Abstract
Rapid expansion of the pulmonary microvasculature through angiogenesis drives alveolarization, the final stage of lung development that occurs postnatally and dramatically increases lung gas-exchange surface area. Disruption of pulmonary angiogenesis induces long-term structural and physiologic lung abnormalities, including bronchopulmonary dysplasia, a disease characterized by compromised alveolarization. Although endothelial cells are primary determinants of pulmonary angiogenesis, mesenchymal cells (MC) play a critical and dual role in angiogenesis and alveolarization. Therefore, we performed single cell transcriptomics and in-situ imaging of the developing lung to profile mesenchymal cells during alveolarization and in the context of lung injury. Specific mesenchymal cell subtypes were present at birth with increasing diversity during alveolarization even while expressing a distinct transcriptomic profile from more mature correlates. Hyperoxia arrested the transcriptomic progression of the MC, revealed differential cell subtype vulnerability with pericytes and myofibroblasts most affected, altered cell to cell communication, and led to the emergence of Acta1 expressing cells. These insights hold the promise of targeted treatment for neonatal lung disease, which remains a major cause of infant morbidity and mortality across the world.
Similar content being viewed by others
Introduction
Lung development requires coordinated interactions between multiple cell types in the epithelial, endothelial, immune and mesenchymal compartments1,2. In utero, the lung is fluid filled and the pulmonary circulation develops in a relatively low oxygen tension environment with low blood flow and high pressure3. Within moments after birth, the perinatal lung undergoes a remarkable transition that enables gas exchange with establishment of a gas–liquid interface, a tenfold increase in pulmonary blood flow and a marked decrease in pulmonary arterial pressure4,5. After birth, the distal lung undergoes structural remodeling including the formation of alveoli by secondary septation and a transition from a double to single capillary layer to increase gas exchange efficiency6.
Lung development requires coordinated interactions between multiple cell types in the epithelial, endothelial, immune and mesenchymal compartments1,2. Lung mesenchymal cells (MC) play a central role in secondary septation, thinning of the interstitium, and transmitting mechanical forces that promote alveolarization7. Although the lung mesenchyme includes multiple, distinct cell types, knowledge surrounding the degree of cellular heterogeneity, dynamic changes in gene expression, and cell–cell interactions during alveolarization remains limited.
Insight into the lung mesenchyme during early postnatal life has significant implications for lung injury and regeneration, across the lifespan, but especially in neonates. Since the lung continues to develop in late pregnancy and through the first decade of life8, premature infants are uniquely susceptible to life-threatening lung disorders, including bronchopulmonary dysplasia (BPD), a lung disease characterized by compromised alveolarization9. Given the importance of the mesenchyme in both physiologic development and developmental disease, we sought to interrogate the lung mesenchyme relative to cellular diversity, dynamic changes in gene expression and cell–cell communication in the perinatal lung, during alveolarization, and in the context of hyperoxia-induced lung injury, a preclinical model of BPD10.
In this report, we combined single cell transcriptomics (scRNA-Seq) with fluorescent multiplexed in situ hybridization (FISH) and computational analyses to characterize changes in composition, localization, and function of MC in the murine lung from just before birth through the first three weeks of postnatal life. Moreover, we applied the same strategies to cells derived from neonatal murine lung after 7 days of hyperoxia (80% oxygen). Consistent with prior studies, MC fell into three broad categories: fibroblasts, airway smooth muscle (ASM)/myofibroblasts (MyoF), and mural cells. Overall, the present strategy identified 14 distinct subtypes, some of which were restricted to perinatal developmental stages. Remarkably dynamic changes in cell phenotype occurred at birth as separate precursor cells for either fibroblasts or ASM/MyoF led to distinct but related cell subtypes between E18.5 and P1. At P7, early ASM cells were localized not only around the airways, but also in the distal lung, spatially proximate to, but transcriptionally distinct from, MyoF. At P7 lung MC were most proliferative and corresponded to the emergence of a novel proliferating population of MyoF. Hyperoxia blocked the maturation-related transcriptomic progression of MC, mirroring the structural arrest in lung development that characterizes BPD in infants, and markedly decreased pericyte and myofibroblast abundance and proliferation. Further, single-cell and bulk transcriptomics and FISH imaging identified previously undescribed Acta1+ MC in hyperoxia-exposed mice. These data demonstrate that the neonatal lung mesenchyme possesses a high degree of cellular diversity and changes dynamically with development, with hyperoxia causing both generalized transcriptomic arrest and the emergence of novel, previously undescribed MC.
Results
The perinatal lung mesenchyme is composed of three groups of cell types
Single cell RNA-Seq (scRNA-Seq) data were generated from eight mice at different stages of perinatal development, with two mice (one female and one male) from four points during development, embryonic day 18.5 (E18.5; early saccular stage), postnatal day of life 1 (P1; late saccular), P7 (early alveolar) and P21 (late alveolar). Lung tissue was resected, perfused, and dissociated as described previously11. Fluorescence activated cell sorting (FACS) was used to enrich mesenchymal cells by depleting cells positive for CD326, CD31, and CD45 (Fig. 1A). Libraries were generated in 384-well plates, combining medium cell throughput with excellent per-cell capture sensitivity as demonstrated previously11. A total of 5493 high-quality cells at an average depth of 980,000 read pairs per cell were analyzed (see “Materials and methods” section), yielding a much larger number of detected genes per cell compared to any other study of the perinatal lung (Fig. S1). Key reagents used in the study are included in Table 1.
The perinatal lung mesenchyme is populated by three groups of cell types. (A) Experimental design including time points sampled, tissue dissociation, and cell sorting to isolate mesenchymal cells. (B) Embedding of lung mesenchymal cells, colored by relative expression of four distinguishing marker genes. (C) As in (B), but colored by cell type group. (D) Dot plot of representative marker genes for each group. (E) Relative abundance for each group in normal development. (F) As in (B), but colored by time point of each cell. At this level of resolution, airway smooth muscle (ASM) and myofibroblasts (MyoF) belong to the same group (ASM/MyoF).
An initial representation of the perinatal lung mesenchyme was constructed by computing a t-distributed stochastic neighbor embedding (t-SNE)12 and categorizing each cell based on expression of established marker genes (Fig. 1B). Three groups of mesenchymal cell types were defined (Fig. 1C). Fibroblasts were marked by Adh113,14. Tgfbi expression marked a heterogeneous group of cells that included including both airway smooth muscle cells and myofibroblasts (ASM/MyoF)15. Mural cells were marked by Pdgfrb16 (Fig. 1D). At all time-points, except for P7, most lung mesenchymal cells were fibroblasts. Airway smooth muscle cells and myofibroblasts abundance peaked at P7, with mural cell abundance increasing slowly with development (Fig. 1E). Both cell type abundance and gene expression changed dynamically with maturation (Fig. 1F). No cluster coexpressed both Tgfbi and Pdgfrb (Fig. 1D), previously described as vascular smooth muscle cells (VSM) markers17, while airway smooth muscle cells and myofibroblasts embedded in distinct locations across development (Fig. 1F), underscoring the importance of smooth muscle cell subtype identification18,19,20,21.
Perinatal and adult mesenchymal cell subtypes are transcriptionally distinct
We then undertook more discrete cell subtype annotation18,19,21,22,23. Leiden clustering24 resulted in 14 distinct clusters (Fig. 2A). Annotation was achieved by harmonized co-embedding with an adult cell atlas, Tabula Muris Senis (TMS, Fig. 2B)19 and marker gene analysis (Fig. 2C). Cell clusters at P21 cell were consistent with the adult atlas, while clusters derived from earlier time points (clusters 1–3, 5, 7–8, 10–11) embedded separately25, supporting the notion that perinatal lung cells possess transcriptomic profiles distinct from adult. Clusters 1–6 were consistent with fibroblasts, clusters 7–11 included airway smooth muscle (ASM) and myofibroblasts, and clusters 12–14 mural cells.
Harmonized fine grained composition of the perinatal mesenchyme. (A) Embedding as in Fig. 1B, but colored by fine cell type and annotated with cluster numbers 1–14. This color code is used in subsequent figures. (B) Harmonized embedding of perinatal mesenchymal cells with mesenchymal cells from TMS colored by source (top panel, gray TMS cells drawn on top) and cell type (bottom, merging concordant cell types between our data and TMS). (C,D) Dotplots of all mesenchymal cell types in this study for (C) marker genes and (D) components of the extracellular matrix. TMS Tabula Muris Senis.
For cluster 12, which co-embedded between smooth muscle-like cells and pericytes (Fig. 2B), the coexpression of both smooth muscle (Tagln, Acta2) and mural (Prgfrb) genes (Fig. 2C) identified this population as vascular smooth muscle26,27. Cluster 9 derived from P21 mice (Fig. 1E) and expressed Hhip, an established airway smooth muscle marker28 but not Pdgfra, a myofibroblast marker, therefore these cells were annotated as ASM (Fig. 2C). This led to the tentative annotation of cluster 8 (Hhip+ Pdgfra−) as early ASM and cluster 10 (Hhip− Pdgfra+) as myofibroblasts (Fig. 2C). Both clusters comprised mostly cells from P1 and P7. At E18.5, a single Tgfbi+ cluster was found (cluster 7). It coexpressed Hhip and Pdgfra and was therefore annotated as ASM/MyoF precursors. Clusters 2, 11, and 14 expressed Mki67 and were therefore labeled as proliferating fibroblasts, myofibroblasts, and pericytes, respectively (Fig. 2C). To contextualize our cell type definitions, we computed dot plots of marker genes from recent publications against our cell types22,29 (Figs. S2, S3).
To further buttress the physiologic relevance of the transcriptomic findings, expression of genes associated with extracellular matrix (ECM) production and remodeling were evaluated (Fig. 2D). Fibroblasts shared ECM components Mfap4, Fn1, Vcan, and Ogn. In alveolar fibroblasts (AlvF), subtype-specific expression included Spon1 which encodes a secreted adhesion protein30. Mature and Early Adventitial fibroblasts (AdvF) expressed Mfap5, which contributes to tissue elasticity31, and Dcn, which confers resistance to tissue compression, while only adventitial fibroblasts from P21 mice expressed Podn, encoding a molecule that constrains smooth muscle cell proliferation and migration32. Early ASM highly expressed Aspn, which inhibits canonical TGFβ and SMAD signaling33 and competes with Dcn for collagen binding34, while ASM from P21 animals had reduced expression. However, mature ASM expressed Lum, which regulates collagen fibril organization. Interestingly, Early ASM, found in P1 and P7 mice, did not express Lum, suggesting other genes might regulate collagen fibrils at those time points35,36. Pericytes, which modulate vascular tone, endothelial cell (EC) migration and angiogenesis37, expressed Mcam, Mfge8, Postn, and Eng. Vascular smooth muscle had the highest expression of elastin (Eln), while all other cell types except pericytes expressed some level of Eln.
Fibroblast and ASM/MyoF precursors differentiate at the onset of air-breathing life
The cell composition of the lung changed dramatically between E18.5 and P1, a time window of just 48 h. Fibroblast precursors, the most abundant mesenchymal type at E18.5, disappeared by P1 (Fig. 3A), concomitant with the appearance of two distinct types of postnatal fibroblasts (Fig. 3B, Fig. S3) distinguished by expression of either Col13a1 or Col14a1 (Fig. 3C), resembling adult alveolar and adventitial fibroblasts, respectively. Col13a1 and Col14a1 were coexpressed in the E18.5 precursors, suggesting a common origin (Fig. 3C), a hypothesis corroborated by trajectory analysis via diffusion maps, pseudotime, and RNA velocity (Fig. S4). In terms of ECM-associated gene expression, fibroblast precursors appeared slightly more similar to alveolar fibroblasts (Fig. 2D, Fig. S4). Differentially expressed genes between embryonic and a balanced mix of postnatal fibroblasts showed a statistical enrichment of mesenchyme-, ECM-, and vascular development-associated pathways in both groups (Fig. 3D, red/blue bars upregulated in precursors/postnatal cells, respectively)38.
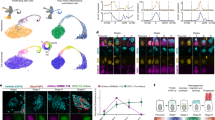
Developmental changes in perinatal fibroblasts, myofibroblasts, and airway smooth muscle. (A) Changes in composition of lung fibroblast subtypes between E18.5 and P21. (B) Embedding of fibroblasts and precursors at E18.5 (left) and P1 (right). (C) Distribution of expression for Col13a1 and Col14a1 in fibroblasts. (D) Pathways enriched among genes up- (red, top) or down-regulated (blue, bottom) in early postnatal fibroblasts versus prenatal fibroblast precursors38. (E) Changes in abundance of ASM, myofibroblasts and their precursors between E18.5 and P1. (F) Embedding of ASM and myofibroblasts at E18.5 (left) and P1 (right). (G) Distribution of expression of Hhip and Pdgfra in myofibroblasts, and early ASM, and their precursors. (H) Pathways enriched among genes up- (red, top) or down-regulated (blue, bottom) in early postnatal myofibroblasts and ASM versus their precursors. (I) Expression changes of Crh between E18.5 and P1. (J) In-situ hybridization of Crh+ Tgfbi+ (solid arrows) and Crh- Tgfbi+ cells (dashed arrows) in E18.5 and P1 lungs. (K) Expression changes of Hsd11b1 in fibroblasts and Hif3a in both fibroblasts, ASM, and myofibroblasts between E18.5 and P1. (L) Fraction of proliferative fibroblasts and myofibroblasts at postnatal time points. (M) In-situ hybridization of nonproliferating (Mki67− solid arrows) and proliferating (Mki67+ dashed arrows) myofibroblasts at P7. (N) Embedding of Early ASM and myofibroblasts, colored by subtype and by expression of select genes. (O) In-situ hybridization of P7 lungs around airways (left) and in the distal lung (right). Dashed arrows: Tgfbi+ Hhip+ cells, solid arrow: Tgfbi+ Pdgfra+ cells. (P) Quantification of the degree of coexpression of Hhip, Tgfbi, and Pdgfra from in-situ images of P7 mice. P-values are Kolmogorov–Smirnov tests with Bonferroni multiple hypothesis correction.
Airway smooth muscle/myofibroblast precursor abundance decreased between E18.5 and P1 (Fig. 3E). Early airway smooth muscle cells and myofibroblasts were rare before birth and increased in abundance by P1 (Fig. 3E,F). Airway smooth muscle/myofibroblast precursors coexpressed Hhip and Pdgfra, while postnatal MyoF expressed only Pdgfra and postnatal Early ASM expressed only Hhip (Fig. 3G). Pathway analysis suggested that ASM/MyoF precursors expressed higher levels of genes related to vascular development and muscle cell proliferation (Fig. 3H, red bars) and lower levels of smooth-muscle contraction genes (blue bars) compared to balanced postnatal ASM/MyoF cells. Airway smooth muscle/myofibroblast precursors but not postnatal counterparts expressed corticotropin releasing hormone (Crh, Fig. 3I), an early signal for glucocorticoid production within the hypothalamic–pituitary–adrenal (HPA) axis. Crh plays an essential role in lung maturation39 via cortisol-induced increases in surfactant protein release40,41. Crhr1 and Crhr2, which encode the Crh receptors, were not expressed by any other cell type in the perinatal lung (Fig. S5)11,17,42, not in the adult atlas19. Tgfbi+ Crh+ cells were detected by in-situ hybridization imaging (RNA-Scope) within the lung parenchyma in E18.5 mouse embryos but not postnatally (Fig. 3J, solid arrows, n = 6 animals per group). Fibroblast precursors but not postnatal fibroblasts expressed high levels of Hsd11b1 (Fig. 3K), the main enzyme responsible for converting cortisone into its bioactive form cortisol, indicating a local amplification of glucocorticoid action43. Expression of Hif3a, a negative regulator of Hif1a, was lost in both fibroblasts and ASM/MyoF cells after birth, suggesting tight regulation of this pathway at the critical transition between fetal development and air-breathing life (Fig. 3K).
Mki67+ proliferating myofibroblasts and pericytes were observed at P1 and increased in abundance at P7 (Fig. 3L). Proliferating myofibroblasts were localized throughout the parenchyma of P7 lungs as Tgfbi+ Pdgfra+ Mki67+ cells (Fig. 3M, dashed arrows, n = 2) adjacent to Tgfbi+ Pdgfra+ Mki67− nonproliferating myofibroblasts (solid arrows). There were no proliferating (myo-)fibroblasts detected via scRNA-Seq from P21 mice (Fig. 3L), consistent with a progression towards cell quiescence towards the end of alveolarization and adulthood.
The relationship between postnatal Early ASM and myofibroblasts was then examined in detail. These cells, in addition to shared Tgfbi expression and exclusive expression of Hhip (for Early ASM) and Pdgfra (for myofibroblasts) as mentioned above, showed partial expression of Actc1, an alpha actin typically expressed in heart muscle44, in a fraction of Early ASM and Stc1, a regulator of calcium levels that has been implicated in myopathy45, in a fraction of myofibroblasts, indicating further layers of latent transcriptomic heterogeneity (Fig. 3N). In-situ hybridization was then used to determine the location of each cluster at P7. As expected, only Tgfbi+ Hhip+ Pdgfra− cells, a profile matching Early ASM, were found encircling airways (Fig. 3O left, dashed arrows, n = 2), buttressing the scRNA-Seq-based annotation. Surprisingly, however, not only Tgfbi+ Hhip− Pdgfra+ (solid arrows), matching myofibroblasts, but also Tfgbi+ Hhip+ Pdgfra− cells (dashed arrows), matching Early ASM, were observed in the distal parenchyma (Fig. 3O, right, n = 2). Automated image analysis on the distal parenchyma only (10 images, no airways) demonstrated a lower correlation between Pdgfra and Hhip than the correlation of either molecule with Tgfbi (Fig. 3P), which mirrors the transcriptomics results (Fig. 3N).
Mural cells change gradually during perinatal development
Mural cells, which include pericytes and vascular smooth muscle, form the outer lining of the vasculature and help support the endothelium and regulate vascular tone. In the perinatal lung, the transcriptomes of mural cells changed slowly between E18.5 and P21. Relative cell type abundance of mural cells (Fig. 1D) and of each mural subtype (pericytes and VSM) (Fig. 4A) increased steadily over time. A cluster of proliferating pericytes peaked in abundance at P7 (Fig. 4B), at the same time as proliferating myofibroblasts. Confocal imaging of P7 lung confirmed the presence of proliferating pericytes that co-expressed Cox4i2 and Mki67 (Fig. 4C, n = 2). A developmental shift in the pericyte transcriptome was visible in our embedding (Fig. 4D, colors as in Fig. 1F). Further analyses revealed specific transcriptomic progressions: genes exhibiting gradual decreases in expression over time (e.g. Aspn), gradual increases over time (e.g. Enpp2), marked downregulation after birth (e.g. Timp3), and either marked down-regulation (e.g. Acta2) or up-regulation (e.g. Cxcl14) between P7 and P21 (Fig. 4E, same colors—all P-values are Bonferroni-adjusted KS tests). In contrast, VSM cells did not exhibit a clear transcriptomic shift over time in the embedding (Fig. 4F) and lacked high expression of Mki67 (Fig. 2A).
Mural cells encompass proliferating pericytes at P7 and harmonized vascular smooth muscle characterized by specific transcription factors. (A) Mural cell type composition between E18.5 and P21. Colors as in Fig. 2A. (B) Fraction of proliferating pericytes at postnatal time points. (C) Representative FISH image of Mki67+ Cox4i2+ proliferating pericytes in P7 lungs. (D) Embedding of pericytes, colored by time point. (D) Violin plots of the expression of select genes that change in expression over perinatal development in pericytes. (F) Embedding of VSM, colored by time point. (G) Violin plots of gene expression in pericytes versus VSM (left) and ASM versus VSM (right) for transcription factors (TFs) marking either cell type. (H) Combined statistical enrichment of expressed transcription factors in VSM versus pericytes (x axis) and versus ASM (y axis). Genes in the upper right quadrant (violet) are specific to VSM versus both ASM and pericytes, while genes in the lower left quadrant (black) are specifically absent in VSM but not in pericytes nor ASM. (I) Harmonized embedding of VSM from this study (red circles) and overlapping cells from Tabula Muris Senis (gray squares), identified using a rigidly expanded principle component analysis (PCA) ellipse. These cells are originally annotated as pericytes, myofibroblasts, or fibroblasts. (J) Differential expression of select transcription factors in newly identified atlas VSM versus atlas pericytes (x axis) and versus atlas myofibroblasts (y axis).
The transcriptional distinction between VSM, pericytes, and ASM remains incompletely defined18,19,26,29. For instance, Guo et al. annotated as “Pericyte-1” a cell cluster which lacks pericyte markers (Fig. S6). To clarify the transcriptional identity of VSM, our analysis focused on transcription factors (TFs) given the capacity to effectively distinguish between cell types20. Several transcription factors were upregulated in VSM (purple) relative to ASM (green) and in VSM compared to pericytes (pink) (Fig. 4G, top) or downregulated in VSM versus the same cell types (Fig. 4G, bottom). VSM also expressed TFs that were absent in both ASM and pericytes (Fig. 4H, purple dots): these included Prrx146, Osr147 and Id448. Tbx449 and other genes were expressed in both ASM and pericytes but specifically downregulated in VSM (black dots). Given the absence of a VSM annotation in adult atlas19, we reanalyzed atlas cells that overlapped in the harmonized embedding (Fig. 2B) with our VSM cells (Fig. 4I). The 8 atlas cells within the principal component analysis-based ellipse (gray squares) were originally annotated as adult myofibroblast or pericyte. However, the co-embedding coordinates overlapped with those of our VSM (red circles). Differential expression of transcription factors between these 8 atlas cells and other atlas pericytes and myofibroblasts resulted in a specific TF fingerprint (Fig. 4J) that was almost identical to the developmental VSM fingerprint previously defined (Fig. 4H), with upregulation of Prrx1, Osr1 and Id4, and downregulation of Tbx4 compared to both other cell types. In-situ hybridization of P7 lungs with Tgfbi, Pdgfra, and hydrazide (which marks vessels but not airways) showed that cells lining the large airways highly expressed Tgfbi (which is expressed by both ASM and myofibroblasts), but cells located between the double elastic laminae of arterioles were Tgfbi negative (Fig. 4, Supplementary Fig. 2). Overall, these data confirmed that VSM differ from both ASM and myofibroblasts not only in their location within the tissue but also in their transcriptional profile, in agreement with some reports26,50.
Hyperoxia inhibits mesenchymal transcriptional progression and pericyte proliferation
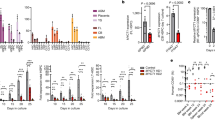
To study the discrete transcriptomic effect of hyperoxia on the developing lung, we performed scRNA-Seq and in-situ analysis on mice exposed to 80% oxygen (hyperoxia, HO) from birth through P710,51,52,53. Early lung growth was impaired by hyperoxia with larger and fewer alveoli in mice exposed to HO, compared to normoxia, while the radial alveolar count was lower (Fig. 5A; P < 0.001, n = 3, student’s test) in hyperoxic compared to normoxic mice. Hyperoxia increased the relative abundance of fibroblasts but decreased the abundance of ASM, proliferating myofibroblasts and pericytes (Fig. 5B). We hypothesized that hyperoxia might render the lung cellular composition at P7 more similar to younger animals. We compared cell type abundance profiles via bootstrapping and found that the transcriptomes of hyperoxia-exposed mice were significantly more similar to healthy P1 mice than P7 mice (P = 0.001, Fig. 5C), indicating that hyperoxia inhibited the cellular diversification of the mesenchyme. We next asked whether hyperoxia delayed developmental changes in the transcriptome within each cell type. We specifically interrogated the differential expression of genes that exhibit transcriptional progression during development in normoxia (e.g. higher at P7 than P1), and found that hyperoxia inhibited this progression in all the mesenchymal populations, especially in the mural cells (Fig. 5D).
Hyperoxia delays mesenchymal transcriptional progression and suppresses pericyte proliferation. (A) Representative histologic sections from murine lungs developing in normoxia (left) or hyperoxia (right) through P7 and radial alveolar counts in normoxic (N) and hyperoxic (HO) P7 mouse lungs. Scale bar: 50 μm. ****P-value < 0.001. (B) Cell subtype composition in between normoxia and hyperoxia at P7. (C) Cell subtype composition changes in hyperoxic mice. The arrows indicate the observed severe loss of pericytes (solid) and complete loss of proliferating pericytes (dashed). (D) Kernel density estimate of 1000 bootstraps over cells (balanced at each time point) for the difference in distance on cell composition between hyperoxic mice and P7, minus hyperoxic mice and P1. A positive number indicated that the hyperoxic mesenchyme composition is closer to a healthy P1 mouse than a P7 one. Only 1 out of 1000 simulations showed hyperoxia closer to P7 than P1. (D) Differential expression between normoxic and hyperoxic cells in each cell type for genes progressing during early postnatal development (upregulated at P7 versus P1). (E) Representative in-situ images of pericytes (solid) and proliferating pericytes (dashed) in P7 normoxia and hyperoxia mice. (F) Quantification of the relative abundance of pericytes and proliferating pericytes in in-situ images from P7 mice using an unsupervised image analysis approach over thousands of cells. (G) Representative confocal imaging of lung tissue at P7 from Notch3-CreERT2-tdT expressing mouse pups who received tamoxifen from P1 to P3 and were placed in either normoxia or hyperoxia for seven days from birth to P7. ⍺-smooth muscle actin (⍺-SMA) shown in green. (H) Number of detected ligand-receptor cell–cell interactions between pericytes and other cell types at each time point during normal development and in hyperoxia mice at P7. (I) Strength of select interactions with pericytes measured as product of gene expression and pericyte abundance in normoxia and hyperoxia. Data sources: endothelial cells17 and immune cells11.
Hyperoxia dramatically decreased pericyte number and especially proliferating pericytes. The scRNA-Seq data (Fig. 5B, solid and dashed arrows, respectively) were confirmed in vivo (Fig. 5E, n = 2 for each group). Hyperoxia reduced the abundance of both total pericytes (P = 0.003, KS test) and proliferating pericytes (P = 0.01, KS test) (Fig. 5F, each dot represents an analyzed image). To further confirm this observation Notch3-CreERT2-tdT mice were given tamoxifen at P0 and P3 and placed in normoxia or hyperoxia from until P7. Deep imaging of thick distal lung section (150 μm) indicated that compared to normoxia, pericyte number was markedly reduced in hyperoxia (Fig. 5G, representative images of n = 3 animals for each group), and the morphology of the pericytes disrupted, with a decrease in the number and length of projections (Fig. 5G, high mag inset).
To determine whether pericyte loss might affect the cell–cell communication networks, we used the CellPhoneDB database54 to count the number of putative ligand–receptor interactions (at least 20% of cells expressing each gene) between pericytes and EC, other mesenchymal cells, and immune cells at all four time points and in hyperoxia at P7. Hyperoxia increased the number of putative interactions between remaining pericytes and all other cell types (Fig. 5H). The overall “interaction strength”, the product of cell type abundance and gene expression levels, varied by gene (Fig. 5I). For some genes, down-regulation intensified the effect of pericyte loss, possibly leading to loss of signaling. This analysis demonstrated a potential decrease in signaling for Pdgfrb, an essential component of a pathway promoting pericyte proliferation and migration towards the endothelium during vascular development55, Gjc1, a connexin that promotes vessel stability and EC quiescence56, and midkine or Mdk, a heparin binding, proangiogenic cytokine57. In contrast, other genes exhibited up-regulation of gene expression, which may serve to mitigate the effect of diminished pericyte type abundance and thereby preserve signaling.
Hyperoxia can cause the emergence of Acta1+ cells in the neonatal lung
In hyperoxic mice, novel populations of Acta1+ cells were observed (Fig. 6A). The Acta1+ cells have not been previously described during lung development or in hyperoxia18. The fraction of mesenchymal cells expressing Acta1+ cells is significantly greater in hyperoxia, compared to normoxia (Fig. 6A; P < 0.001). The two novel clusters (henceforth called HA1 and HA2) were projected in distinct parts of the embedding, with HA2 located close to Alveolar fibroblasts (Fig. 6B). Both clusters expressed the neuron-specific tubulin (Tubb3) and Lgals3, encoding a -galactosidase binding lectin that is up-regulated in tissue fibrosis58 as well as the cardiac muscle gene Actc1 and the membrane channel Aqp3. The HA2 cluster also expressed genes characteristic of alveolar fibroblasts (e.g. Wnt2, Col13a1) (Fig. 6C), explaining its embedding proximity to fibroblasts. Acta1+ cells did not express endothelial (e.g. Pecam1, Cdh5), epithelial (e.g. Epcam), or immune (e.g. Ptprc) marker genes.
Hyperoxia causes the emergence of Acta1+ cells in the lung. (A) Relative cell subtype abundance for broad mesenchymal types in normoxia and hyperoxia. (B) Embedding as in Fig. 2A but with the additional populations hyperoxic Acta1+ 1 (HA1, in purple) and hyperoxic Acta1+ 2 (HA2, in red). Other cells are colored in gray. (C) Dot plot of marker genes for Acta1+ cells. (D) Reanalysis of bulk transcriptomics in P7 lungs exposed to hyperoxia shows up-regulation of Acta1 and other marker genes for the novel cell populations. Data from Ref.59. FDR false discovery rate. Downregulated genes are not shown. (E) Representative in-situ hybridization images of Acta1 (green) expression in normoxia and hyperoxia. Cells with three or more Acta1 molecules are indicated by arrows. (F) Quantification of percentage of Acta1+ cells in images from normoxic and hyperoxic mice. *P-value < 0.05.
To further validate the presence Acta1+ cells, we reanalyzed bulk RNA-Seq data from an independent study on hyperoxia-exposed P10 murine lungs. Acta1 was markedly increased in hyperoxia (n = 6), compared to normoxia, (n = 5) P10 mice. Tubb3, Aqp3, and Lgals3 expression was also increased by hyperoxia59 (Fig. 6D). In-situ hybridization imaging of lungs from a separate group of mice (n = 5 for each group) confirmed an increased density of Acta1+ cells in both male and female hyperoxia-exposed mice versus normal P7 mice, especially near pulmonary blood vessels (Fig. 6E). To further buttress the findings, additional in situ studies were performed on normoxic (n = 3) and hyperoxic (n = 3) P7 mice. The Acta1+ expressing cells, quantified as a percentage of the overall number of cells, were markedly increased in hyperoxia, compared to normoxia (Fig. 6F; P < 0.05, n = 3, student’s test). Thus, with hyperoxia a previously undetected, but transcriptionally distinct Acta1+ cell population emerges in the neonatal lung.
Discussion
Mesenchymal cells are responsible for essential functions in the developing lung, including modulating the composition of the extracellular matrix, providing contractility to airways and vasculature, and supporting angiogenesis and alveolarization5,8,26. Single-cell transcriptomics has been recently applied by several groups to investigate the composition of the lung mesenchyme with high granularity22,26,29,50. Despite similar experimental approaches across studies, there have been substantial differences in the computational analyses and subsequent biological interpretations. This study sought to comprehensively define cellular identity of the perinatal lung mesenchyme in physiologic and pathophysiologic lung development.
The transcriptional identity of several mesenchymal cell subtypes was clarified. A transcriptomic signature was identified for vascular smooth muscle cells that distinguishes the population from airway smooth muscle, myofibroblasts, and pericytes19,20,22,29.Consistent with a prior report alveolar and adventitial fibroblasts were noted20 while lipofibroblasts were not identified26,29,50. Myofibroblasts and airway smooth muscle cells were present and transcriptionally distinct in early postnatal development. Airway smooth muscle cells, but not myofibroblasts were present at P21, suggesting that in the absence of injury ductal myofibroblasts may not persist through adulthood50. Intriguingly, the identification of an embryonic mesenchymal cell population expressing Crh, which encodes a hormone that controls systemic glucocorticoid levels via the hypothalamus—pituitary gland—adrenal gland axis, was identified computationally and in situ. These observations support prior reports and point to a novel role for the cell in promoting perinatal lung epithelial cell maturation26,50. Lung development proceeds normally in Crh-deficient mice through E17.5, but septal thinning and airspace formation is subsequently insufficient and newborn pups die within 24 h60. The significance of Crh expressing lung myofibroblast precursors remains unknown. However, a paradigm wherein cells in the lung call for systemic cortisol release to promote surfactant release immediately prior to parturition merits further study. Interrogation of the function and physiologic role with cell type-specific gain- and loss-of-Crh expression will provide meaningful insight.
While the effects of hyperoxia on the transcriptomes of lung cells have been described in previous publications17,18, this study focused specifically on the effects on hyperoxia on mesenchymal cells. The dramatic loss of proliferating pericytes and the subsequent alteration of cell–cell communication networks are suggestive of a potentially underappreciated role for pulmonary pericytes in protecting against neonatal lung injury61. While the specific functions of lung pericytes are not fully understood, pericyte deficiency can disrupt the blood–brain barrier and contributes to the capillary instability, vascular leak and macular edema of diabetic retinopathy62,63. Ongoing efforts by our groups are aimed at clarifying the physiologic implications of partial pericyte loss on the altered tissue structure and pathologic vascular remodeling observed during lung recovery after hyperoxia. Given that this murine model recapitulates many features of the human disease, elucidation of novel functions for understudied cell types might lead to new strategies to improve clinical care for prematurely born infants with bronchopulmonary dysplasia.
The emergence of Acta1-expressing cells in multiple hyperoxia-exposed mice, seen by single cell and bulk transcriptomics and confirmed via in-situ hybridization, raises questions about developmental ontogeny and putative pathophysiologic function. While much remains to be discovered about these cells, in a previous study cardiomyocyte-associated genes were differentially expressed in a bulk transcriptomic of whole lung after hyperoxia exposure64. Intriguingly, cardiomyocytes of the pulmonary vein, which have been associated with asthma65, express Actc1 and partially Acta1 but not Tubb3 and Aqp323, suggesting that Acta1-expressing cells might be functionally related and possibly developmentally linked to cardiomyocytes. Given the large variability in severity seen in both hyperoxia-exposed mice and infants with bronchopulmonary dysplasia—only one of our two mice sequenced via single cell RNA-Seq contained Acta1+ cells, it is tempting to speculate that these cells might be a stochastic response of the body restricted to the most severe forms of disease. Future experiments using transgenic mice for Acta1, Tubb3, or other marker genes specific for these cells are planned to test this hypothesis. Though prior transcriptomic studies of the neonatal lung have employed hyperoxia as a lung injury model18, this report is the first to identify Acta1+ cells in the lung. The study design likely accounts for the discovery of the Acta1+ cells as the cell digestion protocol was carefully optimized11,66. Moreover, the read depth per cell was significantly greater than reported in prior studies (Fig. S1). Each of these factors likely contributed to our ability to identify previously undetected, relatively low abundance cells.
Overall, this study constitutes a coherent cellular portrait of the perinatal lung mesenchyme that aims to overcome discrepant biologic interpretations in the existing literature and deeply characterize the cellular and transcriptional changes that accompany neonatal hyperoxia exposure, a widely used model for bronchopulmonary dysplasia. The combination of de novo high-quality data collection, integration with and comparative secondary analysis of extant data sets, and validation via in-situ hybridization provides a blueprint that could be used to systematically improve our understanding of the mesenchymal compartment across organs and organisms.
Materials and methods
Mouse lung cell isolation
C57BL/6 mice were obtained from Charles River Laboratories. For studies using E18.5, P1, and P7 murine lungs, pregnant dams were purchased, and pups aged prior to lung isolation. At E18.5, the dam was asphyxiated with CO2 and pups extracted. At P1, P7, and P21 pups were euthanized with euthanasia solution (Vedco Inc). For hyperoxia experiments, pups were exposed to 80% atmosphere for 7 days as previously described17. Genetic sex of mice at developmental stages E18.5, P1 and P7 was determined by performing PCR amplification of the Y chromosome gene Sry. P1 and P7 mice were sexed through identification of a pigment spot on the scrotum of male mice67. For all timepoints, female and male mice were randomly selected for the studies. For all timepoints, except E18.5, the pulmonary circulation was perfused with ice cold heparin in 1× PBS until the circulation was cleared of blood. Lungs were minced and digested with Liberase (Sigma Aldrich) in RPMI for 15 (E18.5, P1, and P7) or 30 (P21) minutes at 37 C, 200 rpm. Lungs were manually triturated and 5% fetal bovine serum (FBS) in 1× PBS was used to quench liberase solution. Red blood cells were lysed with 1× RBC lysis buffer (Invitrogen) as indicated by the manufacturer and total lung cells counted on Biorad cell counter (BioRad). Protocols were prospectively approved by the Institutional Animal Care and Use Committee at Stanford (APLAC #19087). All murine studies adhered to American Physiological Society and the United States National Institutes of Health guidelines for humane use of animals for research. Moreover, the present study is reported in accordance with the ARRIVE guidelines.
Immunostaining and fluorescence-activated cell sorting (FACS) of single cells
Lungs were plated at 1 × 106 cells per well and stained with Fc block (CD16/32, 1:100, Tonbo Biosciences) for 30 min on ice. Cells were surface stained with the endothelial marker CD31 (1:100, eBioscience/ThermoFisher), epithelial marker Epcam (1:100, eBioscience/ThermoFisher), and immune marker CD45 (1:100, eBioscience/ThermoFisher) for 30 min on ice. The live/dead dye, Sytox Blue (Invitrogen), was added to cells and incubated for 3 min prior to sorting into 384-well plates (Bio-Rad Laboratories, Inc) prefilled with lysis buffer using the Sony LE-SH800 cell sorter (Sony Biotechnology Inc), a 100 μm sorting chip (Catalog number: LE-C3110) and ultra-purity mode. Single color controls were used to perform fluorescence compensation and generate sorting gates. 384-well plates containing single cells were spun down, immediately placed on dry ice and stored at − 80 ºC.
cDNA library generation
RNA from sorted cells was reverse transcribed and amplified using an adaptation of the Smart-Seq2 protocol for 384-well plates11. Concentration of cDNA was quantified using Quant-it Picogreen (Life Technologies/Thermo Fisher) to ensure adequate cDNA amplification and cDNA was normalized to 0.4 ng/μL. Tagmentation and barcoding of cDNA was prepared using in-house Tn5 transposase and custom, double barcoded indices20. Library fragment concentration and purity were quantified by Agilent bioanalyzer. Libraries were pooled and sequenced on Illumina NovaSeq 6000 with 2 × 100 base kits and at a depth of around 1 million read pairs per cell.
scRNA-Seq analysis
Sequencing reads were mapped against the mouse genome (GRCm38) using STAR aligner (https://github.com/alexdobin/STAR)68 and gene expression was quantified using HTSeq (https://github.com/simon-anders/htseq)69. To coordinate mapping and counting, snakemake (https://snakemake.readthedocs.io/en/stable/) was used70. Gene expression count tables were converted into loom objects (https://linnarssonlab.org/loompy/) and cells with less than 50,000 uniquely mapped reads or less than 400 genes per cell were discarded. Doublets were discarded by excluding small clusters and single cells that coexpress markers for incompatible cell types. Counts for the remaining NNN cells were normalized to counts per million reads. For t-distributed stochastic embedding (t-SNE)12, 500 features were selected that had a high Fano factor in most mice, and the restricted count matrix was log-transformed with a pseudocount of 0.1 and projected onto the top 25 principal components using scikit-learn (https://scikit-learn.org/)71. Unsupervised clustering was performed using Leiden (https://github.com/vtraag/leidenalg) (C++/Python implementation)24. Singlet (https://github.com/iosonofabio/singlet) and custom Python3 scripts were used: the latter are available at https://github.com/iosonofabio/lung_neonatal_mesenchymal/. Pathway analysis on the differentially expressed genes was performed via Metascape38 on the top 100 most differentially expressed genes for each comparison: precursors versus sample-balanced joint progenies and subsequent time points within each cluster. The most enriched pathways against a permutation test are shown ordered by significance from top to bottom (negative log of the P-value). Batch-corrected KNN25 was used to compare our P21 data with the Smart-seq 2 data from Tabula Muris Senis19.
In-situ validation using RNAscope and immunofluorescence (IF)
Embryonic and post-natal mice were euthanized as described above. Female and male mice were randomly selected from the litter, and at least two litters were used to source the lung tissue for all validation studies. E18.5 lungs were immediately placed in 10% neutral buffered formalin following dissection. P1, P7, and P21 murine lungs were perfused as described above, and P7 and P21 lungs inflated with 2% low melting agarose (LMT) in 1× PBS and placed in 10% neutral buffered formalin. Following 20 h incubation at 4 °C, fixed lungs were washed twice in 1× PBS and placed in 70% ethanol for paraffin-embedding. In situ validation of genes identified by single cell RNA-seq was performed using the RNAscope Multiplex Fluorescent v2 Assay kit (Advanced Cell Diagnostics) and according to the manufacturer’s protocol. Formalin-fixed paraffin-embedded (FFPE) lung section (5 μm) were used within a day of sectioning for optimal results. Nuclei were counterstained with DAPI (Life Technology Corp.) and extracellular matrix proteins stained with hydrazide72. Opal dyes (Akoya Biosciences) were used for signal amplification as directed by the manufacturer. Images were captured with Zeiss LSM 780 and Zeiss LSM 880 confocal microscopes, using 405 nm, 488 nm, 560 nm and 633 nm excitation lasers. For scanning tissue, each image frame was set as 1024 × 1024 and pinhole 1AiryUnit (AU). For providing Z-stack confocal images, the Z-stack panel was used to set z-boundary and optimal intervals, and images with maximum intensity were processed by merging Z-stacks images. For all both merged signals and split channels were collected.
Image quantification
Images were quantified in two ways. First, for pericytes we used automatic image segmentation using cellprofiler73 to identify RNA molecules from RNA-Scope and the cell boundaries, and then counted the number of cells with at least 3 positive RNA molecules in different conditions. Second, for other cells that were challenging to segment (e.g. ASM/MyoF), whole-image correlations of the fluorescence channels were computed.
In-situ validation of pericyte abundance using transgenic mice
Notch3-CreERT2-tdT mice were given tamoxifen at P0 and P3 and placed in normoxia or hyperoxia (0.80 FiO2) from P1 to P7. Lungs were harvested at P7 and inflation fixed. Deep imaging of thick lung section (150 μm) allowed us to visualize the abundance and location of tdT-labeled pericytes in the distal lung.
Statistical analyses
Single cell omics data do not follow a normal distribution and are often difficult to parameterize altogether. Therefore, to ensure statistical soundness of all analyses, a statistical plan based on nonparametric tests was deployed throughout the study, exploiting the fact that hundreds of quasi-replicate cells were sampled. To identify differentially expressed genes within cell populations, between time points, and between normoxia and hyperoxia, nonparametric Kolmogorov–Smirnov (KS) tests on the distributions of gene expression were performed, and either the genes with the largest absolute value of the test statistic or the genes above a certain KS statistic (e.g. 0.3) with the largest average fold change were chosen. Nonparametric bootstrapping was used to evaluate the developmental arrest of hyperoxia. Correlation tests and proportion tests for cell population abundances were used in some image analyses.
Data availability
Raw fastq files, count tables, and metadata are available on NCBI’s Gene Expression Omnibus (GEO) website: GSE172251 and GSE175842 (immune cells). An h5ad file with the processed data is available on FigShare and can be accessed and downloaded at: https://figshare.com/articles/dataset/Cell_atlas_of_the_murine_perinatal_lung/14703792. Bulk transcriptomic data were sourced from and are available in Ref.59.
References
Branchfield, K. et al. A three-dimensional study of alveologenesis in mouse lung. Dev. Biol. 409, 429–441 (2016).
Peng, T. et al. Coordination of heart and lung co-development by a multipotent cardiopulmonary progenitor. Nature 500, 589–592 (2013).
Rudolph, A. M. Distribution and regulation of blood flow in the fetal and neonatal lamb. Circ. Res. 57, 811–821 (1985).
Cassin, S., Dawes, G. S., Mott, J. C., Ross, B. B. & Strang, L. B. The vascular resistance of the foetal and newly ventilated lung of the lamb. J. Physiol. 171, 61–79 (1964).
Dawes, G. S., Mott, J. C., Widdicombe, J. G. & Wyatt, D. G. Changes in the lungs of the new-born lamb. J. Physiol. 121, 141–162 (1953).
Schittny, J. C. Development of the lung. Cell Tissue Res. 367, 427–444 (2017).
Li, R., Li, X., Hagood, J., Zhu, M.-S. & Sun, X. Myofibroblast contraction is essential for generating and regenerating the gas-exchange surface. J. Clin. Investig. 130, 2859–2871 (2020).
Herring, M. J., Putney, L. F., Wyatt, G., Finkbeiner, W. E. & Hyde, D. M. Growth of alveoli during postnatal development in humans based on stereological estimation. Am. J. Physiol. Lung Cell Mol. Physiol. 307, 338–344 (2014).
Thébaud, B. et al. Bronchopulmonary dysplasia. Nat. Rev. Dis. Primers 5, 78 (2019).
Hilgendorff, A., Reiss, I., Ehrhardt, H., Eickelberg, O. & Alvira, C. M. Chronic lung disease in the preterm infant. Lessons learned from animal models. Am. J. Respir. Cell Mol. Biol. 50, 233–245 (2014).
Domingo-Gonzalez, R. et al. Diverse homeostatic and immunomodulatory roles of immune cells in the developing mouse lung at single cell resolution. Elife 9, 890. https://doi.org/10.7554/eLife.56890 (2020).
van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Tsukui, T. et al. Collagen-producing lung cell atlas identifies multiple subsets with distinct localization and relevance to fibrosis. Nat. Commun. 11, 1920 (2020).
Xie, T. et al. Single-cell deconvolution of fibroblast heterogeneity in mouse pulmonary fibrosis. Cell Rep. 22, 3625–3640 (2018).
Desmoulière, A., Geinoz, A., Gabbiani, F. & Gabbiani, G. Transforming growth factor-beta 1 induces alpha-smooth muscle actin expression in granulation tissue myofibroblasts and in quiescent and growing cultured fibroblasts. J. Cell Biol. 122, 103–111 (1993).
Winkler, E. A., Bell, R. D. & Zlokovic, B. V. Pericyte-specific expression of PDGF beta receptor in mouse models with normal and deficient PDGF beta receptor signaling. Mol. Neurodegener. 5, 32 (2010).
Zanini, F. et al. Phenotypic diversity and sensitivity to injury of the pulmonary endothelium during a period of rapid postnatal growth. BioRxiv. https://doi.org/10.1101/2021.04.27.441649 (2021).
Hurskainen, M. et al. Single cell transcriptomic analysis of murine lung development on hyperoxia-induced damage. Nat. Commun. 12, 1565 (2021).
Tabula Muris Consortium. A single-cell transcriptomic atlas characterizes ageing tissues in the mouse. Nature 583, 590–595 (2020).
Tabula Muris Consortium. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature. https://doi.org/10.1038/s41586-018-0590-4 (2018).
Reyfman, P. A. et al. Single-cell transcriptomic analysis of human lung provides insights into the pathobiology of pulmonary fibrosis. Am. J. Respir. Crit. Care Med. 199, 1517–1536 (2019).
Liu, X. et al. Definition and signatures of lung fibroblast populations in development and fibrosis in mice and men. BioRxiv. https://doi.org/10.1101/2020.07.15.203141 (2020).
Negretti, N. M. et al. A single-cell atlas of mouse lung development. Development 148, 512. https://doi.org/10.1242/dev.199512 (2021).
Traag, V. A., Waltman, L. & van Eck, N. J. From Louvain to Leiden: Guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Polański, K. et al. BBKNN: Fast batch alignment of single cell transcriptomes. Bioinformatics 36, 964–965 (2020).
Zepp, J. A. et al. Distinct mesenchymal lineages and niches promote epithelial self-renewal and myofibrogenesis in the lung. Cell 170, 1134–1148 (2017).
Townsley, M. I. Structure and composition of pulmonary arteries, capillaries, and veins. Compr. Physiol. 2, 675–709 (2012).
Danopoulos, S., Bhattacharya, S., Mariani, T. J. & Al, A. D. Transcriptional characterisation of human lung cells identifies novel mesenchymal lineage markers. Eur. Respir. J. 55, 1. https://doi.org/10.1183/13993003.00746-2019 (2020).
Guo, M. et al. Single cell RNA analysis identifies cellular heterogeneity and adaptive responses of the lung at birth. Nat. Commun. 10, 37 (2019).
Klar, A., Baldassare, M. & Jessell, T. M. F-spondin: A gene expressed at high levels in the floor plate encodes a secreted protein that promotes neural cell adhesion and neurite extension. Cell 69, 95–110 (1992).
Combs, M. D. et al. Microfibril-associated glycoprotein 2 (MAGP2) loss of function has pleiotropic effects in vivo. J. Biol. Chem. 288, 28869–28880 (2013).
Ross, M. D. et al. Podocan, a novel small leucine-rich repeat protein expressed in the sclerotic glomerular lesion of experimental HIV-associated nephropathy. J. Biol. Chem. 278, 33248–33255 (2003).
Tomoeda, M. et al. PLAP-1/asporin inhibits activation of BMP receptor via its leucine-rich repeat motif. Biochem. Biophys. Res. Commun. 371, 191–196 (2008).
Kalamajski, S., Aspberg, A., Lindblom, K., Heinegård, D. & Oldberg, A. Asporin competes with decorin for collagen binding, binds calcium and promotes osteoblast collagen mineralization. Biochem. J. 423, 53–59 (2009).
Rada, J. A., Cornuet, P. K. & Hassell, J. R. Regulation of corneal collagen fibrillogenesis in vitro by corneal proteoglycan (lumican and decorin) core proteins. Exp. Eye Res. 56, 635–648 (1993).
Lucariello, A. et al. Small leucine rich proteoglycans are differently distributed in normal and pathological endometrium. In Vivo 29, 217–222 (2015).
Sugden, W. W. et al. Endoglin controls blood vessel diameter through endothelial cell shape changes in response to haemodynamic cues. Nat. Cell Biol. 19, 653–665 (2017).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Muglia, L., Jacobson, L., Dikkes, P. & Majzoub, J. A. Corticotropin-releasing hormone deficiency reveals major fetal but not adult glucocorticoid need. Nature 373, 427–432 (1995).
Emanuel, R. L., Torday, J. S., Asokananthan, N. & Sunday, M. E. Direct effects of corticotropin-releasing hormone and thyrotropin-releasing hormone on fetal lung explants. Peptides 21, 1819–1829 (2000).
Jobe, A. H. & Ikegami, M. Surfactant homeostasis in corticotropin-releasing hormone deficiency in adult mice. Am. J. Respir. Crit. Care Med. 158, 995–997 (1998).
Cohen, M. et al. Lung single-cell signaling interaction map reveals basophil role in macrophage imprinting. Cell 175, 1031-1044.e18 (2018).
Seckl, J. R. & Walker, B. R. Minireview: 11beta-hydroxysteroid dehydrogenase type 1- a tissue-specific amplifier of glucocorticoid action. Endocrinology 142, 1371–1376 (2001).
Matsson, H. et al. Alpha-cardiac actin mutations produce atrial septal defects. Hum. Mol. Genet. 17, 256–265 (2008).
Sheikh-Hamad, D. et al. Stanniocalcin-1 is a naturally occurring L-channel inhibitor in cardiomyocytes: Relevance to human heart failure. Am. J. Physiol. Heart Circ. Physiol. 285, H442–H448 (2003).
Higuchi, M. et al. PRRX1- and PRRX2-positive mesenchymal stem/progenitor cells are involved in vasculogenesis during rat embryonic pituitary development. Cell Tissue Res. 361, 557–565 (2015).
Han, L. et al. Osr1 functions downstream of Hedgehog pathway to regulate foregut development. Dev. Biol. 427, 72–83 (2017).
Yang, J., Li, X. & Morrell, N. W. Id proteins in the vasculature: From molecular biology to cardiopulmonary medicine. Cardiovasc. Res. 104, 388–398 (2014).
Xie, T. et al. Transcription factor TBX4 regulates myofibroblast accumulation and lung fibrosis. J. Clin. Investig. 126, 3063–3079 (2016).
Narvaez Del Pilar, O., Gacha Garay, M. J. & Chen, J. Three-axis classification of mouse lung mesenchymal cells reveals two populations of myofibroblasts. Development 149, 81. https://doi.org/10.1242/dev.200081 (2022).
Warner, B. B., Stuart, L. A., Papes, R. A. & Wispé, J. R. Functional and pathological effects of prolonged hyperoxia in neonatal mice. Am. J. Physiol. 275, L110–L117 (1998).
Ito, R., Barnes, E. A., Che, X., Alvira, C. M. & Cornfield, D. N. SM22α cell-specific HIF stabilization mitigates hyperoxia-induced neonatal lung injury. Am. J. Physiol. Lung Cell Mol. Physiol. 323, 129–141 (2022).
Liu, M. et al. Transforming growth factor-induced protein promotes NF-κB-mediated angiogenesis during postnatal lung development. Am. J. Respir. Cell Mol. Biol. 64, 318–330 (2021).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. Cell PhoneDB: Inferring cell-cell communication from combined expression of multi-subunit ligand-receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Hellström, M., Kalén, M., Lindahl, P., Abramsson, A. & Betsholtz, C. Role of PDGF-B and PDGFR-beta in recruitment of vascular smooth muscle cells and pericytes during embryonic blood vessel formation in the mouse. Development 126, 3047–3055 (1999).
Choudhary, M. et al. Tumor-induced loss of mural Connexin 43 gap junction activity promotes endothelial proliferation. BMC Cancer 15, 427 (2015).
Weckbach, L. T. et al. Midkine acts as proangiogenic cytokine in hypoxia-induced angiogenesis. Am. J. Physiol. Heart Circ. Physiol. 303, H429–H438 (2012).
Li, L.-C., Li, J. & Gao, J. Functions of galectin-3 and its role in fibrotic diseases. J. Pharmacol. Exp. Ther. 351, 336–343 (2014).
Hirani, D. et al. Macrophage-derived IL-6 trans-signaling as a novel target in the pathogenesis of bronchopulmonary dysplasia. Eur. Respir. J. https://doi.org/10.1183/13993003.02248-2020 (2021).
Muglia, L. J. et al. Proliferation and differentiation defects during lung development in corticotropin-releasing hormone-deficient mice. Am. J. Respir. Cell. Mol. Biol. 20, 181–188 (1999).
Li, R. et al. Pdgfra marks a cellular lineage with distinct contributions to myofibroblasts in lung maturation and injury response. Elife 7, 36865. https://doi.org/10.7554/eLife.36865 (2018).
Armulik, A. et al. Pericytes regulate the blood–brain barrier. Nature 468, 557–561 (2010).
Hammes, H.-P. et al. Pericytes and the pathogenesis of diabetic retinopathy. Diabetes 51, 3107–3112 (2002).
Yee, M. et al. Neonatal hyperoxia depletes pulmonary vein cardiomyocytes in adult mice via mitochondrial oxidation. Am. J. Physiol. Lung Cell Mol. Physiol. 314, 846–859 (2018).
Folmsbee, S. S. & Gottardi, C. J. Cardiomyocytes of the heart and pulmonary veins: Novel contributors to asthma? Am. J. Respir. Cell Mol. Biol. 57, 512–518 (2017).
Zanini, F. et al. Developmental diversity and unique sensitivity to injury of lung endothelial subtypes during postnatal growth. iScience 26, 106097 (2023).
Wolterink-Donselaar, I. G., Meerding, J. M. & Fernandes, C. A method for gender determination in newborn dark pigmented mice. Lab. Anim. 38, 35–38 (2009).
Dobin, A. et al. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Putri, G. H., Anders, S., Pyl, P. T., Pimanda, J. E. & Zanini, F. Analysing high-throughput sequencing data in Python with HTSeq 2.0. Bioinformatics 38, 2943. https://doi.org/10.1093/bioinformatics/btac166 (2022).
Köster, J. & Rahmann, S. Snakemake—A scalable bioinformatics workflow engine. Bioinformatics 28, 2520–2522 (2012).
Pedregosa, F. et al. Scikit-learn: Machine learning in python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Clifford, P. S. et al. Spatial distribution and mechanical function of elastin in resistance arteries. Arterioscler. Thromb. Vasc. Biol. 31, 2889–2896 (2011).
Stirling, D. R., Carpenter, A. E. & Cimini, B. A. Cell profiler analyst 3.0: Accessible data exploration and machine learning for image analysis. Bioinformatics 37, 3992–3994 (2021).
Acknowledgements
The authors thank Sai Saroja Kolluru (Stanford University) for assistance with library submission to the Chan Zuckerberg Biohub, Yuan Xue (Stanford University) for assistance with the initial single cell RNA-seq data acquisition, Astrid Gillich for technical support with the RNAscope experiments, and Maya Kumar for providing hydrazide. They also thank the Stanford Shared FACS Facility, Lisa Nichols, Meredith Weglarz, and Tim Knaak for assistance with the flow cytometry instrumentation and antibody panel design. Flow cytometry data was collected on an instrument in the Stanford Shared FACS Facility obtained using NIH S10 Shared Instrument Grant (S10RR027431-01).
Funding
This work was supported by National Institutes of Health grants HL122918 (CMA), HL 154002 (CMA), HL155828 (CMA), HD092316 (CMA, DNC), HL160018 (CMA, DNC) the Stanford Maternal Child Health Institute Tashia and John Morgridge Faculty Scholar Award (CMA), the Crandall Endowed Faculty Scholar Award (CMA), the Karam Family Fund (DNC), the Stanford Center of Excellence in Pulmonary Biology (DNC), Bill and Melinda Gates Foundation (SRQ), the Chan Zuckerberg Biohub (DNC and SRQ), the Ernest and Amelia Gallo Endowed Fellowship (NES), and the Chan Zuckerberg Biohub Physician-Scientist Fellowship (NES). The funding was provided by National Heart, Lung, and Blood Institute (060784, 5T32HL129970).
Author information
Authors and Affiliations
Contributions
F.Z. designed and performed experiments and wrote the main manuscript. X.C. performed experiments, prepared figures. C.K. Designed experiments, performed data analysis, prepared figures. N.S. performed experiments and prepared a figure. P.K. Performed analysis and prepared figures. Y.X. Performed analysis. M.L. Performed experiments. R.D. Performed experiments and designed experiments. R.J. Performed experiments. S.Q. Assisted with experimental design, reviewed the manuscript. C.A. Designed experiments, performed analysis, draft components and reviewed the manuscript. D.C. Designed experiments, performed analysis, draft components and reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zanini, F., Che, X., Suresh, N.E. et al. Hyperoxia prevents the dynamic neonatal increases in lung mesenchymal cell diversity. Sci Rep 14, 2033 (2024). https://doi.org/10.1038/s41598-023-50717-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-50717-w
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.