Abstract
The molecular basis of colorectal cancer (CRC) can guide patient prognosis and therapy. In Brazil, knowledge on the CRC mutation landscape is limited. Here, we investigated the mutation profile of 150 cancer-related genes by next-generation sequencing and associated with microsatellite instability (MSI) and genetic ancestry in a series of 91 Brazilian CRC patients. Driver mutations were found in the APC (71.4%), TP53 (56.0%), KRAS (52.7%), PIK3CA (15.4%) and FBXW7 (10.9%) genes. Overall, genes in the MAPK/ERK, PIK3/AKT, NOTCH and receptor tyrosine kinase signaling pathways were mutated in 68.0%, 23.1%, 16.5%, and 15.3% of patients, respectively. MSI was found in 13.3% of tumors, most of which were proximal (52.4%, P< 0.001) and had a high mutation burden. European genetic ancestry was predominant (median of 83.1%), followed by Native American (4.1%), Asian (3.4%) and African (3.2%). NF1 and BRAF mutations were associated with African ancestry, while TP53 and PIK3CA mutations were inversely correlated with Native American ancestry. Our study suggests that Brazilian CRC patients exhibit a mutation profile similar to other populations and identify the most frequently mutated genes, which could be useful in future target therapies and molecular cancer screening strategies.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) is the third most common type of cancer worldwide1,2. According to GLOBOCAN, over 1.8 million new colorectal cancer cases occurred in 2018. Moreover, the disease is the second cause of death by cancer worldwide2. Increases in both incidence and mortality have been observed over the last 10 years in several Europe, Latin America, and Asia countries1. The most significant increases in incidence were observed in Brazil, Costa Rica and Bulgaria1. In Brazil, the Brazilian National Cancer Institute (INCA) estimated that over 36,000 new cases were expected for 2018, ranking CRC as the third most frequent cancer among men and as the second most frequent cancer among women3.
CRC is more frequently observed in the distal colon (left colon, from splenic flexure to rectum) than in the proximal side (right colon, from the cecum to transverse colon)4. In addition to incidence differences, the tumors arising from the left and right colon are distinct in their epidemiology, biology, histology and microbial diversity4,5,6,7. Consequently, it also influences patients’ prognosis4,8.
Several lifestyles are risk factors associated with CRC, such as red and processed meat consumption, alcohol intake, smoking and body weight9. Moreover, the cumulative acquisition of genetic alterations leads to a progressive tumorigenesis process from normal to precursor lesion, culminating in a malignant tumor10. The majority (80–85%) of CRC cases are sporadic and emerge from somatic alterations in driver genes10. These alterations are linked to three main molecular groups: chromosomal instability, mismatch repair defect and methylator phenotype7,10. In addition to these classic features, the CRC Subtyping Consortium classified four consensus molecular subtypes (CMS) of CRC: CMS1, which are enriched for tumors with MSI, overexpression of DNA damage repair proteins, high immune activation and widespread hypermethylation; CMS2, which have tumors with chromosomal instability and activation of the WNT and MYC signaling pathways; CMS3, with tumors with epithelial and metabolic dysregulation; and CMS4, which have tumors with transforming growth factor-β activation, stromal infiltration and angiogenesis6. The molecular profile of CRCs is distinct between proximal and distal tumors. Proximal CRCs are associated with microsatellite instability (MSI) and activating mutations on the BRAF gene, frequently harboring a high mutation burden, whereas distal CRCs are associated with chromosomal instability and are more frequently non-hypermutated and have less frequently the MSI phenotype6,7. In line with this, proximal CRCs are enriched for CMS1 tumors and, inversely, distal CRC are enriched for CMS2 tumors6.
Molecular characterization is crucial for CRC patient care and prognostic and therapeutic assessment11,12,13. Knowledge of the mutation profile of Brazilian CRC patients is limited. Therefore, the present study aimed to interrogate the mutational profile of CRCs by next-generation sequencing mutation analysis of a large panel of 150 cancer-related genes and to identify significant cancer drivers. Moreover, the molecular profile was associated with clinicopathological features and genetic ancestry markers of CRC patients.
Results
Description of the mutation profile
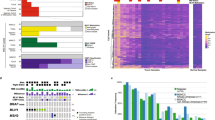
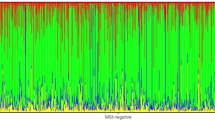
We sequenced 150 cancer-related genes in 91 colorectal tumors. The mean read depth of sequencing was 625.7x per gene and 60.7x per variant. Overall, after filtering out non-driver variants (see Material and Methods), we identified at least one somatic variant for each tumor sample with a mean of 3.6 mutations per patient (range 1–20). Thirteen tumors (14.3%) harbored mutations in single genes, 26 tumors (28.6%) in two genes, 24 tumors (26.4%) in three genes, and 28 tumors (30.8%) harbored alterations in four or more genes. Among the 150 genes sequenced, 46 showed at least one driver somatic mutation, including missense, frameshift, nonsense, in-frame, and splice mutations (54.9%, 22.6%, 21.3%, 0.9%, 0.3%, respectively). The most recurrently altered genes in our study population are shown in Fig. 1 (for a complete list of variants identified, see Supplementary Table S1). APC was the most affected gene, altered in 65 tumor samples (71.4%), followed by TP53 (51 cases, 56.0%), KRAS (48 cases, 52.7%), PIK3CA (14 cases, 15.4%) and FBXW7 (10 cases, 10.9%, Fig. 1). Below, we further describe the most affected genes and pathways.
Waterfall plot (oncoplot) of the distribution of mutations found in our tumor samples. The upper plot shows the frequency of mutation for each tumor sample. Left and right plots show the frequency of samples mutated for proximal colon (left) and distal colon (right). The central plot shows the types of mutations in each tumor sample (when the sample showed more than one mutation in the same gene, only the most deleterious type is shown). The lower part of the figure shows the tumor stage at diagnosis and microsatellite instability status (MSI) of each sample. Tumor stage: stage of the disease at the diagnosis.
APC gene
A total of 86 somatic mutations in APC were found in 65 tumors (71.4%), among which 44 harbored a single mutation, and 21 tumors harbored two mutations in the gene. The majority of the mutations were protein truncating: 52 nonsense (60.5%), 30 frameshift (34.8%), and 4 missense (4.7%) (Supplementary Fig. S1 and Supplementary Table S1). Among the mutations found, 68 were previously reported as somatic mutations in CRC (50 nonsense and 14 frameshift), and 22 were new mutations (16 frameshift, 4 missense and 2 nonsense mutations). The mutations found are spread widely throughout the coding region of the APC gene, and the most affected site was the protein position 1556, which showed frameshift mutations in 6 tumors (Supplementary Fig. S1). In addition, nine APC mutated cases were selected and PCR-followed by direct Sanger sequencing was performed, validating all mutations (Supplementary Fig. S2).
TP53 gene
Fifty-six mutations on TP53 were found affecting 51 tumors (56.0% of the tumors, Fig. 1 and Supplementary Fig. S1), among which 46 harbored a single mutation on the gene and 5 harbored two mutations. The majority of alterations in TP53 were missense mutations (40, 71.4%), followed by nonsense (7 mutations, 12.5%), frameshift (5 mutations, 8.9%), in-frame (3 mutations, 5.4%) and splice region (1 mutation, 1.8%) (Supplementary Fig. S1 and Supplementary Table S1). Among the mutations found, 53 were previously reported as somatic mutations in CRC (40 missense, 7 nonsense, 3 frameshift, 2 in-frame, and 1 splice region mutations), and 3 were new mutations, including 2 frameshift and 1 in-frame mutation. Most mutations were localized on the DNA-binding domain (only one mutation was found outside this domain), and the most frequently altered site was protein position 273, with 6 missense mutations (Supplementary Fig. S1). In addition, nine TP53 mutated cases were selected and PCR-followed by direct Sanger sequencing was performed, validating all mutations (Supplementary Fig S2).
MAPK-ERK signaling pathway alterations
The MAPK-ERK signaling pathway had at least one mutated gene in 62 tumors (68%). The most frequently altered gene in the MAPK/ERK pathway was KRAS, which showed 48 missense mutations, affecting 52.7% of the tumors. All KRAS mutations affected hotspots of the gene at protein sites 12 (33 mutations, 68.7% of the KRAS mutations), 13 (8 mutations, 16.7%), 146 (4 mutations, 8.3%), 61 (2 mutation, 4.2%), and 117 (1 mutation, 2.1%) (Fig. 2, and Supplementary Table S1). All mutations on codons 12/13 and codon 61 of the KRAS gene were confirmed by the cobas® KRAS mutation test. The NF1 gene showed 10 somatic mutations in 8 tumor samples (8.8%), three of which were already known mutations (2 missense and 1 nonsense mutations), four were novel missense mutations, and three were novel frameshift mutations. The mutations were spread throughout the coding region of the gene. All truncating protein mutations in NF1 were mutually exclusive with KRAS, and four tumors had missense mutations on both KRAS and NF1 (Fig. 2a). The BRAF gene was mutated in 8 tumor samples (8.8%), showing 8 missense somatic mutations mutually exclusive with KRAS, all previously reported as somatic mutations in CRC. The p.Val600Glu mutation (p.V600E) was the most prevalent mutation, occurring in 6 tumors (75% of all mutations on the BRAF gene). The two variants, different from V600E mutations, also affected the kinase domain of the BRAF protein (p.Asp594Glu and p.Gly469Val). Mutations in the NRAS gene were found in 4 tumor samples (4.4%), 3 at codon 12 (75%) and one at codon 61. NRAS mutations were mutually exclusive with KRAS, NF1 and BRAF mutations. Finally, the ARAF gene had one previously known missense mutation that was located outside of the main domains of the protein (Fig. 2 and Supplementary Table S1).
PI3K-AKT signaling pathway alterations
Genes in the PI3K-AKT signaling pathway were altered in 21 tumors (23.1%). The most frequently altered gene in the PI3K-AKT signaling pathway was PIK3CA, with alterations in 14 tumors (15.4%, Fig. 3a), all harboring already known somatic mutations. The PIK domain of the protein harbored most of the mutations found in PIK3CA, 6 mutations at codon 545 (42.9% of all PIK3CA mutations), 2 mutations at codon 542 (14.3%), and 1 mutation at codon 546 (7.1%). The p85-binding domain of the protein harbored 1 mutation (7.1%), and the other 4 (28.6%) mutations affected sites outside the main domains of the protein (Fig. 3a). The PTEN gene was mutated in 4.4% of the tumors (Fig. 3), showing one previously reported missense mutation and one frameshift mutation, along with two novel frameshift and nonsense mutations. The MTOR gene was mutated in 2 tumors (2.2%, Fig. 3), showing two previously described missense mutations in CRC, one affecting the PI3K/PI4K domain and one outside the main domains of the protein. AKT1 was mutated in 1 tumor (1.1%, Fig. 3) and showed one already known missense mutation affecting the Pleckstrin homology (PH) domain of the protein.
Notch signaling pathway alterations
The Notch signaling pathway was altered in 15 tumors (16.5%). FBXW7 was the most affected gene in this pathway, mutated in 10 tumors (10.9%, Fig. 4), showing nine already known mutations (7 missense and 2 frameshift mutations) and one novel somatic missense mutation. Most of the FBXW7 mutations (7, 70%) affected one of the repeated beta-transducin domains. One mutation (10%) affected the F-box-like domain, and 2 mutations were localized outside the main domains of the protein (Fig. 4b). The NOTCH2 gene was altered in 3 tumors (3.3%, Fig. 4), all harboring novel missense mutations located at one of the repeats of the EGF-like domain (2 mutations) and one at the ankyrin repeat. The NOTCH1 gene was altered in three tumors (3.3%, Fig. 4) with 4 mutations, three already known mutations (2 missense and one nonsense mutation) and one novel missense mutation. Two NOTCH1 mutations were localized at one of the ankyrin repeats and two outside of the main domains of the protein. Finally, the CREBBP gene was altered in 1 tumor (1.1%, Fig. 4), showing one already known frameshift mutation localized outside the main domains of the protein.
Receptor tyrosine kinase alterations
The receptor tyrosine kinase (RTK) family showed 9 altered genes (Fig. 5). ROS1 was altered in 3 tumors (3.3%, Fig. 5), showing one previously described missense mutation in the protein tyrosine kinase domain and two novel missense mutations outside the main domains of the protein. The ERBB4 gene was altered in 3 tumors (3.3%, Fig. 5), showing one novel missense mutation in the furin-like cysteine rich region of the protein, one previously described missense mutation in the protein tyrosine kinase domain, and one previously described missense mutation outside the main domains of the protein. ERBB3 was altered in 3 tumors (3.3%, Fig. 5), showing 1 already known missense mutation in the furin-like cysteine rich region of the protein and 2 novel missense mutations in growth factor receptor domain IV. The EGFR gene was altered in 2 tumors (2.2%, Fig. 5), showing 2 known missense mutations in the protein tyrosine kinase domain. The RET gene was mutated in 1 tumor (1.1%, Fig. 5), showing two novel mutations in the protein tyrosine kinase domain. The NTRK1 gene was mutated in 1 tumor (1.1%, Fig. 5), showing one novel missense mutation outside the main domains of the protein. The FLT3 gene was mutated in 1 tumor (1.1%, Fig. 5), showing one novel missense mutation in the protein tyrosine kinase domain. The FLT4 gene was altered in 1 tumor (1.1%, Fig. 5), showing one already known missense mutation in the immunoglobulin I-set domain. Finally, the ALK gene was also altered in 1 tumor (1.1%, Fig. 5) and showed one previously described missense mutation in the protein tyrosine kinase domain.
MSI status and its association with tumor mutation burden
We assessed the MSI status of tumor samples by multiplex PCR. Seventy-eight did not exhibit microsatellite instability (MSI-negative, 86.7%), and twelve tumors showed microsatellite instability (MSI-positive, 13.3%). The mean mutation burden of tumors that were MSI-positive was higher than MSI-negative tumors (7.1 ± 1.4 and 3.1 ± 0.2, respectively, P = 0.002, Supplementary Fig. S3). Tumors with MSI-positive status also showed a higher frequency of frameshift mutations (49.4% versus 13.2% in MSI-negative tumors, P < 0.001) and a lower frequency of missense (43.5% vs 59.1%, P = 0.01) and nonsense mutations (5.9% versus 26.4%, P < 0.001, Supplementary Fig. S3).
The profile of somatic mutations between tumors with and without MSI was also distinct. Somatic mutations of mismatch repair (MMR) genes were found only in MSI-positive tumors, which showed four frameshift mutations on MSH6 and one nonsense in MLH1. Likewise, TGFBR2, NOTCH2, and STAT5B genes were exclusively mutated in MSI-positive tumors (P < 0.001, P = 0.002 and P = 0.002, respectively, Table 1). Other genes also showed mutations only in MSI-positive tumors; however, there was no significant difference (Supplementary Table S2). MSI-positive tumors also had higher rates of BRAF mutations (41.7% vs 3.8%, P < 0.001), ATM mutations (25% vs 3.8%, P = 0.029), BLM mutations (50% vs 1.3%, P < 0.001), NOTCH1 mutations (16.7% vs 1.3%, P = 0.046), CHEK2 mutations (16.7% vs 1.3%, P = 0.046) and ROS1 mutations (16.7% vs 0.3%, P = 0.046, Table 1).
In contrast, MSI-negative tumors showed a higher frequency of APC (75.6% vs 41.7%, p = 0.034) and TP53 mutations (65.4% vs 8.3%, p < 0.001, Table 1), and some genes were exclusively altered in MSI-negative tumors, but without statistical significance (Supplementary Table S2).
Differences between proximal and distal colon
Of the 91 tumor samples sequenced, 71 (78%) were from the distal colon, and 20 (22%) were from the proximal colon. The proximal colon had a higher frequency of MSI-positive tumors than the distal colon (50% versus 2.9%, P < 0.001). We found significant differences between the mutation profile of tumors from the proximal and distal colon (Fig. 1 and Table 1). Proximal colon tumors showed higher mutation frequencies than distal tumors on TGFBR2 (30.0% vs 2.8%, P < 0.001), ATM (20.0% vs 2.8%, P = 0.020), BLM (25.0% vs 2.8%, P = 0.005), and PTEN (15.0% vs 1.4%, P = 0.032) genes. BRAF mutations were exclusively found in tumors from the proximal colon (40.0% vs 0.0%, P < 0.001), as well as mutations in the EGFR, JAK1, MAP2K4, MAP3K1, NBN, ALK genes (each of these mutated in 2 tumors), and BRCA1, BRIP1, CREBBP, FLT3, FLT4, MLH1, MPL and NT5C2 genes (each of these mutated in 1 tumor).
In contrast, the distal colon showed a higher frequency than the proximal colon of APC (77.5% vs 50.0%, P = 0.025), TP53 (67.6% vs 20.0%, P < 0.001) and KRAS (63.4% vs 15.0%, P < 0.001) gene mutations (Table 1). Mutations in NRAS (mutated in 4 tumors), ERBB3 (mutated in 3 tumors), RET, AKT1, AR, ARAF, BARD1, CDKN2A, GATA2, IDH2 MRE11A, NTRK1, PALB2 and SMO genes (each of these mutated in 1 tumor) were exclusively found in the distal colon.
Mutation profile and genetic ancestry markers
We further assessed the ancestry of the patients by a panel of ancestry informative markers (AIMs) to estimate the proportion of ancestral background. The proportion of European genetic ancestry was predominant in our study population (median 83.1%, interquartile range 76.6% and 87.8%), followed by Native American (median 4.1%, interquartile range 2.9% and 7.2%), Asian (median 3.4%, interquartile range 2.8% and 5.1%) and African (median 3.2%, interquartile 2.1% and 7.9%) (Supplementary Fig. S4).
Patients with the highest proportion of African ancestry (those with African ancestry proportions higher than 7.9%) had more frequent NF1 and BRAF mutations when compared with patients with intermediate and lowest proportions of African ancestry (P = 0.045 and 0.031, respectively, Supplementary Table S3). Patients with the highest proportion of Asian ancestry (those with a proportion of Asian ancestry higher than 5.1%) also had more NF1 mutations than those patients with intermediate and low Asian ancestry (P = 0.037, Supplementary Table S3). Patients with the highest proportion of Native American ancestry (those with a proportion of Native American ancestry higher than 7.2%) had fewer TP53 and PIK3CA mutations than those patients with the intermediate and the lowest proportion of Native American ancestry (P = 0.002 and 0.024, respectively, Supplementary Table S3).
Discussion
In this study, we investigated mutations within 150 selected oncogenes and tumor suppressor genes in a Brazilian cohort. To the best of our knowledge, this is the largest mutation profile study in this population. Overall, our results showed that our population had a similar mutation profile to other populations (Table 2), with alterations in canonical driver genes, such as APC, KRAS, TP53, PIK3CA, and FBXW7. The differences between the proximal and distal colon were also highlighted in Brazilian patients, and the admixed features of our population were revealed by ancestry markers.
APC mutations are a common event in CRC, being an early and a general event for tumorigenesis progression14. Here, we showed that APC was the most frequently mutated gene, similar to several other studies demonstrated in different populations7,15,16,17 (Table 2). Additionally, as previously reported, we found that APC mutations were more frequent in distal colon and MSI-negative tumors6,16.
TP53 is a ubiquitous driver gene in several cancer types18, and one of the most frequently altered genes in colorectal cancer, ranging from 51.7 to 68% of the cases7,16,19,20 (Table 2). Our study population showed a similar frequency of TP53 gene alterations when compared with the TCGA frequency and studies in other populations7,21, showing the highest frequency in non-hypermutated tumors (MSI-negative tumors) and in distal colon tumors7,17,22,23.
The MAPK-ERK signaling pathway is significantly altered in CRCs and other tumors18. This pathway is constitutively activated by alterations in several driver genes, including KRAS, BRAF, and NRAS7. We observed that 68% of cases had a mutation in at least one gene in the MAPK-ERK pathway, in accordance with TCGA data (59.3%)7,17. Individually, we observed a high frequency of KRAS mutations (52.7%) that was slightly higher than the one reported in TCGA data (42%) and those found in other populations15,17,21,24 (Table 2). Moreover, previous studies on the prevalence of KRAS mutations carried out in Brazilian populations showed a lower frequency (31.9–49.2%) of mutations25,26. The higher frequency observed in the present study in comparison to those observed in other Brazilian studies may be associated with the higher sensitivity and coverage of the NGS approach used in our study and the high prevalence of early-stage and localized disease cases included in our study (78% stages 0, I and II). In agreement with other studies, most mutations in the KRAS gene were found at protein residues 12 and 13, with predominant Gly12Asp mutations11,27. Epidemiological and molecular observations showed that the proximal and distal colon exhibited distinct features. KRAS mutations are slightly higher in the proximal colon20,29,,28,29,,29. However, in our study, a predominance of KRAS mutations was observed in tumors from the distal colon. We also found a high mutation frequency in the NF1 gene (Table 2), and interestingly, this frequency was associated with a high proportion of Asian ancestry, corroborating the known higher frequency of NF1 mutations in the Asian population30.
BRAF mutations are reported in less than 10% of cases (Table 2) and have been related to MSI-positive and proximal colon tumors in several studies6,7,17,20. Here, we confirmed these observations and found a similar overall frequency of BRAF mutations, including the predominance of activating V600E type mutation31. Some non-V600E mutations, such as mutations at the D594 site, were shown to have either impaired or complete loss of kinase activity32, although constitutive activation of MAPK-ERK by these mutations has been shown to coincide with other mechanisms of activation, such as NF1 deletions33. Non-V600E, such as the G469V mutation found here, has high kinase activity but acts in a different way than the V600E mutation33. The NRAS gene showed a lower frequency of mutation compared to the TCGA data, but it was similar to other large-scale studies (Table 2). Moreover, the significant pattern of mutually exclusivity of KRAS, BRAF and NRAS mutations found here has been observed in several studies7,15,17,34.
The PI3K pathway is another common pathway altered in CRC, leading to cell survival when activated7,10,18. Data from TCGA showed that genes from the PI3K signaling pathway were altered in approximately 25.9% of patients7,17, similar to the frequency reported in our study (23.1%). The PIK3CA gene seems to be the principal gene mutated in this pathway in CRCs7,15, showing activating mutations mainly on the PI3Ka domain (helicase), in addition to other already known activating mutations, such as H1047R and N345K/S found in the present study7,35. The PTEN gene is known to revert the action of PI3Ks36, and its inactivation is observed at a lower frequency in CRCs7 (1.5–9.3%, Table 2) by mutation7,37, as reported in our study (4.4%). The association between PTEN mutations and proximal colon localization has been observed previously23. Although MTOR was found to be mutated in CRCs with a range of 7.6–13.1% in large cohort studies (Table 2), these mutations are poorly characterized38. Mutations in MTOR site S2215, as we found here, seems to be the most frequently altered site across diverse cancer types and has been shown to increase the activity of mTOR protein38. The other MTOR (T2444M) mutation observed here has not been well characterized, but it affects the C-terminal region related to mTOR activation38. A low frequency of AKT1 has been found across large cohort studies (Table 2), and the E17K mutation observed here has been related to the constitutive activation of AKT1 protein39.
We found that 15.4% of tumors harboring missense mutations in members of the RTK family were lower than the frequency observed in TCGA data (22.3%)7,17. Although activating events in some RTK proteins can be observed by rearrangements in CRCs, such as ROS140, NTRK140, ALK40, and RET41, missense mutations in these genes are found across several studies (Table 2), but the oncogenic potential of such mutations is not completely understood in CRC. RET missense mutations promote tumor progression in thyroid cancer42 and are shown to reduce apoptosis in colorectal cancer cell lines42. Interestingly, ALK, ROS1, and NTRK1 missense mutations have been observed in solid tumors resistant to tyrosine receptor kinase inhibitors43,44,45.
We found known activating mutations present in the protein kinase domain EGFR T725M17,46 and P848L47, as well as in FLT317,46. Other mutations affecting the protein kinase domain of the ROS1, ERBB4, and ALK genes were also observed, but their role in oncogeneses is unknown47. Mutations that occur outside the protein kinase domain also play a role in the activation of RTKs, such as ERBB3 mutations, which promote the growth of colonic cells48 and ERBB4 mutations, which promote an increase in the kinase activity and transformation ability in melanoma cells49.
The Notch signaling pathway is important in maintaining the proliferative compartment of intestinal crypts50 and has been linked to tumor progression51. Here, we observed gene alterations in this pathway in 16.5% of the tumors, which was slightly lower when compared to the frequency observed on TCGA data for the same pathway (20.1%)7,17. FBXW7 is one of the regulators of Notch protein52,53, and its loss seems to lead to the activation of this pathway54. The frequency of FBXW7 mutations in our population was compared with those reported in other populations (Table 2)55. We observed mutations affecting hotspots of FBXW7, such as residues 465 and 505, that are important sites for substrate binding56, and frameshift mutations that lead to premature termination of the protein and have also been found affecting this gene7,15,17. The frequency of alterations in NOTCH1 and NOTCH2 genes is variable (Table 2), and missense activating mutations in both NOTCH1 and NOTCH2 have been observed in hematological tumors57,58. The pattern of mutual exclusivity between mutations in the NOTCH2 and FBXW7 genes that we found here has not been observed in large cohort studies7,15,17.
The presence of MSI was found in 13.3% of cases, in accordance with previous studies4,6,59. In addition, MSI was associated with tumors in the proximal colon and with those exhibiting a higher mutation burden6,7,19. As previously reported, we observed an association between ATM7,17, BLM7,17, and TGFRB26,7,17 mutations and MSI and the proximal colon.
Epidemiological evidence suggests that ethnicity has a high impact on cancer incidence and mortality60, and differences in the prevalence of cancer driver mutations have been observed in several cancer types, including CRC61. Our population is known to be diverse62, and our results are in agreement with these observations, showing differences in the proportions of each of the main four population origins compared with previous studies62,63. Our patients with a higher African ancestry proportion had a higher frequency of BRAF mutations, contrasting with a previous study by Staudacher et al.64. We did not observe an association between the frequency of KRAS mutations and the African ancestry that was reported previously64. These disparities may reside in the fact that, although we have an admixture population study, we have a low frequency of self-declared African ancestry and a low frequency of proportion of African ancestry than observed in other population studies62,63. In contrast, the association of NF1 mutations and African ancestry has been observed previously65. Moreover, the observation that NF1 mutations were associated with the high Asian ancestry proportion was consistent with the data from ICGC30. Although ethnicity association with PIK3CA mutations has been observed in different populations65,66, this is the first study to demonstrate an inverse correlation between PIK3CA mutations and a high Native American ancestry background.
Despite the major findings, one limitation of the present study lies in the preselection of a panel of known cancer genes, and known genes related to CCR carcinogenesis were not included in the analysis, such as SMAD4 (altered in 11.6–12.9% of CRC), TTN (35.7–48.1%) and SYNE1 (21.0–48.1%)7,15,17. Moreover, using whole-genome or exome approaches, other putative CCR cancer-related genes could have been identified in the Brazilian context.
In conclusion, our study constitutes the largest mutation landscape of Brazilian colorectal cancer patients. This study paves the way for a better comprehension of the major alterations identified and could guide better-tailored therapy for colorectal cancer in the Brazilian population.
Materials and Methods
Tissue samples
Ninety-one patients diagnosed with colorectal adenocarcinomas and admitted for a surgical procedure at Barretos Cancer Hospital, Barretos, SP, Brazil, were evaluated. The main clinicopathological features are summarized in Table 3. The mean age of patients was 61 years old, and the majority of cases had early-stage and localized disease (0, I and II).
All included patients signed an informed consent form. Both tumors and blood were obtained and immediately processed and stored at −80 °C at Barretos Cancer Hospital Biobank. The present study was approved by the Barretos Cancer Hospital Institutional Review Board (Project n° 1060/2015, protocol CAAE: 51770115.6.0000.5437). All methods were performed in accordance with the relevant guidelines and regulations.
DNA isolation
Tumor DNA was isolated from fresh-frozen tissue using the QIAsymphony DNA Mini Kit following the Tissue_200 protocol for automated isolation in the QIAsymphony (QIAGEN, Hilden, Germany). DNA from leucocytes of peripheral blood was isolated using the QIAmp DNA Blood Mini Kit (QIAGEN, Hilden, Germany) following the manufacturer’s instructions. DNA quantity and quality were assessed by Qubit (Thermo Fisher Scientific, Waltham, MA, USA).
Microsatellite instability analyses
The MSI status was evaluated using a multiplex PCR comprising six quasi-monomorphic mononucleotide repeat markers (BAT-25, BAT-26, NR-21, NR-24, NR-27 and HSP110) as described previously59. The MSI analyses were performed using GeneMapper v4.1 software (Applied Biosystems), and the status was classified as stable (MSS) when none of the markers were unstable. The status was MSI-Low (MSI-L) when one of the markers was unstable and MSI-positive when two or more of the markers were unstable. MSS and MSI-L were considered MSI-negative59.
Mutation profile
The mutation profile of a commercial panel of 150 cancer-related genes (for a list of genes, see Supplementary Information) was conducted at the Mendelics Genetics company (São Paulo, SP, Brazil, (https://www.mendelics.com/oncologia/). The panel analyzed all coding regions of the 150 cancer-related genes. For sequencing, paired tumor and blood DNA libraries were prepared using the Nextera Rapid Capture Custom Enrichment Kit (Illumina, San Diego, CA, USA). Libraries were quantified using a Qubit fluorometer (Thermo Fisher Scientific, Waltham, MA, USA), and their quality was evaluated using an Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA). Cluster generation and sequencing were performed on the Illumina HiSeq. 4000 following the manufacturer’s instructions. Paired-end reads from Illumina sequencing were processed by script bcl2fastq (v. 2.17.1.14) and aligned against the human genome reference build GRCh37 using Burrows-Wheeler Aligner (BWA, version 0.7.13)67. The somatic variants were called by a VarScan2 algorithm68. The variants with artifacts due to indel reads at their position or less than 10% or more than 90% of variant supporting reads on one strand were removed. The variants were further filtered to remove those with fewer than 10 reads covering the variant and with less than 5% variant allele frequency. The annotation of variants was performed using Ensembl Variant Effect Predictor (VEP)69.
To identify driver mutations on tumors, we used the Cancer Genome Interpreter – CGI70. After the classification of mutations by the CGI, we excluded mutations that were not classified as cancer driver mutations or not predicted as a driver by the OncoMut algorithm that CGI used. Therefore, mutations identified as polymorphisms (high allele frequency) or those predicted as neutral or passenger for oncogenesis, and those found in DNA sequences outside coding regions were excluded.
We further validated mutations on codons 12/13 and codon 61 of the KRAS gene using the cobas® KRAS Mutation Test (F. Hoffmann-La Roche, Basel, Switzerland) following the manufacturer’s recommendations. We also performed PCR-followed by direct Sanger sequencing for a subset of samples to confirm mutations in the APC and TP53 genes. The PCR and Sanger sequencing conditions were previously reported by our group71,72
Genetic ancestry markers
The genetic ancestry of patients was determined by 46 autosomal ancestry informative markers (AIMs), consisting of insertion and deletion polymorphisms and using the genetic data of the Human Genome Diversity Project – center d’Etude du Polymorphisme Humain (HGDP-CEPH), according to Pereira et al.63. We classified the proportion of ancestry by tertiles as high, intermediary and low for each of the main populations: African, European, Asian and Native American73.
Statistical analyses
Chi-square and Fisher’s exact tests were used to set differences between mutated and nonmutated genes and patient characteristics. Mutation types and mutation burden differences between MSI-positive and MSI-negative tumors were determined by chi-square and Mann-Whitney tests, respectively. All statistical analyses were performed using SPSS software (v.20) and R 3.5.0 software. Mutation mapper figures were generated using cBioPortal. Oncoplots and ancestry charts were generated using GenVisR and ggplot2 packages (versions 1.14 and 3.1.0, respectively) in R 3.5.0 software.
Data Availability
Data that support the findings are available from Dr. Denise Peixoto Guimarães and Dr. Rui Manual Reis and are not publicly available due patient personal information. However, these data are available upon reasonable request and with permission of the Dr. Rui Manuel Reis (Scientific and Executive Director of the Molecular Oncology Research Center, Barretos Cancer Hospital).
References
Arnold, M. et al. Global patterns and trends in colorectal cancer incidence and mortality. Gut 66, 683–691 (2017).
Bray, F. et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA A J. Clin. https://doi.org/10.3322/caac.21492 (2018).
MINISTÉRIO DA SAÚDE: Instituto Nacional de Câncer José Alencar Gomes da Silva. Estimativa 2018-Incidência de câncer no Brasil. (2017).
Missiaglia, E. et al. Distal and proximal colon cancers differ in terms of molecular, pathological, and clinical features. Ann. Oncol. 25, 1995–2001 (2014).
Drewes, J. L., Housseau, F. & Sears, C. L. Sporadic colorectal cancer: Microbial contributors to disease prevention, development and therapy. Br. J. Cancer 115, 273–280 (2016).
Guinney, J. et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 21, 1350–1356 (2015).
Muzny, D. M. et al. Comprehensive molecular characterization of human colon and rectal cancer. Nature 487, 330–337 (2012).
Benedix, F. et al. Comparison of 17,641 patients with right- and left-sided colon cancer: Differences in epidemiology, perioperative course, histology, and survival. Dis. Colon Rectum 53, 57–64 (2010).
Song, M., Garrett, W. S. & Chan, A. T. Nutrients, Foods, and Colorectal Cancer Prevention. Gastroenterology 148, 1244–1260.e16 (2015).
Kuipers, E. J. et al. Colorectal cancer. Nat. Rev. Dis. Prim. 1, (2015).
Boeckx, N. et al. Prognostic and Predictive Value of RAS Gene Mutations in Colorectal Cancer: Moving Beyond KRAS Exon 2. Drugs 75, 1739–1756 (2015).
Le, D. T. et al. PD-1 Blockade in Tumors with Mismatch-Repair Deficiency. N. Engl. J. Med. 372, 2509–2520 (2015).
Sargent, D. J. et al. Defective mismatch repair as a predictive marker for lack of efficacy of fluorouracil-based adjuvant therapy in colon cancer. J. Clin. Oncol. 28, 3219–3226 (2010).
Cross, W. et al. The evolutionary landscape of colorectal tumorigenesis. Nat. Ecol. Evol. 2, 1661–1672 (2018).
Giannakis, M. et al. Genomic Correlates of Immune-Cell Infiltrates in Colorectal Carcinoma. Cell Rep. 15, 857–865 (2016).
Liu, Y. et al. Comparative Molecular Analysis of Gastrointestinal Adenocarcinomas. Cancer Cell 33, 721–735.e8 (2018).
Gao, J. et al. Integrative Analysis of Complex Cancer Genomics and Clinical Profiles Using the cBioPortal. Sci. Signal. 6, pl1–pl1 (2013).
Bailey, M. H. et al. Comprehensive Characterization of Cancer Driver Genes and Mutations. Cell 173, 371–385.e18 (2018).
Seshagiri, S. et al. Recurrent R-spondin fusions in colon cancer. Nature 488, 660–664 (2012).
Salem, M. E. et al. Comparative molecular analyses of left-sided colon, right-sided colon, and rectal cancers. Oncotarget 8, 86356–86368 (2017).
Liu, Z. et al. The landscape of somatic mutation in sporadic Chinese colorectal cancer. bioRxiv 9, 155671 (2018).
Russo, A. et al. The TP53 colorectal cancer international collaborative study on the prognostic and predictive significance of p53 mutation: Influence of tumor site, type of mutation, and adjuvant treatment. J. Clin. Oncol. 23, 7518–7528 (2005).
Loree, J. M. et al. Classifying colorectal cancer by tumor location rather than sidedness highlights a continuum in mutation profiles and consensus molecular subtypes. Clin. Cancer Res. 24, 1062–1072 (2018).
Al-Shamsi, H. O. et al. Molecular spectrum of KRAS, NRAS, BRAF, PIK3CA, TP53, and APC somatic gene mutations in Arab patients with colorectal cancer: Determination of frequency and distribution pattern. J. Gastrointest. Oncol. 7, 882–902 (2016).
Yamane, L. S. et al. KRAS and BRAF mutations and MSI status in precursor lesions of colorectal cancer detected by colonoscopy. Oncol. Rep. 1419–1426, https://doi.org/10.3892/or.2014.3338 (2014).
Gil Ferreira, C. et al. KRAS mutations: variable incidences in a Brazilian cohort of 8,234 metastatic colorectal cancer patients. BMC Gastroenterol. 14, 73 (2014).
Loupakis, F. et al. KRAS codon 61, 146 and BRAF mutations predict resistance to cetuximab plus irinotecan in KRAS codon 12 and 13 wild-type metastatic colorectal cancer. Br. J. Cancer 101, 715–721 (2009).
Rosty, C. et al. Colorectal carcinomas with KRAS mutation are associated with distinctive morphological and molecular features. Mod. Pathol. 26, 825–834 (2013).
Samowitz, W. S. et al. Relationship of Ki- ras Mutations in Colon Cancers to Tumor Location, Stage, and Survival: A Population-based Study Relationship of Ki- ras Mutations in Colon Cancers to Tumor Location,. Cancer Epidemiol. Biomarkers Prev. 9, 1193–1197 (2000).
Kasprzyk, A. et al. International Cancer Genome Consortium Data Portal–a one-stop shop for cancer genomics data. Database 2011, bar026–bar026 (2011).
Davies, H. et al. Mutations of the BRAF gene in human cancer. Nature 417, 949–954 (2002).
Wan, P. T. C. et al. Mechanism of activation of the RAF-ERK signaling pathway by oncogenic mutations of B-RAF. Cell 116, 855–867 (2004).
Yao, Z. et al. Tumours with class 3 BRAF mutants are sensitive to the inhibition of activated RAS. Nature 548, 234–238 (2017).
Malapelle, U. et al. Less frequently mutated genes in colorectal cancer: evidences from next-generation sequencing of 653 routine cases. J. Clin. Pathol. 69, 767–771 (2016).
Rosty, C. et al. PIK3CA Activating Mutation in Colorectal Carcinoma: Associations with Molecular Features and Survival. PLoS One 8, 1–9 (2013).
Vivanco, I. & Sawyers, C. L. The phosphatidylinositol 3-Kinase–AKT pathway in human cancer. Nat. Rev. Cancer 2, 489–501 (2002).
Rodon, J., Dienstmann, R., Serra, V. & Tabernero, J. Development of PI3K inhibitors: lessons learned from early clinical trials. Nat. Rev. Clin. Oncol. 10, 143–153 (2013).
Grabiner, B. C. et al. A diverse array of cancer-associated MTOR mutations are hyperactivating and can predict rapamycin sensitivity. Cancer Discov. 4, 554–563 (2014).
Carpten, J. D. et al. A transforming mutation in the pleckstrin homology domain of AKT1 in cancer. Nature 448, 439–444 (2007).
Pietrantonio, F. et al. ALK, ROS1, and NTRK rearrangements in metastatic colorectal cancer. J. Natl. Cancer Inst. 109, 1–10 (2017).
Stransky, N., Cerami, E., Schalm, S., Kim, J. L. & Lengauer, C. The landscape of kinase fusions in cancer. Nat. Commun. 5, 4846 (2014).
Mulligan, L. M. RET revisited: Expanding the oncogenic portfolio. Nat. Rev. Cancer 14, 173–186 (2014).
Russo, M. et al. Acquired Resistance to the TRK Inhibitor Entrectinib in Colorectal Cancer. Cancer Discov. 6, 36–44 (2016).
Facchinetti, F. et al. Crizotinib-resistant ROS1 mutations reveal a predictive kinase inhibitor sensitivity model for ROS1- and ALK-rearranged lung cancers. Clin. Cancer Res. 22, 5983–5991 (2016).
Gainor, J. F. et al. Molecular Mechanisms of Resistance to First- and Second-Generation ALK Inhibitors in ALK-Rearranged Lung Cancer. Cancer Discov. 6, 1118–1133 (2016).
Chakravarty, D. et al. OncoKB: A Precision Oncology Knowledge Base HHS Public Access. 1–23 (2017).
Chen, H. et al. Systematic Functional Annotation of Somatic Mutations in Cancer. Cancer Cell 33, 450–462.e10 (2018).
Jaiswal, B. S. et al. Oncogenic ERBB3 Mutations in Human Cancers. Cancer Cell 23, 603–617 (2013).
Prickett, T. D. et al. Analysis of the tyrosine kinome in melanoma reveals recurrent mutations in ERBB4. Nat. Genet. 41, 1127–1132 (2009).
Zeki, S. S., Graham, T. A. & Wright, N. A. Stem cells and their implications for colorectal cancer. Nat. Rev. Gastroenterol. Hepatol. 8, 90–100 (2011).
Zhang, Y., Li, B., Ji, Z. Z. & Zheng, P. S. Notch1 regulates the growth of human colon cancers. Cancer 116, 5207–5218 (2010).
Mu, Y., Zou, H., Chen, B., Fan, Y. & Luo, S. FAM83D knockdown regulates proliferation, migration and invasion of colorectal cancer through inhibiting FBXW7/Notch-1 signalling pathway. Biomed. Pharmacother. 90, 548–554 (2017).
Sancho, R. et al. F-box and WD repeat domain-containing 7 regulates intestinal cell lineage commitment and is a haploinsufficient tumor suppressor. Gastroenterology 139, 929–941 (2010).
Babaei-Jadidi, R. et al. FBXW7 influences murine intestinal homeostasis and cancer, targeting Notch, Jun, and DEK for degradation. J. Exp. Med. 208, 295–312 (2011).
Korphaisarn, K. et al. FBXW7 missense mutation: a novel negative prognostic factor in metastatic colorectal adenocarcinoma. Oncotarget 8, 39268–39279 (2017).
Welcker, M. & Clurman, B. E. FBW7 ubiquitin ligase: A tumour suppressor at the crossroads of cell division, growth and differentiation. Nat. Rev. Cancer 8, 83–93 (2008).
Lee, S. Y. et al. Gain-of-function mutations and copy number increases of Notch2 in diffuse large B-cell lymphoma. Cancer Sci. 100, 920–926 (2009).
Weng, A. P. et al. Activating Mutations of NOTCH1 in Human T Cell Acute Lymphoblastic Leukemia. Science (80-.). 306, 269–271 (2004).
Berardinelli, G. N. et al. Advantage of HSP110 (T17) marker inclusion for microsatellite instability (MSI) detection in colorectal cancer patients. Oncotarget 9, 28691–28701 (2018).
Wiencke, J. K. Impact of race/ethnicity on molecular pathways in human cancer. Nat. Rev. Cancer 4, 79–84 (2004).
Özdemir, B. C. & Dotto, G. P. Racial Differences in Cancer Susceptibility and Survival: More Than the Color of the Skin? Trends in Cancer 3, 181–197 (2017).
Campanella, N. C. et al. Optimization of a pentaplex panel for MSI analysis without control DNA in a Brazilian population: correlation with ancestry markers. Eur. J. Hum. Genet. 22, 875–880 (2014).
Pereira, R. et al. Straightforward inference of ancestry and admixture proportions through ancestry-informative insertion deletion multiplexing. PLoS One 7, (2012).
Staudacher, J. J. et al. Increased Frequency of KRAS Mutations in African Americans Compared with Caucasians in Sporadic Colorectal Cancer. Clin. Transl. Gastroenterol. 8, e124 (2017).
Heath, E. et al. Racial Disparities in the Molecular Landscape of Cancer. Anticancer Res. 38, 2235–2240 (2018).
Chong, M.-L. et al. Phosphatidylinositol-3-kinase pathway aberrations in gastric and colorectal cancer: Meta-analysis, co-occurrence and ethnic variation. Int. J. Cancer 134, 1232–1238 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Koboldt, D. C. et al. VarScan 2: Somatic mutation and copy number alteration discovery in cancer by exome sequencing VarScan 2: Somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576 (2012).
McLaren, W. et al. The Ensembl Variant Effect Predictor. Genome Biol. 17, 122 (2016).
Tamborero, D. et al. Cancer Genome Interpreter annotates the biological and clinical relevance of tumor alterations. Genome Med. 10, 1–8 (2018).
de Oliveira, J. C. et al. Genotype-phenotype correlation in 99 familial adenomatous polyposis patients: A prospective prevention protocol. Cancer Med. 8, 2114–2122 (2019).
Lacerda, C. F. et al. Mutational profile of TP53 in esophageal squamous cell carcinoma associated with chagasic megaesophagus. Dis. Esophagus 30, 1–9 (2017).
Lima-Costa, M. F. et al. Genomic ancestry, Self-rated health and its association with mortality in an admixed population: 10 year follow-up of the Bambui-Epigen (Brazil) cohort study of ageing. PLoS One 10, 1–14 (2015).
Acknowledgements
We are thankful to Barretos Cancer Hospital. This work was supported by the Brazilian Federal Agency for the Support and Evaluation of Graduate Education (CAPES, Brazil), the National Council for Scientific and Technological Development (CNPq, Brazil), and the Public Ministry of Labor Campinas (Research, Prevention and Education of Occupational Cancer, Brazil).
Author information
Authors and Affiliations
Contributions
W.S., D.P.G. and R.M.R. designed and wrote the main manuscript text. D.P.G. and R.M.R. supervised the study. W.S., G.N.B. and T.S. performed the experiments. M.M. performed the histopathological analysis. W.S. and A.F.E. carried out the bioinformatics analysis. M.A.O. and W.S. performed the statistical analysis. W.S., A.C.C., D.P.G., R.M.R. and A.F.E. contributed to the interpretation of the results. The manuscript was reviewed by all authors.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
dos Santos, W., Sobanski, T., de Carvalho, A.C. et al. Mutation profiling of cancer drivers in Brazilian colorectal cancer. Sci Rep 9, 13687 (2019). https://doi.org/10.1038/s41598-019-49611-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-49611-1
This article is cited by
-
Assessing the applicability and interobserver variability of tumor budding and poorly differentiated clusters in colorectal cancer
Surgical and Experimental Pathology (2024)
-
Genomic ancestry and cancer among Latin Americans
Clinical and Translational Oncology (2024)
-
Racial disparities in colorectal cancer clinicopathological and molecular tumor characteristics: a systematic review
Cancer Causes & Control (2024)
-
Proteomic analysis of low- and high-grade human colon adenocarcinoma tissues and tissue-derived primary cell lines reveals unique biological functions of tumours and new protein biomarker candidates
Clinical Proteomics (2022)
-
Signaling pathways and their potential therapeutic utility in esophageal squamous cell carcinoma
Clinical and Translational Oncology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.