upon Sec13 silencing, respectively. (D) Immunoblots (left panel) and quantifications of protein level (right panel) of individual COPII components following siSec13 treatment. Data are presented in right panel as fold-change relative to siScr control transfection (mean ± SEM, n ≥ 3). (E) Immunoblots (left panel) and quantifications of protein level (right panel) localized in different compartments of the cell following siSec13 treatment. Data are presented in right panel as fold-change relative to siScr control transfection (mean ± SEM, n ≥ 3). (F) Protein network of all lysosomal proteins that significantly decreased upon Sec13 silencing. (G) CFTR immunoblot of HBE41o- cells treated with DMSO or chloroquine following siRNA transfection with siSec13 or siScr. (H) Transferrin-receptor (TFRC) immunoblot of HBE41o- cells after siSec13 or siScr transfection. See Fig. S8 for full non-cropped blots used to generate figure.
We used the levels of differentially expressed proteins to calculate enrichment upon siSec13 treatment through the STRING analysis tool103 (Fig. 2A, C; Supplementary Fig. S3). We observed a significant increase in a large number of proteins in these cellular complexes, including components affecting ER to Golgi transport (Fig. 2A, Supplementary Fig. S3) and proteins localized to the lumen of the lysosome reflecting the role of Sec13 cage assembly in ER-associated autophagy-lysosome pathways (Fig. 2C)21,31,38,39,47,66,109,110,111,112,113. These results suggest a role for Sec13 in more globally managing protein fold stability (Fig. 1). To further refine the analysis, we compared the overlap between Sec13 interactors (Fig. 1A) to the changes in the global HBE41o- proteome following Sec13 silencing (Fig. 2; Supplementary Fig. S4, fold-change range < 0.8 or > 1.2), a cut-off to focus on more abundant interactors. This comparison revealed a common subset of Sec13-protein interactions affected by siSec13 silencing (Supplementary Fig. S4). As expected, we observed a prominent decrease in the Sec31A component involved in COPII cage assembly that requires Sec13 for stability2,131,132,133. We also observed changes in a number of Sec13 interactors found in the nucleus, ER, cytoplasm and in cytoskeletal organization. The combined results support a potential role for Sec13 in the stabilization and coordination of activities central to managing endomembrane function and structure.
Effect of siSec13 on COPII mediated trafficking
To begin to address the apparent complex role(s) of Sec13 in facilitating endomembrane biology, we first focused on its known role in ER export by quantifying the effect of siSec13 on the expression level of the COPII components using immunoblotting (Fig. 2D). We noted a modest increase of Sec24A and a small decrease of Sec24C, but a drastic decrease of Sec31A, a result consistent with the proteomic datasets (Figs. 2A, 3A, B) and, as indicated above, the dependance of Sec31A on Sec13 for its stability32,114,115 (Fig. 2D). Strikingly, within 24 h we saw a strong (ten-fold) stabilization of the mature band C glycoform of WT CFTR in response to Sec13 reduction reflecting successful export from the ER and processing by Golgi-associated glycosylation enzymes (Fig. 2D). This response was similar to the strong increase in the level of the mature ABCC4 glycoform, also known as multidrug resistance protein 4 (MDRP4)116, a protein similar to CFTR as part of the ABC transporter family (ABCC7)117. We also determined the level of select proteins localized in different compartments of the cell or extracellular space following secretion (Fig. 2E). For example, we observed a significant decrease in the level of TGN38 impacting post-trans Golgi network function and in maintaining TGN morphology118,119. Interestingly, we observed a drastic diminution of Niemann-Pick disease Type C1 (NPC1), a protein that is a resident protein in the lysosome (Fig. 2E), suggesting an important role of Sec13 cage assembly in managing resident cargo flow to the lysosomal network to prevent degradation in the ER13,21,23,31,39,112,120 and/or, for example, the general stability of the lysosomal system in response to the dependence of GATOR2 complex function on Sec1365,66,67.
siSec13 increases CFTR export and function. (A, C) Immunoblot analysis (upper) and quantifications (lower) of CFTR expression following transfection of HBE41o- (WT CFTR) (A) and CFBE41o- (F508del CFTR) (C) cells with the independent siRNA targeting the different COPII core subunits. Data are presented in lower panel as fold-change of CFTR band B and band C relative to siScr control transfection (mean ± SEM, n ≥ 3). (B, D) Representative fluorescent imaging plate reader traces (left) and quantification (right) of YFP-quenching following transfection of HBE41o- (WT CFTR) (B) and CFBE410- (F508del) (D) YFP cells with either siSec13 or siScr control compared to the -/- YFP cells not expressing CFTR. Data are presented as % relative to baseline (mean ± SEM, n ≥ 3). In all panels, ∗ indicates significant differences (p < 0.05) relative to siScr transfection determined by two-tailed t-test. See Fig. S9 for non-cropped blots used to generate figure.
To determine if Sec13 silencing affected WT CFTR degradation in lysosomes, which could be either due to trafficking directly from ER or during recycling between the cell surface through the endosomal-lysosomal system, we treated WT expressing HBE41o- cells with the lysosome-autophagosome inhibitor chloroquine121. In the absence of siSec13 silencing we observed a strong stabilization of CFTR band C glycoform to chloroquine which remained unaltered in response to siSec13 silencing (Fig. 2G). These results suggest that the post-ER cell surface recycling pool degraded by the lysosome confers increased stability to mature CFTR during the time-frame of siSec13 treatment (Fig. 2G). In contrast, we noted only a minor change in the ER band B CFTR glycoform (Fig. 2D), indicating that Sec13 silencing had little to no effect on degradation of ER residual glycoforms normally processed through ER-associated degradation (ERAD)20,68,88,90,122,123,124,125,126,127,128,129. These observations are consistent with a previous report showing that Sec13 silencing leads to a strong defect of autophagy in Saccharomyces cerevisiae130, and consistent with the observation that autophagy modulation can partially correct F508del CFTR stability and trafficking82. Interestingly, we saw no effect on the total secreted pool of the ER-generated extracellular protease alpha-1-antitrypsin (AAT). These results raise the possibility that the recycling pool of its abundant ER export receptors LMAN146 and/or SURF446,131,132,133 may be insensitive to the time-frame (24 h) of siSec13 silencing used to follow trafficking of CFTR and ABCC4 that utilize unique ER exit motifs for direct delivery to cell surface34,41,70. Likewise, siRNA Sec13 had no effect on the transferrin-receptor (TFRC)134,135 protein level in the time frame (24 h) used to detect increased WT CFTR band C stability, suggesting that Sec13 has no or minimal role in managing the long-term stability of late endocytic recycling pathway components (Fig. 2H).
The effect of Sec13 silencing on variant F508del CFTR export and function
To address how the COPII system through Sec 13 can discriminate between differentially folded states of protein cargo in vivo, we dissected the functional contribution of each of the COPII components for variant CFTR proteins. There are over 2000 variants of CFTR in the CF population that differentially contribute to disease onset. An analysis of the allele frequency of CF-causing mutations reveals that approximately 70–80% of patients carry at least one copy of a three base pair deletion leading to the loss of a phenylalanine at position 508 (F508del) in NBD1136,137,138,139. The F508del mutation disrupts the folding of the variant protein, leading to its loss of engagement by COPII in the ER compartment and clearance by ER-associated degradation (ERAD)76,77,140,141,142,143,144. Using Gaussian process (GP)-based machine learning with 71 variants triggering CF in the population4, we recently demonstrated1,2,3,4,5,6,7,145 the existence of a thermodynamically sensitive region of the CFTR fold involving the interfaces between nucleotide binding domain 1 (NBD1) contributed by the F508 and R560 residues, and the fourth intracellular loop 4 (ICL4) of transmembrane (TM) domain 2 (TMD2), the latter being responsible for COPII engagement for trafficking from the ER via the di-acidic exit motif3,4. We have shown that loss of function of the di-acidic code in the NBD1 domain is the primary defect responsible for CF in F508del-expressing patients3,4,7,41,50,70. This observation is consistent with MS analysis of residues undergoing methylation and ubiquitination where ubiquitination of di-acidic motif residues targets CFTR for degradation when challenged through misfolding by the F508del allele change87,146, and consistent with the need to assemble the ICL4 and NBD1 domains to achieve export73,74,75,77,85,86.
Using SDS-PAGE followed by immunoblotting to capture the conversion of the ER-localized band B glycoform to the Golgi-associated band C glycoform in response to export (Fig. 3A, B (% band B)), HBE41o- cells stably expressing WT CFTR (Fig. 3A) and CFBE41o- stably expressing F508del CFTR (Fig. 3B) were transfected with siRNA specific for each of the COPII components (Fig. 3A, B; Supplementary Fig. S5, WT CFTR; Supplementary S5, F508del CFTR). The level of silencing of each component was validated by immunoblotting (Supplementary Fig. S5). While we observed that silencing of various COPII components differentially modulate WT and F508del CFTR stabilization and trafficking (Fig. 3, Supplementary Fig. S5), only silencing of Sec13 yielded a strikingly strong (~ ten-fold) increase in stabilization of WT CFTR band C glycoform (Fig. 3A). Correspondingly, it also yielded a nearly 2.5-fold increase in total F508del band B glycoform (Fig. 3B). The next highest effect, but only for F508del CFTR, was silencing of Sar1b, potentially reflecting the increased sensitivity of general COPII assembly for recruitment of folding defective proteins in response to decreased Sec13/31 cage assembly complex (Fig. 3A, B; Supplementary Fig. S5). Specificity of Sec13 silencing was validated by using an siRNA recognizing a different region of Sec13 coding sequence (Supplementary Fig. S6). Notably, whereas WT-CFTR accumulated in the band C post-ER glycoform in response to siSec13, F508del accumulated largely as the band B glycoform with a modest but significant increase in band C (Fig. 3B, upper panel, siSec13 lane (band C arrow); quantitation lower panel). These results indicate that siSec13 is not rescuing F508del misfolding for export as do proteostasis components68,82,88,89,90 and CFTR modulators3,4,7,42,125,126,147,148,149,150,151,152.
To determine whether changes in steady state levels of Sec13 are responsible for protein stability and/or trafficking of WT CFTR, we performed a time-course of Sec13 silencing for 2, 3 and 4 days. We observed that decreasing levels of Sec13 leads to a correspondingly increase in WT CFTR band C that peaked by day 3 (Supplementary Fig. S6). To determine if the siSec13 stabilization and trafficking of CFTR was dependent on COPII assembly through the cargo collecting components Sec23/24, we knocked-down all Sec24 isoforms, thereby preventing CFTR WT or F508del recognition by the COPII system (Supplementary Fig. S6). We observed that the deletion of Sec24 isoforms eliminated siSec13 enhancement of CFTR stabilization and export for both WT and F508del, suggesting that the CFTR stability following Sec13 silencing requires the full functionality of the COPII machinery (Supplementary Fig. S6). The combined results suggest that reduction of the cage assembly complex Sec13/31 in response to decreasing levels Sec13 during silencing improves stability of WT CFTR pool in the ER for export, but only weakly corrects the ability of misfolded F508del for export.
We next determined if the partial rescue of band B F508del and the substantial increased band C fraction of WT CFTR represented a functional cell surface-localized chloride channel. In order to assess the functional status of WT and F508del CFTR treated with siSec13, we used HBE41o- and CFBE41o- cells stably expressing the halide-sensitive YFP-H148Q/I152L, a yellow fluorescent protein (YFP) variant whose fluorescence can be quenched in response to iodide influx entering the cell through a functional, cell surface localized CFTR channel153. When compared to cells lacking CFTR where quenching is not observed (Fig. 3C, orange), we observed that Sec13 silencing led to an increase in function at the cell surface for both WT (Fig. 3C) and F508del (Fig. 3D) consistent with its improved stability.
Combined, these results suggest that the role of Sec13 cage assembly is distinct from features such as the Sec23/24 cargo selection machinery dictating initial recruitment of cargo, but is consistent with a role in improving the overall availability of CFTR for ER export by reducing degradation at the ERES-ERGIC interface where COPII-COPI exchange differentiates export from recycling for degradation34,49,51,57,82,89,91,154,155,156, or in the case of WT CFTR, post-ER recycling to the lysosome sensitive to Sec13’s role in maintaining the lysosome. These results surprisingly suggest that steady-state levels of the Sec13 affecting the generation of the Sec13/31 complex regulates CFTR stability in both healthy and misfolded states and the more global state of the function of the early endomembrane system.
Sec13 silencing decreases CFTR ubiquitination and degradation via ERAD
To address the mechanism of action of Sec13 management of CFTR stability, chaperone assisted protein folding through proteostasis and ERAD have been shown to play an important role in triaging proteins for export by the COPII export machinery62,76,82,123,157. Newly synthesized proteins that do not fold correctly in the ER are generally targeted to the ubiquitin–proteasome ERAD pathway associated with retrograde trafficking from ERES and ERGIC20,51,57,62,158 facilitated by COPII-COPI exchange34,49,51,57,82,89,91,154,155,156. In addition, there is now recognized a COPII dependent autophagy pathway that delivers ER cargo directly to the lysosome13,21,23,31,39,112,120. One unusual feature of CFTR protein biogenesis is that during translation in heterologous expression systems, and as observed in primary cells expressing endogenous levels of CFTR, approximately 75% of WT CFTR and most variant F508del CFTR protein is thought to be degraded by the ERAD pathway- suggesting the existence of metastable state in fold design that is not responsive to the steady-state ER associated proteostasis machinery facilitating normal folding in multiple cell types3,4,49,77,78,82,89,90,159,160.
To understand the stabilization of WT CFTR upon siSec13 treatment, we first investigated the ubiquitination status of CFTR after silencing Sec13 by immunoprecipitation (Fig. 4A, Fig. S10B). siSec13 treated HBE41o- (WT) cell lysates were incubated with ubiquitin antibody and eluates were separated on SDS-PAGE gels under denaturing conditions (Fig. 4A). We observed that Sec13 silencing promoted a significant reduction of WT CFTR band B and band C levels recovered in ubiquitin immunoprecipitates (Fig. 4A), indicating that abrogation of Sec13 decreased the ubiquitination of WT CFTR, limiting its degradation leading to its increased stabilization. In addition, we investigated the possible additive and/or synergistic effect of siSec13 treatment with the proteasome inhibitor compound MG132 known to increase WT CFTR stability161 (Fig. 4B). Here, and as expected, we observed a strong MG132 effect on increasing WT CFTR stability and CFTR trafficking in HBE41o- (WT) lung cell lines (Fig. 4B). Interestingly, addition of MG132 to siSec13 treated cells did not result in any additive or synergistic effect between the two treatments alone (Fig. 4B). Given the observation that the impact of siSec13 on WT CFTR was much higher than the impact observed after MG132 treatment, these results suggest siSec13 modulates additional pathways managing CFTR stability- perhaps reflecting the role of the Sec13/31 complex in ER function at the ERES-ERGIC interface involved in COPII-COPI exchange9,51,57,77,78,89,90,155,156, and/or in its involvement ER-autophagy pathways.
siSec13 decreases CFTR ubiquitination, degradation via ERAD and increase CFTR stability. (A) Immunoblot (upper) and quantification (Lower) of CFTR expression in ubiquitin-containing immunoprecipitates following siSec13 transfection in HBE410- cells compared to siScr. Data are presented in lower panel as fold-change of CFTR band B and band C relative to siScr control transfection (mean ± SEM, n ≥ 2). (B) Immunoblot (left) and quantification (right) of CFTR expression (high and low exposure) following transfection of HBE41o- cells with siSec13 in combination with MG132 treatment compared to DMSO and siScr controls. Data are presented in the right panel as fold-change relative of CFTR band B and band C to siScr control transfection in combination with DMSO treatment (mean ± SEM, n ≥ 3). (C) Immunoblot (left) and quantification analysis (right) for CFTR, Hsp90, Hsc/p70 and Hsp40 in CFTR-containing immunoprecipitates following siSec13 transfection of HBE410- (WT) and CFBE41o- (F508del) cells compared to siScr (set to a value of 1, y-axis) (mean ± SEM, n ≥ 2). (D) (Upper panels) Representative immunoblot (left panel) from two experimental replicates of HBE41o- before and after proteolysis digestion with increasing concentration of trypsin (mg/ml) showing digestion pattern of the 60 kDa NBD1 domain and its protease resistant 30 kDa fragment (middle and right panels, respectively) using 3G11 antibody Two-Way ANOVA was applied to compare protein levels between siRNA Sec13 and controls (***, p < 0.001; **, 0.001 < p < 0.01; *, 0.01p < 0.05). (Lower panels) 60 and 30 kDa trypsin fragments and their quantitation recognized by a polyclonal antibody (Mr. Pink) that recognizes multiple CFTR epitopes75,85,86,171. See Fig. S10 for non-cropped blots used to generate figure.
One possibility for enhanced stability of CFTR is that Sec13 silencing induces ER stress leading to alteration of chaperone levels supporting ER function. We followed the siSec13 response levels of calnexin (a resident luminal chaperone), luminal Grp94 (an Hsp90 paralog) and ribophorin B (a resident membrane protein associated with the ER) (Supplementary Fig. S7) with Hsp90 serving as a cytosolic reporter. We did not observe any modification of the level of these proteins upon Sec13 silencing suggesting that siSec13 does not impact the general ER proteostasis environment through stress related unfolded protein response (UPR) pathways in the short experimental timeframe (24 h) impacting CFTR stability (Fig. 3, Supplementary Fig. S7). The possibility of a change in CFTR mRNA levels upon Sec13 silencing was examined by quantitative RT-qPCR (Supplementary Fig. S7B). There was no difference in CFTR mRNA levels in the Sec13 depleted samples compared to controls, suggesting that the upstream roles of Sec13 affecting chromatin architecture and transcription potentially impacting the level of the Sec13 transcript do not play a role in CFTR stabilization in the limited (24 h) time-frame of experiments.
Sec13 promotes CFTR rescue through managing its response to chaperones
Given the extensive cytoplasmic orientation of its non-transmembrane domains, CFTR normally interacts with the chaperone network found in the cytoplasm76,77,82,142,161,162. We have previously shown that chronic activation of the cytosolic heat shock response (HSR) in F508del expressing cells is counterproductive—referred to as the maladaptive stress response (MSR)161. In the augmented proteostasis environment in response to misfolding of F508del, the variant is retained in the ER by an increase in Hsp70 and Hsp90 chaperone pools forming a ‘chaperone trap’ that we have posited is attempting to resolve unwanted structural intermediates that are a challenge to the cytosolic folding machinery rendering it susceptible to degradation by ERAD89.
To address the potential role of cytosolic chaperones in response to Sec13 silencing, we compared the chaperone pools associated with either WT or F508del CFTR in siRNA control or siRNA Sec13 treated cells. CFTR was immunoprecipitated from control or Sec13 depleted HBE41o- and CFBE41o- cells, respectively, and the bound chaperone pools were analyzed using chaperone specific antibodies to the cytosolic Hsp40, Hsp70 and Hsp9082. As expected69,89, analysis of CFTR bound chaperone proteins in the control revealed that mutant F508del protein shows a stronger association with chaperones in comparison to WT CFTR (Fig. 4C). Upon Sec13 silencing, quantification of the bound chaperone proteins in the immunoblots relative to that of siScr control (set to a value of 1) showed a decreased association of Hsp40, Hsp70 and Hsp90 to both WT- and F508del, indicating that reduction of Sec13 levels decreases the interactions between CFTR and proteostasis components directing the maladaptive chaperone response161, and therefore limits interaction of CFTR in the ER with the chaperone trap89,161. Loss of this interaction impacts COPII/COPI exchange for targeting to degradation at the ERES-ERGIC interface9,20,51,57,77,78,89,90,155,156,161.
Protease digestion reveals increased CFTR fold stability in response to siSec13
Since Sec13 silencing reduced the targeting of both WT and F508del CFTR for degradation, we assessed the impact of Sec13 silencing on improving CFTR conformational stability. We followed CFTR stability using protease digestion74,75,77 to detect the conformation status of CFTR protein in response to reduced Sec13 proteins levels. Conformationally stable proteins generally have more compact three-dimensional structures than their variant counterparts, leading to enhanced resistance towards protease digestion. Using the NBD1 60 kDa domain specific antibody 3G11 (Fig. 4D, upper panels) and a polyclonal antibody (Mr. Pink) (Fig. 4D, lower panels)73,74,75,83,84,163,164, we observed the appearance of a more slowly migrating trypsin resistant bands in Sec13 depleted samples compared to siScr control, indicating an increase in conformation stability (Fig. 4D). For example, we observed the appearance of an increased level of a protease resistant band of 60 kDa, characteristic of a stabilized NBD1 domain, and 30 kDa stable region (Fig. 4D, left panel immunoblots, middle panel 60 kDa band quantitation, right panel 30 kDa band quantitation).These results suggests that siSec13 treatment leads to enhanced protease protection of more folded state of NBD1 (Fig. 4D) that, while insufficient to rescue the key YKDAD di-acidic code exit motif defective in the F508del variant for export to the surface3,4,7, helps to prevent delivery of CFTR to ERAD and/or COPII sensitive autophagic pathways at the COPII/COPI ERES-ERGIC interface122.
Discussion
Using CFTR as a model COPII dependent cargo we have defined the role of Sec13 to determine where in the hierarchy of its interactions it exerted its strongest and more immediate effects to understand its mechanism(s) of action in the cell. We screened a library of siRNAs silencing of each of COPII subunits and their isoforms to only reduce their expression. Complete silencing of Sec13 would likely be lethal to the cell as it would eliminate ER export for all proteins, consistent with results in yeast114,115,165. Surprisingly, we found that by reducing Sec13 protein levels, unlike any of the other COPII components such as Sec23/24 involved in cargo selection, we significantly improved both WT and F508del CFTR stability by reducing its degradation. Consistent with this conclusion, partial Sec13 silencing also resulted in a decrease in HSPs bound to CFTR, helping to avoid the chaperone trap89. Whereas WT CFTR showed improved accumulation as the cell surface band C glycoform, F508del largely accumulated as an ER localized band B in response to reduced levels of Sec13/31 cage assembly complexes, although we did detect a modest increase of band C and improved cell surface channel activity, reflecting the stabilizing and trafficking effect observed for WT CFTR. The stability increase in WT and F508del variant CFTR could reflect a contribution of both prevention of delivery to ERAD and/or autophagic/lysosomal including the mTOR pathways and potentially linked to nutrient responses166 managed by Sec13 at the ERES-ERGIC interface responsible for COPII-COPI exchange20,48,51,57,91,155. Moreover, it remains possible that non-conventional trafficking pathways from the ER could also be susceptible to increased activity in response to reduction of Sec13 levels16,166,167.
Intriguingly, the reduced level of Sec13 still allowed for delivery of WT CFTR to the cell surface demonstrating that the cargo selection step for export through the di-acidic code was complete and flow to cell surface is unimpeded following partial Sec13 silencing, suggesting that WT CFTR escapes the COPI dependent recycling step for degradation. Moreover, Sec13 sensitive post-Golgi endo-lysosome trafficking pathways sensitive to Sec13 function may also contribute to stability reflecting its more global role(s) in management of endomembrane composition. A similar result was observed for the ABC transporter family member ABCC4. These results are consistent with improved stability of the fold in response to protease digestion, suggesting that CFTR matures past early intermediate metastable forms in the presence of reduced Sec13/31 complex.
We now suggest that Sec13/31 cage assembly, globally managed by Sec13 levels in the cell, impacts the dynamics between COPII forward and COPI reverse recycling pathways at the ERES-ERGIC interface as originally suggested57 and later supported the efforts of many others20,48,51,57,91,155. The effect on different cargos will require further investigation given that neither AAT, a secreted protein, nor TRF, a plasma membrane recycling protein, were impacted by siRNA Sec13 in the time-frame used to capture impact on nascent synthesis of CFTR. These results potentially reflect the role of recycling cargo receptors such as LMAN1 and SURF4 for AAT18,46, or endosomal recycling pathways for TRF134. In addition, reduction of Sec13 may influence the activity of bypass pathways that we and others have reported51,166. We conclude an important role for Sec13 is in establishing a ‘set-point’ for differentiation between export and degradation with reduced levels of cage assembly favoring cargo stability associated with improved export through intrinsic export motifs such as the di-acidic code for CFTR (Fig. 5, asterisk).
Sec13 is a central regulator of early information flow in eukaryotic cells. Sec13 participates in a wide range of cellular functions related to initiation of information flow from the genome through the endomembrane pathway starting with chromatin condensation/de-condensation, modulation of transcription, RNA maturation and degradation, RNA/protein transport between nucleus and cytoplasm, translation of protein, protein trafficking and modulation of key signaling pathways affecting post-ER endomembrane pathways. Proteins associated with these functions were recovered in the Sec13 interactome. The combined results suggest that Sec13 provides pivotal framework around which endomembrane trafficking could be coordinated through a SCV set-point6,7 defined by Sec13 activity in each cell type in generating and mobilizing cargo that contributes as a collective to cellular endomembrane design and function as a quality system3,4.
While we focused on the role of Sec13 in COPII mediation of cargo traffic from the ER, from a more global perspective based on our Sec13 interactome hits and the response of the cell proteome to siSec13 based silencing, we now appreciate that Sec13 is involved in a large number of activities that could also influence the organization of information flow initiated by expression from the genome (Fig. 5, see summary Supplementary Fig. S1). While we did not detect a change in Sec13 mRNA levels during the short 24 h window leading to the improved stability of CFTR, the more global changes in Sec13’s physical association with chromatin, gene transcription factors affecting RNA maturation, RNA/protein transport from the nucleus to the cytoplasm, as well as translational and trafficking components, in response to Sec13 silencing highlights its potential for a much broader role in coordinating endomembrane structure to function relationships. One possibility is that Sec13’s WD40 propeller domain, a domain that is known to function in generating protein–protein interactions, plays a key role in coordinating information flow by as yet unanticipated mechanisms, to build as a collective endomembrane systems that are dependent on the genome encoded specialized cargo composition of different cell types49. This conclusion is consistent with our recent results where we have used a Gaussian process (GP) machine learning framework to describe the spatial covariance (SCV) features dictating sequence-to-function-to-structure relationships4 responsible for endomembrane trafficking of CFTR3,4,7 and other cargo1,5,6,145 on a residue-by-residue basis as an integrated quality system (QS) in each cell type in response to inherited genetic variation triggering human disease3,4. By being positioned at the root of the genome encoded exocytic and endocytic endomembrane system that are different for each cell type and highly responsive to their local environments, we suggest that Sec13 serves as an unanticipated key and dynamic regulatory node to achieve globally the genome to proteome transformation responsible for health and disease. Such flexibility at the first step in the exocytic pathway allows each cell type to (re)program SCV relationships in response to variation across the population to provide a precision response to natural selection responsible for survival and fitness1,2,3,4,5,6,7 (Fig. 5).
Materials and methods
Cell culture, transfection, and treatment with compounds
Cell culture
HBE41o- and CFBE41o- expressing WT and F508del CFTR, respectively, were obtained from Dr. J Clancy (University of Alabama). Cells were cultured in α-MEM containing 10% fetal bovine serum (FBS) and 2 mM l-glutamine. HBE41o- and CFBE41o- containing YFP-H148Q were engineered and cultured as previously described168. siRNA-mediated silencing. HBE41o- and CFBE41o- cells were plated in 12-well dishes and grown to 60–70% confluence. COPII subunit silencing was performed using 50 nm of validated siRNA for the indicated COPII subunit as per manufacturer’s directions. Cells were cultured for 5 h in the serum-free opti-MEM containing transfection complexes and subsequently washed and cultured for an addition of 48 h in the presence of growth medium prior to processing for immunoblot analysis. Treatment. Where indicated, cells were treated with MG132 (10 µM), chloroquine (50 µM) in complete growth media and incubated at 37 °C, 5% CO2 for 24 h. MG132 and chloroquine were procured from MedChemExpress.
Immunoblot and immunoprecipitation
Preparation of cell lysate and immunoblotting
As previously described3,4, cells were washed twice with ice-cold PBS, and 50 µl of lysis buffer (50 mM Tris–HCl, pH 7.4, 150 mM NaCl, 1% Triton X-100, and 2 mg/ml of complete protease inhibitor mixture) was added to each well. Cells were lysed on ice for 30 min with occasional mixing. The lysates were collected and spun at 14,000 × g for 20 min at 4 °C. The protein concentration in the supernatants was assessed by Bradford assay using the Coomassie Protein Assay Reagent (Pierce). 15 µg of total protein were separated by 7% SDS-PAGE, transferred to nitrocellulose, and incubated with specific primary antibody (Abcam) over-night at 4 °C in TBS + 0.1% Tween 20 + 5% milk. Nitrocellulose was subsequently incubated with the appropriate secondary antibody (Abcam) in TBS + 0.1%Tween 20 + 5% milk at room temperature for 1 h. Detection was performed by chemiluminescence using ECL reagent. Immunoprecipitation. Lysates were prepared as described for immunoblotting. 1.0 mg of total protein was pre-cleared by incubating with γ-bind beads (GE Healthcare) for 1 h at 4 °C and subsequently incubated for 16 h at 4 °C with 20 µl of either CFTR, ubiquitin or Sec13 antibody cross-linked to γ-bind beads (GE Healthcare). The monoclonal antibody to Sec13 was generated and characterized using purified Sec13 as described previously99. The beads were washed with two changes of lysis buffer without protease inhibitor mixture. Beads were pelleted at 2000 × g for 1 min, and bound proteins were eluted by incubation with 50 mM Tris–HCl, pH 6.8, containing 1% SDS at rt for 30 min and at 37 °C for 5 min. Immunoblots of eluted proteins were analyzed as described above. The immunoblots were developed using x-ray film- therefore, the markers do not appear on the blots. The membranes were cut (using Ponceau for protein visualization) before probing with primary antibodies allowing us to probe different regions of the same membrane for different proteins.
CFTR trypsin proteolysis
CFTR trypsin digestion has been performed as previously described161. Briefly, HBE41o- cells treated with siSec13 or control siScr, were first lysed for 30 min at 4 °C with lysis buffer (50 mM Tris–HCl pH 7.4, 150 mM NaCl, 1% Triton X-100,2 mg/ml Protease Inhibitor cocktail [Roche]) and harvested at 20,000 g for 20 min at 4 °C. Total protein concentration of pre-cleared lysates was determined by Bradford. Proteolysis was performed by incubating 80 mg of total protein with increasing concentration of Trypsin in PBS (0.05–0.35 mg/ml) at 4 °C for15 min. Proteolysis was stopped by adding 1 mM of PMSF and 6 × SDS-PAGE sample buffer. Samples were loaded onto 12% SDS-PAGE for separation of the proteolytic fragments and probed with CFTR antibodies for NBD1 (3G11: epitope 396–405).
q-RT-PCR
Quantitative Reverse Transcription PCR (qRT-PCR) was performed using the iScript One-Step RT-PCR kit with SYBR green (BioRad, Hercules, CA). RNA was standardized by quantification of beta-glucuronidase (GUS) mRNA, and all values were expressed relative to GUS. CFTR Forward (GTGGCTGCTTCTTTG-GTTGT) and reverse (CGAACTGCTGCTGGTGATAA) primers were used as indicated.
Functional YFP quenching assay
As described previously3, YFP expressing F508del-CFBE41o- and WT-CFBE41o- cells stably transfected with pcDNA3.1 containing YFP-H148Q were provided by Dr. L. Galietta, Telethon University for Genetics and Medicine, Puzzuoli, IT. The stable cell lines could be passed at least 15 times without a decrease in YFP-H148Q fluorescence. F508del-CFBE -YFP and WT-CFBE-YFP were cultured in the same media as the YFP non-expressing cells with addition 0.75 mg/ml G418. Cells were treated with indicated drug and siRNA. Following treatment, cells were stimulated with a final concentration of 10 μM Fsk and 50 μM Gen for 15 min prior to addition of PBS + NaI (replacement of NaCl with 137 mm NaI). Fluorescence was monitored each second for a total of 30 s (3 s prior to addition of NaI and 27 s after addition of NaI) using a Synergy H1 HybridReader (BioTek, Winooski, VT). As a negative control, identical experiments were performed with null -/- HBE41o- -YFP cells not expressing F508del CFTR.
Immunoprecipitation and sample preparation for Sec13 interactome
HBE41o- cell lysates were prepared as described for immunoblotting89,108,146. The lysates were pre-cleared using 50 μl of GammaBind Plus Sepharose beads for 1 h at 4 °C with mixing. The beads were pelleted at 500 × g for 5 min at 4 °C and the pre-cleared lysate was transferred to a new tube. The Sec13 immunoprecipitation was performed by adding the Sec13 antibody pre-crosslinked to GammaBind Plus Sepharose beads to the precleared lysates (4 mg total protein per IP). Control IPs were performed by using mock-IPs, in which no antibody is coupled to the beads, to identify bead- and cell-specific background. The lysates were incubated with or without the antibody overnight at 4 °C with end-over-end mixing. The beads were pelleted at 500 × g for 5 min at 4 °C and washed twice with 10 bead volumes of lysis buffer and twice with 10 bead volumes of lysis buffer without IGEPAL-CA630. Following immunoprecipitation washes with lysis buffer and 50 mM ammonium bicarbonate, proteins were digested directly on-beads as previously described169. Briefly, proteins bound to the beads were resuspended with 8 M urea and 50 mM ammonium bicarbonate, and Cys disulfide bonds were reduced with 10 mM Tris (2-carboxyethyl) phosphine at 30 °C for 60 min followed by cysteine alkylation with 30 mM iodoacetamide in the dark at room temperature for 30 min. Following alkylation, urea was diluted to 1 M urea using 50 mM ammonium bicarbonate, and proteins were finally subjected to overnight digestion with mass spec grade Trypsin/Lys-C mix (Promega, Madison, WI). Finally, beads were pulled down and the solution with peptides collected into a new tube. The beads were then washed once with 50 mM ammonium bicarbonate to increase peptide recovery. The digested samples were desalted using a C18 TopTip (PolyLC, Columbia, MD), and the organic solvent was removed in a SpeedVac concentrator prior to LC–MS/MS analysis.
Sample preparation and TMT labelling for proteome profiling
Protein extracts were obtained from HBE41o- cells treated with siSec13 or siScr (control). Cell lysis was performed as described above. TMT labelling was prepared as previously described89,108,146. 50 µg of a protein aliquot from each sample was acetone precipitated. The protein pellets were resolubilized in 100 mM TEAB supplemented with 0.1% SDS, reduced with TCEP for 1 h at 55 °C, and then alkylated with iodoacetamide for 30 min in the dark. The denatured proteins were digested with trypsin by overnight incubation at 37 °C. Resulting peptides were labelled with 6-plex Tandem Mass Tag (TMT) labelling reagent (Thermo-Fisher) according to the manufacturer’s recommendations. Subsequently, insoluble precipitates were removed by centrifugation (15 min, 18,000 g) and samples reduced to near dryness in vacuo.
2D LC MS/MS analysis
2D LC MS/MS analysis was performed as previously described89,108,146. Briefly, dried samples were reconstituted in 100 mM ammonium formate (pH ~ 10) and analyzed by 2D LC–MS/MS using a 2D nanoACQUITY Ultra Performance Liquid Chromatography system (Waters Corp., Milford, MA) coupled to a Q-Exactive Plus mass spectrometer (Thermo Fisher Scientific). Peptides were loaded onto the first-dimension column, XBridge BEH130 C18 NanoEase (300 μm × 50 mm, 5 μm) equilibrated with solvent A [20 mM ammonium formate (pH 10), first dimension pump] at 2 μl/min. The first fraction was eluted from the first-dimension column at 17% of solvent B (100% acetonitrile) for 4 min and transferred to the second dimension Symmetry C18 trap column 0.180 × 20 mm (Waters Corp., Milford, MA) using a 1:10 dilution with 99.9% second dimensional pump solvent A (0.1% formic acid in water) at 20 μl/min. Peptides were then eluted from the trap column and resolved on the analytical C18 BEH130 PicoChip column 0.075 × 100-mm, 1.7-μm particles (NewObjective, MA) at low pH by increasing the composition of solvent B (100% acetonitrile) from 2 to 26% over 94 min at 400 nl/min. Subsequent fractions were carried with increasing concentrations of solvent B. The following 4 first dimension fractions were eluted at 19.5%, 22%, 26% and 65% solvent B. The mass spectrometer was operated in positive data-dependent acquisition mode. MS1 spectra were measured with a resolution of 70,000, an AGC target of 1,000,000, and a mass range from 350 to 1700 m/z. Up to 12 MS2 spectra per duty cycle were triggered, fragmented by HCD, and acquired with a resolution of 17,500 and an AGC target of 5e4, an isolation window of 2.0 m/z and a normalized collision energy of 25. Dynamic exclusion was enabled with duration of 20 s.
Data analysis
As previously describe89,108,146, mass spectra were analyzed with MaxQuant software version 1.5.5.1. MS/MS spectra were searched against the Homo sapiens Uniprot protein sequence database (version July 2016) and GPM cRAP sequences (commonly known protein contaminants). Precursor mass tolerance was set to 20 and 4.5 ppm for the first search where initial mass recalibration was completed and for the main search, respectively. Product ions were searched with a mass tolerance 0.5 Da. The maximum precursor ion charge state used for searching was 7. Carbamidomethylation of cysteines was searched as a fixed modification, while oxidation of methionines and acetylation of protein N-terminal were searched as variable modifications. Enzyme was set to trypsin in a specific mode and a maximum of two missed cleavages were allowed for searching. The target-decoy-based false discovery rate filter for spectrum and protein identification was set to 1%. As previously described170, normalized spectral abundance factor (NSAF) values were calculated for proteins in each sample to account for protein size and variability between runs. The NSAF for a protein p is the number of spectral counts (SpC, the total number of MS/MS spectra) identifying a protein, p, divided by the protein length (L), divided by the sum of SpC/L for all N proteins in the experimental design. The data sets were statistically compared to determine the significance of the change between the two groups using Student’s t test (two tailed unpaired t test). To determine the relative abundance of interacting proteins on Sec13 cross-linked Gamma Bind Plus Sepharose beads relative to the Gamma Bind Plus Sepharose beads alone, the data sets were first filtered to include only those proteins that were detected in all three replicates for each condition. Then the ratio of the mean of the NSAF values from three biological replicates of Sec13 immunoprecipitates to the mean of NSAF values from three biological replicates of the control was computed to calculate the fold-change.
Data availability
All data generated or analyzed during this study are included in the published article and its supplementary information file.
References
Sun, S. et al. Capturing the conversion of the pathogenic alpha-1-antitrypsin fold by ATF6 enhanced proteostasis. Cell Chem. Biol. 30, 22–42 (2023).
Loguercio, S. et al. Understanding the host-pathogen evolutionary balance through Gaussian process modeling of SARS-CoV-2. Patterns N. Y. 4, 100800 (2023).
Wang, C., Angles, F. & Balch, W. E. Triangulating variation in the population to define mechanisms for precision management of genetic disease. Structure 30, 1190–1207 (2022).
Anglès, F., Wang, C. & Balch, W. E. Spatial covariance analysis reveals the residue-by-residue thermodynamic contribution of variation to the CFTR fold. Commun. Biol. 5, 1–16 (2022).
Wang, C. et al. Individualized management of genetic diversity in Niemann-Pick C1 through modulation of the Hsp70 chaperone system. Hum. Mol. Genet. 29, 1–19 (2020).
Wang, C. et al. Quantitating the epigenetic transformation contributing to cholesterol homeostasis using Gaussian process. Nat. Commun. 10, 5052 (2019).
Wang, C. & Balch, W. E. Bridging Genomics to Phenomics at Atomic Resolution through Variation Spatial Profiling. Cell Rep. 24, 2013–2028 (2018).
Stagg, S. M. et al. Structural basis for cargo regulation of COPII coat assembly. Cell 134, 474–484 (2008).
Gürkan, C., Stagg, S. M., LaPointe, P. & Balch, W. E. The COPII cage: Unifying principles of vesicle coat assembly. Nat. Rev. Mol. Cell Biol. 7, 727–738 (2006).
Aridor, M. A tango for coats and membranes: New insights into ER-to-Golgi traffic. Cell Rep. 38, 110258 (2022).
Chatterjee, S., Choi, A. J. & Frankel, G. A systematic review of Sec24 cargo interactome. Traffic 22, 412–424 (2021).
Lu, C. L. & Kim, J. Consequences of mutations in the genes of the ER export machinery COPII in vertebrates. Cell Stress Chaperones 25, 199–209 (2020).
Li, Z., Huang, W. & Wang, W. Multifaceted roles of COPII subunits in autophagy. Biochim. Biophys. Acta Mol. Cell Res. 1867, 118627 (2020).
Pantazopoulou, A. & Glick, B. S. A kinetic view of membrane traffic pathways can transcend the classical view of golgi compartments. Front. Cell Dev. Biol. 7, 153 (2019).
McCaughey, J. & Stephens, D. J. COPII-dependent ER export in animal cells: Adaptation and control for diverse cargo. Histochem. Cell Biol. 150, 119–131 (2018).
Van der Verren, S. E. & Zanetti, G. The small GTPase Sar1, control centre of COPII trafficking. FEBS Lett. 597, 865–882 (2023).
Tang, V. T. & Ginsburg, D. Cargo selection in endoplasmic reticulum-to-Golgi transport and relevant diseases. J. Clin. Invest. 133, 1 (2023).
Shen, Y., Gu, H. M., Qin, S. & Zhang, D. W. Surf4, cargo trafficking, lipid metabolism, and therapeutic implications. J. Mol. Cell Biol. 2023, 14 (2023).
Raote, I., Saxena, S. & Malhotra, V. Sorting and export of proteins at the endoplasmic reticulum. Cold Spring Harb. Perspect. Biol. 15, 823 (2023).
Ogen-Shtern, N. et al. COP I and II dependent trafficking controls ER-associated degradation in mammalian cells. iScience 26, 106232 (2023).
Parashar, S. & Ferro-Novick, S. Architecture of the endoplasmic reticulum plays a role in proteostasis. Autophagy 18, 937–938 (2022).
Malis, Y., Hirschberg, K. & Kaether, C. Hanging the coat on a collar: Same function but different localization and mechanism for COPII. Bioessays 44, e2200064 (2022).
Li, X., Li, X., Fan, B., Zhu, C. & Chen, Z. Specialized endoplasmic reticulum-derived vesicles in plants: Functional diversity, evolution, and biotechnological exploitation. J. Integr. Plant Biol. 64, 821–835 (2022).
Wang, L. et al. WFS1 functions in ER export of vesicular cargo proteins in pancreatic beta-cells. Nat. Commun. 12, 6996 (2021).
Shomron, O. et al. COPII collar defines the boundary between ER and ER exit site and does not coat cargo containers. J. Cell Biol. 220, 87 (2021).
Raote, I. & Malhotra, V. Tunnels for protein export from the endoplasmic reticulum. Annu. Rev. Biochem. 90, 605–630 (2021).
Joiner, A. M. N. & Fromme, J. C. Structural basis for the initiation of COPII vesicle biogenesis. Structure 29, 859–872 (2021).
Duan, J. & Lambright, D. G. Propelling COPII vesicle biogenesis at the endoplasmic reticulum. Structure 29, 779–781 (2021).
Stancheva, V. G. et al. Combinatorial multivalent interactions drive cooperative assembly of the COPII coat. J. Cell Biol. 219, 85 (2020).
Gomez-Navarro, N. et al. Cargo crowding contributes to sorting stringency in COPII vesicles. J. Cell Biol. 219, 89 (2020).
Cui, Y., Parashar, S. & Ferro-Novick, S. A new role for a COPII cargo adaptor in autophagy. Autophagy 16, 376–378 (2020).
Barlowe, C. Twenty-five years after coat protein complex II. Mol. Biol. Cell 31, 3–6 (2020).
Stagg, S. M. et al. Structure of the Sec13/31 COPII coat cage. Nature 439, 234–238 (2006).
Wang, X. et al. COPII-dependent export of cystic fibrosis transmembrane conductance regulator from the ER uses a di-acidic exit code. J. Cell Biol. 167, 65–74 (2004).
Nie, C., Wang, H., Wang, R., Ginsburg, D. & Chen, X. W. Dimeric sorting code for concentrative cargo selection by the COPII coat. Proc. Natl. Acad. Sci. U. S. A. 115, E3155–E3162 (2018).
Khoriaty, R. et al. Functions of the COPII gene paralogs SEC23A and SEC23B are interchangeable in vivo. Proc. Natl. Acad. Sci. U. S. A. 115, E7748–E7757 (2018).
Paraan, M., Bhattacharya, N., Uversky, V. N. & Stagg, S. M. Flexibility of the Sec13/31 cage is influenced by the Sec31 C-terminal disordered domain. J. Struct. Biol. 204, 250–260 (2018).
Parashar, S. et al. Endoplasmic reticulum tubules limit the size of misfolded protein condensates. Elife 10, 123 (2021).
Cui, Y. et al. A COPII subunit acts with an autophagy receptor to target endoplasmic reticulum for degradation. Science 365, 53–60 (2019).
Farhan, H., Kundu, M. & Ferro-Novick, S. The link between autophagy and secretion: A story of multitasking proteins. Mol. Biol. Cell 28, 1161–1164 (2017).
Nishimura, N. & Balch, W. E. A di-acidic signal required for selective export from the endoplasmic reticulum. Science 277, 556–558 (1997).
Saha, K. et al. Pharmacological chaperone-rescued cystic fibrosis CFTR-F508del mutant overcomes PRAF2-gated access to endoplasmic reticulum exit sites. Cell Mol. Life Sci. 79, 530 (2022).
Lagunas-Gomez, D., Yanez-Dominguez, C., Zavala-Padilla, G., Barlowe, C. & Pantoja, O. The C-terminus of the cargo receptor Erv14 affects COPII vesicle formation and cargo delivery. J. Cell Sci. 136, 796 (2023).
Tempio, T. et al. A virtuous cycle operated by ERp44 and ERGIC-53 guarantees proteostasis in the early secretory compartment. iScience 24, 102244 (2021).
Sargeant, J. et al. ALG-2 and peflin regulate COPII targeting and secretion in response to calcium signaling. J. Biol. Chem. 297, 101393 (2021).
Ordonez, A., Harding, H. P., Marciniak, S. J. & Ron, D. Cargo receptor-assisted endoplasmic reticulum export of pathogenic alpha1-antitrypsin polymers. Cell Rep. 35, 109144 (2021).
Nalbach, K. et al. Spatial proteomics reveals secretory pathway disturbances caused by neuropathy-associated TECPR2. Nat. Commun. 14, 870 (2023).
Aridor, M. et al. The Sar1 GTPase coordinates biosynthetic cargo selection with endoplasmic reticulum export site assembly. J. Cell Biol. 152, 213–229 (2001).
Aridor, M., Bannykh, S. I., Rowe, T. & Balch, W. E. Cargo can modulate COPII vesicle formation from the endoplasmic reticulum. J. Biol. Chem. 274, 4389–4399 (1999).
Aridor, M., Weissman, J., Bannykh, S., Nuoffer, C. & Balch, W. E. Cargo selection by the COPII budding machinery during export from the ER. J. Cell Biol. 141, 61–70 (1998).
Aridor, M., Bannykh, S. I., Rowe, T. & Balch, W. E. Sequential coupling between COPII and COPI vesicle coats in endoplasmic reticulum to Golgi transport. J. Cell Biol. 131, 875–893 (1995).
Hutchings, J., Stancheva, V., Miller, E. A. & Zanetti, G. Subtomogram averaging of COPII assemblies reveals how coat organization dictates membrane shape. Nat. Commun. 9, 4154 (2018).
Mehrani, A. & Stagg, S. M. Probing intracellular vesicle trafficking and membrane remodelling by cryo-EM. J. Struct. Biol. 214, 107836 (2022).
Bhattacharya, N. J. O. D. & Stagg, S. M. The structure of the Sec13/31 COPII cage bound to Sec23. J. Mol. Biol. 420, 324–334 (2012).
O’Donnell, J., Maddox, K. & Stagg, S. The structure of a COPII tubule. J. Struct. Biol. 173, 358–364 (2011).
Russell, C. & Stagg, S. M. New insights into the structural mechanisms of the COPII coat. Traffic 11, 303–310 (2010).
Rowe, T. et al. COPII vesicles derived from mammalian endoplasmic reticulum microsomes recruit COPI. J. Cell Biol. 135, 895–911 (1996).
Xu, F. et al. COPII mitigates ER stress by promoting formation of ER whorls. Cell Res. 31, 141–156 (2021).
Phuyal, S. & Farhan, H. Want to leave the ER? We offer vesicles, tubules, and tunnels. J. Cell Biol. 220, 6 (2021).
Newstead, S. & Barr, F. Molecular basis for KDEL-mediated retrieval of escaped ER-resident proteins - SWEET talking the COPs. J. Cell Sci. 133, 19 (2020).
Peotter, J., Kasberg, W., Pustova, I. & Audhya, A. COPII-mediated trafficking at the ER/ERGIC interface. Traffic 20, 491–503 (2019).
Bethune, J. & Wieland, F. T. Assembly of COPI and COPII vesicular coat proteins on membranes. Annu. Rev. Biophys. 47, 63–83 (2018).
Niu, X. et al. The nuclear pore complex function of Sec13 protein is required for cell survival during retinal development. J. Biol. Chem. 289, 11971–11985 (2014).
Lin, D. H. & Hoelz, A. The structure of the nuclear pore complex (an update). Annu. Rev. Biochem. 88, 725–783 (2019).
Bar-Peled, L. et al. A Tumor suppressor complex with GAP activity for the Rag GTPases that signal amino acid sufficiency to mTORC1. Science 340, 1100–1106 (2013).
Parmigiani, A. et al. Sestrins inhibit mTORC1 kinase activation through the GATOR complex. Cell Rep. 9, 1281–1291 (2014).
Valenstein, M. L. et al. Structure of the nutrient-sensing hub GATOR2. Nature 607, 610–616 (2022).
Wang, X., Koulov, A. V., Kellner, W. A., Riordan, J. R. & Balch, W. E. Chemical and biological folding contribute to temperature-sensitive DeltaF508 CFTR trafficking. Traffic 9, 1878–1893 (2008).
Wang, X. et al. Hsp90 cochaperone Aha1 downregulation rescues misfolding of CFTR in cystic fibrosis. Cell 127, 803–815 (2006).
Nishimura, N. et al. A di-acidic (DXE) code directs concentration of cargo during export from the endoplasmic reticulum. J. Biol. Chem. 274, 15937–15946 (1999).
Fiedorczuk, K. & Chen, J. Mechanism of CFTR correction by type I folding correctors. Cell 185, 158–168 (2022).
Fiedorczuk, K. & Chen, J. Molecular structures reveal synergistic rescue of Delta508 CFTR by Trikafta modulators. Science 378, 284–290 (2022).
Im, J. et al. ABC-transporter CFTR folds with high fidelity through a modular, stepwise pathway. Cell Mol. Life Sci. 80, 33 (2023).
Hwang, T. C., Braakman, I., van der Sluijs, P. & Callebaut, I. Structure basis of CFTR folding, function and pharmacology. J. Cyst. Fibros 22(Suppl 1), S5–S11 (2023).
Kleizen, B. et al. Co-translational folding of the first transmembrane domain of ABC-transporter CFTR is supported by assembly with the first cytosolic domain. J. Mol. Biol. 433, 166955 (2021).
Baaklini, I., Goncalves, C. C., Lukacs, G. L. & Young, J. C. Selective binding of HSC70 and its co-chaperones to structural hotspots on CFTR. Sci. Rep. 10, 4176 (2020).
Bagdany, M. et al. Chaperones rescue the energetic landscape of mutant CFTR at single molecule and in cell. Nat. Commun. 8, 398 (2017).
Amaral, M. D. & Balch, W. E. Hallmarks of therapeutic management of the cystic fibrosis functional landscape. J. Cystic Fibrosis 14, 687–699 (2015).
Shishido, H., Yoon, J. S., Yang, Z. & Skach, W. R. CFTR trafficking mutations disrupt cotranslational protein folding by targeting biosynthetic intermediates. Nat. Commun. 11, 4258 (2020).
Santos, J. D. et al. Folding status is determinant over traffic-competence in defining CFTR interactors in the endoplasmic reticulum. Cells 8, 353 (2019).
Schwarzenberg, S. J. et al. Elexacaftor/tezacaftor/ivacaftor and gastrointestinal outcomes in cystic fibrosis: Report of promise-GI. J. Cyst. Fibros 22, 282–289 (2023).
Hutt, D. M. et al. Silencing of the Hsp70-specific nucleotide-exchange factor BAG3 corrects the F508del-CFTR variant by restoring autophagy. J. Biol. Chem. 293, 13682–13695 (2018).
Hillenaar, T., Beekman, J., van der Sluijs, P. & Braakman, I. Redefining Hypo- and Hyper-Responding Phenotypes of CFTR Mutants for Understanding and Therapy. Int. J. Mol. Sci. 23, 145 (2022).
Sabusap, C. M. et al. The CFTR P67L variant reveals a key role for N-terminal lasso helices in channel folding, maturation, and pharmacologic rescue. J. Biol. Chem. 296, 100598 (2021).
van Willigen, M. et al. Folding-function relationship of the most common cystic fibrosis-causing CFTR conductance mutants. Life Sci. Alliance 2019, 2 (2019).
Hoelen, H. et al. The primary folding defect and rescue of ΔF508 CFTR emerge during translation of the mutant domain. PLoS One 5, e15458 (2010).
Pankow, S. et al. ∆ F508 CFTR interactome remodelling promotes rescue of cystic fibrosis. Nature 528, 510–516 (2015).
Hutt, D. M. et al. FK506 binding protein 8 peptidylprolyl isomerase activity manages a late stage of cystic fibrosis transmembrane conductance regulator (CFTR) folding and stability. J. Biol. Chem. 287, 21914–21925 (2012).
Coppinger, J. A. et al. A chaperone trap contributes to the onset of cystic fibrosis. PLoS One 7, e37682 (2012).
Koulov, A. V. et al. Biological and structural basis for Aha1 regulation of Hsp90 ATPase activity in maintaining proteostasis in the human disease cystic fibrosis. Mol. Biol. Cell 21, 871–884 (2010).
Bannykh, S. I., Rowe, T. & Balch, W. E. The organization of endoplasmic reticulum export complexes. J. Cell Biol. 135, 19–35 (1996).
Bi, X., Mancias, J. D. & Goldberg, J. Insights into COPII coat nucleation from the structure of Sec23.Sar1 complexed with the active fragment of Sec31. Dev. Cell 13, 635–645 (2007).
Fath, S., Mancias, J. D., Bi, X. & Goldberg, J. Structure and organization of coat proteins in the COPII cage. Cell 129, 1325–1336 (2007).
Brohawn, S. G., Leksa, N. C., Spear, E. D., Rajashankar, K. R. & Schwartz, T. U. Structural evidence for common ancestry of the nuclear pore complex and vesicle coats. Science 322, 1369–1373 (2008).
Enninga, J., Levay, A. & Fontoura, B. M. Sec13 shuttles between the nucleus and the cytoplasm and stably interacts with Nup96 at the nuclear pore complex. Mol. Cell. Biol. 23, 7271–7284 (2003).
Hsia, K.-C., Stavropoulos, P., Blobel, G. & Hoelz, A. Architecture of a coat for the nuclear pore membrane. Cell 131, 1313–1326 (2007).
Zhu, X. et al. Structure of the cytoplasmic ring of the Xenopus laevis nuclear pore complex. Science 376, 8280 (2022).
Capelson, M. et al. Chromatin-bound nuclear pore components regulate gene expression in higher eukaryotes. Cell 140, 372–383 (2010).
Gurkan, C. & Balch, W. E. Recombinant production in baculovirus-infected insect cells and purification of the mammalian Sec13/Sec31 complex. Methods Enzymol. 404, 58–66 (2005).
Jain, B. P. & Pandey, S. WD40 repeat proteins: Signalling scaffold with diverse functions. Protein J. 37, 391–406 (2018).
Xu, C. & Min, J. Structure and function of WD40 domain proteins. Protein Cell 2, 202–214 (2011).
Stirnimann, C. U., Petsalaki, E., Russell, R. B. & Müller, C. W. WD40 proteins propel cellular networks. Trends Biochem. Sci. 35, 565–574 (2010).
Szklarczyk, D. et al. STRING v11: Protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47, D607–D613 (2019).
Pavitt, G. D. Regulation of translation initiation factor eIF2B at the hub of the integrated stress response. Wiley Interdiscipl. Rev. RNA 9, e1491 (2018).
Gamper, A. M., Kim, J. & Roeder, R. G. The STAGA subunit ADA2b is an important regulator of human GCN5 catalysis. Mol. Cell Biol. 29, 266–280 (2009).
Zhao, Y. et al. A TFTC/STAGA module mediates histone H2A and H2B deubiquitination, coactivates nuclear receptors, and counteracts heterochromatin silencing. Mol. Cell 29, 92–101 (2008).
Gamper, A. M. & Roeder, R. G. Multivalent binding of p53 to the STAGA complex mediates coactivator recruitment after UV damage. Mol. Cell Biol. 28, 2517–2527 (2008).
Subramanian, K., Rauniyar, N., Lavalleé-Adam, M., Yates, J. R. III. & Balch, W. E. Quantitative analysis of the proteome response to the histone deacetylase inhibitor (HDACi) vorinostat in Niemann-Pick Type C1 disease. Mol. Cell. Proteom. 16, 1938–1957 (2017).
Li, S., Zhang, M. & Ge, L. A new type of membrane contact in the ER-Golgi system regulates autophagosome biogenesis. Autophagy 17, 4499–4501 (2021).
Ahsan, A. et al. Natural compounds modulate the autophagy with potential implication of stroke. Acta Pharm. Sin. B 11, 1708–1720 (2021).
Yamamoto, Y. H. & Noda, T. Autophagosome formation in relation to the endoplasmic reticulum. J. Biomed. Sci. 27, 97 (2020).
Shima, T., Kirisako, H. & Nakatogawa, H. COPII vesicles contribute to autophagosomal membranes. J. Cell Biol. 218, 1503–1510 (2019).
Zahoor, M. & Farhan, H. Crosstalk of autophagy and the secretory pathway and its role in diseases. Int. Rev. Cell Mol. Biol. 337, 153–184 (2018).
Salama, N. R., Chuang, J. S. & Schekman, R. W. Sec31 encodes an essential component of the COPII coat required for transport vesicle budding from the endoplasmic reticulum. Mol. Biol. Cell 8, 205–217 (1997).
Pryer, N. K., Salama, N. R., Schekman, R. & Kaiser, C. A. Cytosolic Sec13p complex is required for vesicle formation from the endoplasmic reticulum in vitro. J. Cell Biol. 120, 865–875 (1993).
Nguyen, J. P., Kim, Y., Cao, Q. & Hirota, J. A. Interactions between ABCC4/MRP4 and ABCC7/CFTR in human airway epithelial cells in lung health and disease. Int. J. Biochem. Cell Biol. 133, 105936 (2021).
Russel, F. G., Koenderink, J. B. & Masereeuw, R. Multidrug resistance protein 4 (MRP4/ABCC4): A versatile efflux transporter for drugs and signalling molecules. Trends Pharmacol. Sci. 29, 200–207 (2008).
Banting, G. & Ponnambalam, S. TGN38 and its orthologues: Roles in post-TGN vesicle formation and maintenance of TGN morphology. Biochim. Biophys. Acta BBA Mol. Cell Res. 1355, 209–217 (1997).
Pfeffer, S. R. Multiple routes of protein transport from endosomes to the trans Golgi network. FEBS Lett. 583, 3811–3816 (2009).
Jing, J., Wang, B. & Liu, P. The functional role of SEC23 in vesicle transportation, autophagy and cancer. Int. J. Biol. Sci. 15, 2419–2426 (2019).
Mauthe, M. et al. Chloroquine inhibits autophagic flux by decreasing autophagosome-lysosome fusion. Autophagy 14, 1435–1455 (2018).
He, L. et al. DNAJB12 and Hsp70 triage arrested intermediates of N1303K-CFTR for endoplasmic reticulum-associated autophagy. Mol. Biol. Cell 32, 538–553 (2021).
Li, C., Xia, B., Wang, S. & Xu, J. Folded or degraded in endoplasmic reticulum. Adv. Exp. Med. Biol. 1248, 265–294 (2020).
Poothong, J., Jang, I. & Kaufman, R. J. Defects in protein folding and/or quality control cause protein aggregation in the endoplasmic reticulum. Prog. Mol. Subcell Biol. 59, 115–143 (2021).
Estabrooks, S. & Brodsky, J. L. Regulation of CFTR biogenesis by the proteostatic network and pharmacological modulators. Int. J. Mol. Sci. 21, 452 (2020).
Estabrooks, S. K. & Brodsky, J. L. Ubiquitination of disease-causing CFTR variants in a microsome-based assay. Anal. Biochem. 604, 113829 (2020).
Goeckeler-Fried, J. L. et al. Improved correction of F508del-CFTR biogenesis with a folding facilitator and an inhibitor of protein ubiquitination. Bioorg. Med. Chem. Lett. 48, 128243 (2021).
Hou, X. et al. Dissection of the role of VIMP in endoplasmic reticulum-associated degradation of CFTRDeltaF508. Sci. Rep. 8, 4764 (2018).
Huang, Y. et al. Targeting DNAJB9, a novel ER luminal co-chaperone, to rescue DeltaF508-CFTR. Sci. Rep. 9, 9808 (2019).
Suzuki, K., Akioka, M., Kondo-Kakuta, C., Yamamoto, H. & Ohsumi, Y. Fine mapping of autophagy-related proteins during autophagosome formation in Saccharomyces cerevisiae. J. Cell Sci. 126, 2534–2544 (2013).
Saegusa, K. et al. Cargo receptor Surf4 regulates endoplasmic reticulum export of proinsulin in pancreatic beta-cells. Commun. Biol. 5, 458 (2022).
Ginsberg, H. N. ApoB SURFs a Ride from the ER to the Golgi. Cell Metab. 33, 231–233 (2021).
Saegusa, K., Sato, M., Morooka, N., Hara, T. & Sato, K. SFT-4/Surf4 control ER export of soluble cargo proteins and participate in ER exit site organization. J. Cell Biol. 217, 2073–2085 (2018).
Richard, C. & Verdier, F. Transferrin receptors in erythropoiesis. Int. J. Mol. Sci. 21, 9713 (2020).
Gammella, E., Buratti, P., Cairo, G. & Recalcati, S. The transferrin receptor: The cellular iron gate. Metallomics 9, 1367–1375 (2017).
Qu, B.-H., Strickland, E. H. & Thomas, P. J. Localization and suppression of a kinetic defect in cystic fibrosis transmembrane conductance regulator folding. J. Biol. Chem. 272, 15739–15744 (1997).
Sosnay, P. R. et al. Defining the disease liability of variants in the cystic fibrosis transmembrane conductance regulator gene. Nat. Genet. 45, 1160–1167 (2013).
Joynt, A. T., Cutting, G. R. & Sharma, N. Genetics of cystic fibrosis: Clinical implications. Clin. Chest Med. 43, 591–602 (2022).
Cutting, G. R. Cystic fibrosis genetics: From molecular understanding to clinical application. Nat. Rev. Genet. 16, 45–56 (2015).
Ahner, A., Nakatsukasa, K., Zhang, H., Frizzell, R. A. & Brodsky, J. L. Small heat-shock proteins select ΔF508-CFTR for endoplasmic reticulum-associated degradation. Mol. Biol. Cell 18, 806–814 (2007).
Brodsky, J. L. & Wojcikiewicz, R. J. Substrate-specific mediators of ER associated degradation (ERAD). Curr. Opin. Cell Biol. 21, 516–521 (2009).
Gong, X. et al. Non-native conformers of cystic fibrosis transmembrane conductance regulator NBD1 are recognized by Hsp27 and conjugated to SUMO-2 for degradation. J. Biol. Chem. 291, 2004–2017 (2016).
Turnbull, E. L., Rosser, M. F. & Cyr, D. M. The role of the UPS in cystic fibrosis. BMC Biochem. 8, 1–10 (2007).
Vembar, S. S. & Brodsky, J. L. One step at a time: Endoplasmic reticulum-associated degradation. Nat. Rev. Mol. Cell Biol. 9, 944–957 (2008).
Subramanian, K. et al. Correction of Niemann-Pick type C1 trafficking and activity with the histone deacetylase inhibitor valproic acid. J. Biol. Chem. 295, 8017–8035 (2020).
Pankow, S., Bamberger, C. & Yates, J. R. A posttranslational modification code for CFTR maturation is altered in cystic fibrosis. Sci. Signal. 12, 45 (2019).
Badr, A. et al. CFTR modulators restore acidification of autophago-lysosomes and bacterial clearance in cystic fibrosis macrophages. Front. Cell Infect. Microbiol. 12, 819554 (2022).
Kapouni, N., Moustaki, M., Douros, K. & Loukou, I. Efficacy and safety of elexacaftor-tezacaftor-ivacaftor in the treatment of cystic fibrosis: A systematic review. Children Basel 10, 745 (2023).
Bacalhau, M. et al. Elexacaftor-tezacaftor-ivacaftor: A life-changing triple combination of CFTR modulator drugs for cystic fibrosis. Pharmaceut. Basel 16, 78 (2023).
Ensinck, M. M. & Carlon, M. S. One size does not fit all: The past, present and future of cystic fibrosis causal therapies. Cells 11, 14 (2022).
Dawood, S. N. et al. Newly discovered cutting-edge triple combination cystic fibrosis therapy: A systematic review. Cureus 14, e29359 (2022).
Zaher, A., ElSaygh, J., Elsori, D., ElSaygh, H. & Sanni, A. A review of Trikafta: Triple cystic fibrosis transmembrane conductance regulator (CFTR) modulator therapy. Cureus 13, e16144 (2021).
Galietta, L. V., Jayaraman, S. & Verkman, A. Cell-based assay for high-throughput quantitative screening of CFTR chloride transport agonists. Am. J. Physiol.-Cell Physiol. 281, C1734–C1742 (2001).
Singh, J. K. et al. Management of Hsp90-dependent protein folding by small molecules targeting the Aha1 Co-Chaperone. Cell Chem. Biol. 27, 292–305 (2020).
Bannykh, S. I. & Balch, W. E. Membrane dynamics at the endoplasmic reticulum-Golgi interface. J. Cell Biol. 138, 1–4 (1997).
Peter, F., Plutner, H., Zhu, H., Kreis, T. E. & Balch, W. E. Beta-COP is essential for transport of protein from the endoplasmic reticulum to the Golgi in vitro. J. Cell Biol. 122, 1155–1167 (1993).
Wu, Y. et al. Sec16A, a key protein in COPII vesicle formation, regulates the stability and localization of the novel ubiquitin ligase RNF183. PLoS One 13, e0190407 (2018).
Cao, Q. et al. PAQR3 regulates endoplasmic reticulum-to-golgi trafficking of COPII vesicle via interaction with Sec13/Sec31 coat proteins. iScience 9, 382–398 (2018).
Dascher, C. & Balch, W. E. Dominant inhibitory mutants of ARF1 block endoplasmic reticulum to Golgi transport and trigger disassembly of the Golgi apparatus. J. Biol. Chem. 269, 1437–1448 (1994).
Ward, C. L. & Kopito, R. R. Intracellular turnover of cystic fibrosis transmembrane conductance regulator. Inefficient processing and rapid degradation of wild-type and mutant proteins. J. Biol. Chem. 269, 25710–25718 (1994).
Roth, D. M. et al. Modulation of the maladaptive stress response to manage diseases of protein folding. PLoS Biol. 12, e1001998 (2014).
Veit, G. et al. A precision medicine approach to optimize modulator therapy for rare CFTR folding mutants. J. Pers. Med. 11, 673 (2021).
Kleizen, B. et al. Transmembrane Helices 7 and 8 confer aggregation sensitivity to the cystic fibrosis transmembrane conductance regulator. Int. J. Mol. Sci. 24, 15741 (2023).
McCaul, N. & Braakman, I. Hold the fold: How delayed folding aids protein secretion. EMBO J. 41, e112787 (2022).
Antonny, B., Gounon, P., Schekman, R. & Orci, L. Self-assembly of minimal COPII cages. EMBO Rep. 4, 419–424 (2003).
Gee, H. Y., Noh, S. H., Tang, B. L., Kim, K. H. & Lee, M. G. Rescue of DeltaF508-CFTR trafficking via a GRASP-dependent unconventional secretion pathway. Cell 146, 746–760 (2011).
Cruz-Garcia, D., Malhotra, V. & Curwin, A. J. Unconventional protein secretion triggered by nutrient starvation. Semin. Cell Dev. Biol. 83, 22–28 (2018).
Anglès, F., Hutt, D. M. & Balch, W. E. HDAC inhibitors rescue multiple disease-causing CFTR variants. Hum. Mol. Genet. 28, 1982–2000 (2019).
Hutt, D. M., Loguercio, S., Campos, A. R. & Balch, W. E. A proteomic variant approach (ProVarA) for personalized medicine of inherited and somatic disease. J. Mol. Biol. 430, 2951–2973 (2018).
Zhu, W., Smith, J. W. & Huang, C.-M. Mass spectrometry-based label-free quantitative proteomics. J. Biomed. Biotechnol. 2010, 6 (2009).
Oliver, K. E. et al. Slowing ribosome velocity restores folding and function of mutant CFTR. J. Clin. Invest. 129, 5236–5253 (2019).
Acknowledgements
Grant support provided by NIH HL095524, DK051870, AG070209, AG049665, HL141810, HL166410 and HG10881 to WEB. FA and CW were recipients of Cystic Fibrosis Foundation (CFF) Fellowships. We acknowledge the expertise of Dr. Junyan Hu in helping to finalize Supplementary data for publication.
Author information
Authors and Affiliations
Contributions
WB led the project. WB, FA, VG and CW contributed to conceptualization and design of experiments. FA and VG performed the experiments and contributed to the analysis. WB, FA, and CW contributed to writing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Anglès, F., Gupta, V., Wang, C. et al. COPII cage assembly factor Sec13 integrates information flow regulating endomembrane function in response to human variation. Sci Rep 14, 10160 (2024). https://doi.org/10.1038/s41598-024-60687-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-60687-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.