Abstract
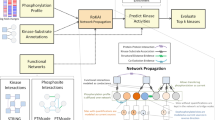
Phosphoprotein phosphatases (PPPs) execute >90% of serine/threonine dephosphorylation in cells and tissues. While the role of PPPs in cell biology and diseases such as cancer, cardiac hypertrophy and Alzheimer’s disease is well established, the molecular mechanisms governing and governed by PPPs still await discovery. Here we describe a chemical proteomic strategy, phosphatase inhibitor beads and mass spectrometry (PIB-MS), that enables the identification and quantification of PPPs and their posttranslational modifications in as little as 12 h. Using a specific but nonselective PPP inhibitor immobilized on beads, PIB-MS enables the efficient affinity-capture, identification and quantification of endogenous PPPs and associated proteins (‘PPPome’) from cells and tissues. PIB-MS captures functional, endogenous PPP subunit interactions and enables discovery of new binding partners. It performs PPP enrichment without exogenous expression of tagged proteins or specific antibodies. Because PPPs are among the most conserved proteins across evolution, PIB-MS can be employed in any cell line, tissue or organism.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Brautigan, D. L. & Shenolikar, S. Protein serine/threonine phosphatases: keys to unlocking regulators and substrates. Annu. Rev. Biochem. 87, 921–964 (2018).
Virshup, D. M. & Shenolikar, S. From promiscuity to precision: protein phosphatases get a makeover. Mol. Cell 33, 537–545 (2009).
Brautigan, D. L. Protein Ser/Thr phosphatases—the ugly ducklings of cell signalling. FEBS J. 280, 324–345 (2013).
Sun, L. et al. Inhibition of protein phosphatase 2A- and protein phosphatase 1-induced tau hyperphosphorylation and impairment of spatial memory retention in rats. Neuroscience 118, 1175–1182 (2003).
Kamat, P. K., Rai, S. & Nath, C. Okadaic acid induced neurotoxicity: an emerging tool to study Alzheimer’s disease pathology. Neurotoxicology 37, 163–172 (2013).
Sontag, J. M., Nunbhakdi-Craig, V., White, C. L., Halpain, S. & Sontag, E. The protein phosphatase PP2A/Bα binds to the microtubule-associated proteins Tau and MAP2 at a motif also recognized by the kinase Fyn: Implications for tauopathies. J. Biol. Chem. 287, 14984–14993 (2012).
Chen, W. et al. Identification of specific PP2A complexes involved in human cell transformation. Cancer Cell 5, 127–136 (2004).
Arroyo, J. D. & Hahn, W. C. Involvement of PP2A in viral and cellular transformation. Oncogene 24, 7746–7755 (2005).
Molkentin, J. D. et al. A calcineurin-dependent transcriptional pathway for cardiac hypertrophy. Cell 93, 215–228 (1998).
Lyons, S. P. et al. A quantitative chemical proteomic strategy for profiling phosphoprotein phosphatases from yeast to humans. Mol. Cell Proteomics 17, 2448–2461 (2018).
Nasa, I. et al. Quantitative kinase and phosphatase profiling reveal that CDK1 phosphorylates PP2Ac to promote mitotic entry. Sci. Signal. 13, eaba7823 (2020).
Frohner, I. E., Mudrak, I., Kronlachner, S., Schüchner, S. & Ogris, E. Antibodies recognizing the C terminus of PP2A catalytic subunit are unsuitable for evaluating PP2A activity and holoenzyme composition. Sci. Signal. 13, eaax6490 (2020).
Couzens, A. L. et al. Protein interaction network of the mammalian Hippo pathway reveals mechanisms of kinase-phosphatase interactions.Sci. Signal. 6, rs15 (2013).
St-Denis, N. et al. Phenotypic and interaction profiling of the human phosphatases identifies diverse mitotic regulators. Cell Rep. 17, 2488–2501 (2016).
Yadav, L. et al. Systematic analysis of human protein phosphatase interactions and dynamics. Cell Syst. 4, 430–444 e5 (2017).
Heroes, E. et al. The PP1 binding code: a molecular-lego strategy that governs specificity. FEBS J. 280, 584–595 (2013).
Swingle, M., Ni, L. & Honkanen, R. E. Small-molecule inhibitors of ser/thr protein phosphatases: Specificity, use and common forms of abuse. Methods Mol. Biol. 365, 23–38 (2007).
Moorhead, G. B. G., Haystead, T. A. J. & MacKintosh, C. Synthesis and use of the protein phosphatase affinity matrices microcystin-sepharose and microcystin-biotin-sepharose. Methods Mol. Biol. 365, 39–45 (2007).
Duncan, J. S. et al. Dynamic reprogramming of the kinome in response to targeted MEK inhibition in triple-negative breast cancer. Cell 149, 307–321 (2012).
Godl, K. et al. An efficient proteomics method to identify the cellular targets of protein kinase inhibitors. Proc. Natl Acad. Sci. USA 100, 15434–15444 (2003).
Bantscheff, M. et al. Quantitative chemical proteomics reveals mechanisms of action of clinical ABL kinase inhibitors. Nat. Biotechnol. 25, 1035–1044 (2007).
Kruse, U. et al. Chemoproteomics-based kinome profiling and target deconvolution of clinical multi-kinase inhibitors in primary chronic lymphocytic leukemia cells. Leukemia 25, 89–100 (2011).
Midland, A. A. et al. Defining the expressed breast cancer kinome. Cell Res. 22, 620–623 (2012).
Stuhlmiller, T. J. et al. Inhibition of lapatinib-induced kinome reprogramming in ERBB2-positive breast cancer by targeting BET family bromodomains. Cell Rep. 11, 390–404 (2015).
Cooper, M. J. et al. Application of multiplexed kinase inhibitor beads to study kinome adaptations in drug-resistant leukemia. PLoS One 8, e66755 (2013).
Johnson, G. L., Stuhlmiller, T. J., Angus, S. P., Zawistowski, J. S. & Graves, L. M. Molecular pathways: adaptive kinome reprogramming in response to targeted inhibition of the BRAF-MEK-ERK pathway in cancer. Clin. Cancer Res. 20, 2516–2522 (2014).
Klaeger, S. et al. The target landscape of clinical kinase drugs. Science 358, eaan4368 (2017).
Hughes, P. F. et al. A highly selective Hsp90 affinity chromatography resin with a cleavable linker. Bioorganic Med. Chem. 20, 3298–3305 (2012).
Ranjitkar, P., Brock, A. M. & Maly, D. J. Affinity reagents that target a specific inactive form of protein kinases. Chem. Biol. 17, 195–206 (2010).
Moorhead, G., MacKintosh, R. W., Morrice, N., Gallagher, T. & MacKintosh, C. Purification of type 1 protein (serine/threonine) phosphatases by microcystin-Sepharose affinity chromatography. FEBS Lett 356, 46–50 (1994).
Hughes, C. S. et al. Single-pot, solid-phase-enhanced sample preparation for proteomics experiments. Nat. Protoc. 14, 68–85 (2019).
Ting, L., Rad, R., Gygi, S. P. & Haas, W. MS3 eliminates ratio distortion in isobaric multiplexed quantitative proteomics. Nat. Methods 8, 937–940 (2011).
Eng, J. K., Jahan, T. A. & Hoopmann, M. R. Comet: an open-source MS/MS sequence database search tool. Proteomics 13, 22–24 (2013).
Valot, B., Langella, O., Nano, E. & Zivy, M. MassChroQ: a versatile tool for mass spectrometry quantification. Proteomics 11, 3572–3577 (2011).
R Core Team. The R Project for Statistical Computing. https://www.R-project.org/ (2019).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Yu, S. H. et al. Expanding the Perseus software for omics data analysis with custom plugins. Curr. Protoc. Bioinformatics 71, 1–29 (2020).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906 (2007).
Zecha, J. et al. TMT labeling for the masses: a robust and cost-efficient, in-solution labeling approach. Mol. Cell Proteomics 18, 1468–1478 (2019).
Kettenbach, A. N. & Gerber, S. A. Rapid and reproducible single-stage phosphopeptide enrichment of complex peptide mixtures: application to general and phosphotyrosine-specific phosphoproteomics experiments. Anal. Chem. 83, 7635–7644 (2011).
Bollen, M., Peti, W., Ragusa, M. J. & Beullens, M. The extended PP1 toolkit: designed to create specificity. Trends Biochem. Sci. 35, 450–458 (2010).
Nilsson, J. Protein phosphatases in the regulation of mitosis. J. Cell Biol. 218, 395–409 (2019).
Krystkowiak, I. & Davey, N. E. SLiMSearch: a framework for proteome-wide discovery and annotation of functional modules in intrinsically disordered regions. Nucleic Acids Res. 45, W464–W469 (2017).
Chen, M. J., Dixon, J. E. & Manning, G. Genomics and evolution of protein phosphatases. Sci. Signal. 10, eaag1796 (2017).
Uhlen, M. et al. Tissue-based map of the human proteome. Science 347, 1260419–1260419 (2015).
Nusinow, D. P. et al. Quantitative proteomics of the Cancer Cell Line Encyclopedia. Cell 180, 387–402 (2020).
Wang, D. et al. A deep proteome and transcriptome abundance atlas of 29 healthy human tissues. Mol. Syst. Biol. 15, e8503 (2019).
Xu, Y. et al. Structure of the protein phosphatase 2A holoenzyme. Cell 127, 1239–1251 (2006).
Cho, U. S. & Xu, W. Crystal structure of a protein phosphatase 2A heterotrimeric holoenzyme. Nature 445, 53–57 (2007).
Seshacharyulu, P., Pandey, P., Datta, K. & Batra, S. K. Phosphatase: PP2A structural importance, regulation and its aberrant expression in cancer. Cancer Lett. 335, 9–18 (2013).
da Cruz e Silva, E. F. & Cohen, P. T. Isolation of a cDNA likely to encode a novel Ca2+-dependent/calmodulin-stimulated protein phosphatase. Biochim. Biophys. Acta 1009, 293–296 (1989).
Klee, C. B., Ren, H. & Wang, X. Regulation of the calmodulin-stimulated protein phosphatase, calcineurin. J. Biol. Chem. 273, 13367–13370 (1998).
Rusnak, F. & Mertz, P. Calcineurin: form and function. Physiol. Rev. 80, 1483–1521 (2000).
Li, H., Rao, A. & Hogan, P. G. Interaction of calcineurin with substrates and targeting proteins. Trends Cell Biol. 21, 91–103 (2011).
Gingras, A. C. et al. A novel, evolutionarily conserved protein phosphatase complex involved in cisplatin sensitivity. Mol. Cell Proteomics 4, 1725–1740 (2005).
Kloeker, S. & Wadzinski, B. E. Purification and identification of a novel subunit of protein serine/threonine phosphatase 4. J. Biol. Chem. 274, 5339–5347 (1999).
Chen, M. X. et al. A novel human protein serine/threonine phosphatase, which possesses four tetratricopeptide repeat motifs and localizes to the nucleus. EMBO J. 13, 4278–4290 (1994).
Das, A. K., Cohen, P. W. & Barford, D. The structure of the tetratricopeptide repeats of protein phosphatase 5: implications for TPR-mediated protein-protein interactions. EMBO J. 17, 1192–1199 (1998).
Guergnon, J., Derewenda, U., Edelson, J. R. & Brautigan, D. L. Mapping of protein phosphatase-6 association with its SAPS domain regulatory subunit using a model of helical repeats. BMC Biochem. 10, 24 (2009).
Stefansson, B., Ohama, T., Daugherty, A. E. & Brautigan, D. L. Protein phosphatase 6 regulatory subunits composed of ankyrin repeat domains. Biochemistry 47, 1442–1451 (2008).
Huang, X. & Honkanen, R. E. Molecular cloning, expression, and characterization of a novel human serine/threonine protein phosphatase, PP7, that is homologous to Drosophila retinal degeneration C gene product (rdgC). J. Biol. Chem. 273, 1462–1468 (1998).
Verbinnen, I., Ferreira, M. & Bollen, M. Biogenesis and activity regulation of protein phosphatase 1. Biochem. Soc. Trans. 45, 89–99 (2017).
Hertz, E. P. T. et al. A conserved motif provides binding specificity to the PP2A-B56 phosphatase. Mol. Cell 63, 686–695 (2016).
Wang, X., Bajaj, R., Bollen, M., Peti, W. & Page, R. Expanding the PP2A interactome by defining a B56-specific SLiM. Structure 24, 2174–2181 (2016).
Roy, J. & Cyert, M. S. Cracking the phosphatase code: docking interactions determine substrate specificity. Sci. Signal. 2, re9 (2009).
Roy, J., Li, H., Hogan, P. G. & Cyert, M. S. A conserved docking site modulates substrate affinity for calcineurin, signaling output, and in vivo function. Mol. Cell 25, 889–901 (2007).
Ueki, Y. et al. A consensus binding motif for the PP4 protein phosphatase. Mol. Cell 76, 953–964 (2019).
Acknowledgements
A.N.K. and S.A.G. acknowledge support from NCI R33 CA225458. S.A.G. received support from NCI P30 CA0231008. K.W. was supported by a Burroughs-Wellcome Big Data in the Life Sciences Fellowship. We thank K. Smolen for assistance with video editing.
Author information
Authors and Affiliations
Contributions
B.L.B., S.M., G.B.M. and A.N.K. developed and wrote the protocol. B.L.B. drew the illustrations. K.W. and S.A.G. contributed to the design of the data analysis workflow. S.A.G. contributed to the editing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Albert J. R. Heck, Martin R. Larsen and the other, anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Lyons, S. P. et al. Mol. Cell Proteomics 17, 2448–2461 (2018): https://doi.org/10.1074/mcp.RA118.000822
Nasa, I. et al. Sci. Signal. 13, eaba7823 (2020): https://doi.org/10.1126/scisignal.aba7823
Supplementary information
Supplementary Table 1
Annotated PPP subunits and interactors. Uniprot ID and entry name are given for each protein, alongside its gene name and description. PPP subunit indicates the PPP the proteins belong to (PP1-7), whether it is a catalytic subunit (C), a regulatory subunit (R), an inhibitory protein (i), or an activating protein (a).
Supplementary Video 1
A step-by-step demonstration of making a StageTip and using it to desalt a TMT label check sample.
Rights and permissions
About this article
Cite this article
Brauer, B.L., Wiredu, K., Mitchell, S. et al. Affinity-based profiling of endogenous phosphoprotein phosphatases by mass spectrometry. Nat Protoc 16, 4919–4943 (2021). https://doi.org/10.1038/s41596-021-00604-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-021-00604-3
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.