Abstract
RNA fluorescence in situ hybridization (FISH) and antibody staining/immunofluorescence (IF) are widely used to detect distributions of mRNAs and proteins. Here we describe a combined FISH and IF protocol to simultaneously detect multiple mRNAs and proteins in whole-mount zebrafish embryos and larvae. In our approach, FISH is performed before IF to prevent mRNA degradation during the IF procedure. Instead of proteinase K digestion, Triton X-100 treatment and skin removal are used to permeate tissues and preserve antigen epitopes, making this protocol applicable to both whole-mount embryos and larvae. Off-target hybridization and FISH background are reduced by using PCR-amplified DNA templates and stringent buffers. This protocol simultaneously detects multiple mRNAs and proteins with high sensitivity, and enables detection at single-cell resolution. The protocol can be completed within 6 days, overcoming the shortage of reliable antibodies available for zebrafish and exploiting the advantages of zebrafish for studying organ development and regeneration.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The authors declare that the data supporting the findings of this study are available within the paper and its Supplementary Information files. Any additional data, if needed, will be provided upon request.
References
Harding, K., Wedeen, C., McGinnis, W. & Levine, M. Spatially regulated expression of homeotic genes in Drosophila. Science 229, 1236–1242 (1985).
Vize, P. D., McCoy, K. E. & Zhou, X. Multichannel wholemount fluorescent and fluorescent/chromogenic in situ hybridization in Xenopus embryos. Nat. Protoc. 4, 975–983 (2009).
Broadbent, J. & Read, E. M. Wholemount in situ hybridization of Xenopus and zebrafish embryos. Methods Mol. Biol. 127, 57–67 (1999).
Fienberg, A. A. et al. Homeo box genes in murine development. Curr. Top. Dev. Biol. 23, 233–256 (1987).
Speel, E. J., Hopman, A. H. & Komminoth, P. Amplification methods to increase the sensitivity of in situ hybridization: play card(s). J. Histochem. Cytochem. 47, 281–288 (1999).
Speel, E. J., Saremaslani, P., Roth, J., Hopman, A. H. & Komminoth, P. Improved mRNA in situ hybridization on formaldehyde-fixed and paraffin-embedded tissue using signal amplification with different haptenized tyramides. Histochem. Cell Biol. 110, 571–577 (1998).
Baldino, F. Jr., Chesselet, M. F. & Lewis, M. E. High-resolution in situ hybridization histochemistry. Methods Enzymol. 168, 761–777 (1989).
McDougall, J. K., Dunn, A. R. & Jones, K. W. In situ hybridization of adenovirus RNA and DNA. Nature 236, 346–348 (1972).
Wilkinson, D. G., Bhatt, S., Chavrier, P., Bravo, R. & Charnay, P. Segment-specific expression of a zinc-finger gene in the developing nervous system of the mouse. Nature 337, 461–464 (1989).
Springer, J. E., Robbins, E., Gwag, B. J., Lewis, M. E. & Baldino, F. Jr. Non-radioactive detection of nerve growth factor receptor (NGFR) mRNA in rat brain using in situ hybridization histochemistry. J. Histochem. Cytochem. 39, 231–234 (1991).
Lauter, G., Soll, I. & Hauptmann, G. Sensitive whole-mount fluorescent in situ hybridization in zebrafish using enhanced tyramide signal amplification. Methods Mol. Biol. 1082, 175–185 (2014).
Bobrow, M. N., Litt, G. J., Shaughnessy, K. J., Mayer, P. C. & Conlon, J. The use of catalyzed reporter deposition as a means of signal amplification in a variety of formats. J. Immunol. Methods 150, 145–149 (1992).
Zaidi, A. U., Enomoto, H., Milbrandt, J. & Roth, K. A. Dual fluorescent in situ hybridization and immunohistochemical detection with tyramide signal amplification. J. Histochem. Cytochem. 48, 1369–1375 (2000).
Pepe, F. A. Analysis of antibody staining patterns obtained with striated myofibrils in fluorescence microscopy and electron microscopy. Int. Rev. Cytol. 24, 193–231 (1968).
Haffter, P. et al. The identification of genes with unique and essential functions in the development of the zebrafish, Danio rerio. Development 123, 1–36 (1996).
Howe, K. et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature 496, 498–503 (2013).
Thisse, B. & Thisse, C. In situ hybridization on whole-mount zebrafish embryos and young larvae. Methods Mol. Biol. 1211, 53–67 (2014).
Toledano, H., D’Alterio, C., Loza-Coll, M. & Jones, D. L. Dual fluorescence detection of protein and RNA in Drosophila tissues. Nat. Protoc. 7, 1808–1817 (2012).
Wang, F. et al. RNAscope: a novel in situ RNA analysis platform for formalin-fixed, paraffin-embedded tissues. J. Mol. Diagn. 14, 22–29 (2012).
Lauter, G., Soll, I. & Hauptmann, G. Sensitive multiplexed fluorescent in situ hybridization using enhanced tyramide signal amplification and its combination with immunofluorescent protein visualization in zebrafish. Methods Mol. Biol. 2047, 397–409 (2020).
Gross-Thebing, T., Paksa, A. & Raz, E. Simultaneous high-resolution detection of multiple transcripts combined with localization of proteins in whole-mount embryos. BMC Biol. 12, 55 (2014).
He, J. et al. Mammalian target of rapamycin complex 1 signaling is required for the dedifferentiation from biliary cell to bipotential progenitor cell in zebrafish liver regeneration. Hepatology 70, 2092–2106 (2019).
He, J., Lu, H., Zou, Q. & Luo, L. Regeneration of liver after extreme hepatocyte loss occurs mainly via biliary transdifferentiation in zebrafish. Gastroenterology 146, 789–800 e788 (2014).
Chen, J. et al. Cerebrovascular injuries induce lymphatic invasion into brain parenchyma to guide vascular regeneration in zebrafish. Dev. Cell 49, 697–710 (2019).
Julich, D. et al. beamter/deltaC and the role of Notch ligands in the zebrafish somite segmentation, hindbrain neurogenesis and hypochord differentiation. Dev. Biol. 286, 391–404 (2005).
Brend, T. & Holley, S.A. Zebrafish whole mount high-resolution double fluorescent in situ hybridization. J. Vis. Exp. 25, e1229 (2009).
Thisse, C. & Thisse, B. High-resolution in situ hybridization to whole-mount zebrafish embryos. Nat. Protoc. 3, 59–69 (2008).
Furthauer, M., Reifers, F., Brand, M., Thisse, B. & Thisse, C. sprouty4 acts in vivo as a feedback-induced antagonist of FGF signaling in zebrafish. Development 128, 2175–2186 (2001).
Furthauer, M., Lin, W., Ang, S. L., Thisse, B. & Thisse, C. Sef is a feedback-induced antagonist of Ras/MAPK-mediated FGF signalling. Nat. Cell Biol. 4, 170–174 (2002).
Tsang, M., Friesel, R., Kudoh, T. & Dawid, I. B. Identification of Sef, a novel modulator of FGF signalling. Nat. Cell Biol. 4, 165–169 (2002).
Kikuchi, Y. et al. casanova encodes a novel Sox-related protein necessary and sufficient for early endoderm formation in zebrafish. Genes Dev. 15, 1493–1505 (2001).
Donovan, A. et al. The zebrafish mutant gene chardonnay (cdy) encodes divalent metal transporter 1 (DMT1). Blood 100, 4655–4659 (2002).
Lauter, G., Soll, I. & Hauptmann, G. Two-color fluorescent in situ hybridization in the embryonic zebrafish brain using differential detection systems. BMC Dev. Biol. 11, 43 (2011).
Cao, Z., Mao, X. & Luo, L. Germline stem cells drive ovary regeneration in zebrafish. Cell Rep. 26, 1709–1717 e1703 (2019).
Cha, Y. R. & Weinstein, B. M. Use of PCR template-derived probes prevents off-target whole mount in situ hybridization in transgenic zebrafish. Zebrafish 9, 85–89 (2012).
Field, H. A., Ober, E. A., Roeser, T. & Stainier, D. Y. Formation of the digestive system in zebrafish. I. Liver morphogenesis. Dev. Biol. 253, 279–290 (2003).
Cooper, M. S. et al. Visualizing morphogenesis in transgenic zebrafish embryos using BODIPY TR methyl ester dye as a vital counterstain for GFP. Dev. Dyn. 232, 359–368 (2005).
Godinho, L. et al. Targeting of amacrine cell neurites to appropriate synaptic laminae in the developing zebrafish retina. Development 132, 5069–5079 (2005).
Her, G. M., Chiang, C. C., Chen, W. Y. & Wu, J. L. In vivo studies of liver-type fatty acid binding protein (L-FABP) gene expression in liver of transgenic zebrafish (Danio rerio). FEBS Lett. 538, 125–133 (2003).
Acknowledgements
We thank K. D. Poss for providing pCS2-ctgfa:GFP plasmid. This work was supported by the National Key Research and Development Program of China (2017YFA0106600), the National Natural Science Foundation of China (31730060, 31970784, 31801214, 91739304) and the 111 Program (B14037).
Author information
Authors and Affiliations
Contributions
L.L., J.H. and D.M. designed the experimental strategy, analysed data and wrote the manuscript. J.C. performed the experiments shown in Fig. 6 and Extended Data Fig. 6. D.M. and J.H. performed all the other experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Uwe Strähle and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
He, J. et al. Hepatology 70, 2092−2106 (2019): https://doi.org/10.1002/hep.30790
Chen, J. et al. Dev. Cell 49, 697−710 (2019): https://doi.org/10.1016/j.devcel.2019.03.022
He, J., Lu, H., Zou, Q. & Luo, L. Gastroenterology 146, 789−800 (2014): https://doi.org/10.1053/j.gastro.2013.11.045
Extended data
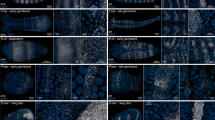
Extended Data Fig. 1 FISH in combination with antibody staining of brain neuron and spinal cord at late embryonic and larval stage.
a, Labelling of FISH-huc (a gene specific to brain neurons and spinal cord), anti-GFP and DAPI in the whole body in the Tg(RasGFP) transgenic line at 4 dpf (3D stack of images; 448 µm, 32 slices, 14 µm each slice). b, Labelling of FISH-huc, anti-GFP and DAPI in the whole body in the Tg(RasGFP) transgenic line at 10 dpf (3D stack of images; 448 µm, 32 slices, 14 µm each slice). Note that huc mRNA is expressed in the brain neuron, spinal cord and intestine; the GFP and DAPI stained the whole body. Scale bar, 100 µm.
Extended Data Fig. 2 FISH in combination with antibody staining of notochord in larvae.
a, The Kaede expresses in the notochord at 5 and 10 dpf in the transgenic line Tg(ctgfa:Kaede). b, Labelling of FISH-ctgfa, anti-Kaede and DAPI in the notochord (dashed line) in Tg(ctgfa:Kaede) at 5 dpf (3D stack of images; 68 µm, 17 slices, 4 µm each slice). c, Labelling of FISH-ctgfa, anti-Kaede and DAPI in the notochord (dashed line) in Tg(ctgfa:Kaede) at 10 dpf (3D stack of images; 68 µm, 17 slices, 4 µm each slice). Scale bar, 100 µm.
Extended Data Fig. 3 FISH in combination with antibody staining of pectoral fin, skin and hair cells at 10 dpf.
a, The Tomato expresses in skin and hair cells at 10 dpf in the transgenic line Tg(krt18:Tomato). b, Labelling of FISH-ctgfa (a pectoral fin marker), anti-Tomato and DAPI in the pectoral fin (dashed line, top view) in Tg(krt18:Tomato) at 10 dpf (3D stack of images; 152 µm, 38 slices, 4 µm each slice; dorsal view). c, Labelling of FISH-ctgfa (a pectoral fin marker), anti-Tomato and DAPI in the pectoral fin (dashed line, front view) in Tg(krt18:Tomato) at 10 dpf (3D stack of images; 152 µm, 38 slices, 4 µm each slice; ventral view). Note that the Tomato expresses in the skin and FISH-ctgfa expresses in the pectoral fin under the skin. d, Labelling of FISH-krt18 (the skin and hair cells specific-gene), anti-Tomato and DAPI in skin and hair cells in Tg(krt18:Tomato) at 10 dpf (3D stack of images; 232 µm, 58 slices, 4 µm each slice). Note that the FISH-krt18 merges with Tomato and expresses strongly in hair cells (arrowheads) and weekly in the skin. BF, bright field. Scale bar, 100 µm.
Extended Data Fig. 4 FISH in combination with antibody staining of hepatic duct cells in the liver and secretory cells in the intestine at 20 dpf.
a–d, Labelling of FISH-anxa4 (the antigen of 2F11), 2F11 (the marker of hepatic duct cells and secretory cells in the intestine) and DAPI in hepatic duct cells in the liver and secretory cells in the intestine in wild type at 20 dpf (3D stack of images; 337.33 µm, 38 slices, 8.877 µm each slice; left view). BF, bright field. Scale bar, 100 µm.
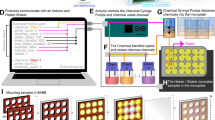
Extended Data Fig. 5 Mounting the embryos or larvae.
a, A glass-bottom microwell dish (30 mm) was used for mounting the embryos and larvae. The samples were mounted in the middle of the dish with 100–120 µl 1.2% LMP agarose.
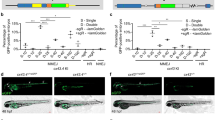
Extended Data Fig. 6 Comparison of the current protocol with the traditional FISH protocol combined with antibody staining in the liver and meningeal lymphatic and vascular system at the larval stage.
a, The liver after triple labelling of FISH-cp, 2F11, anti-GFP and with DAPI staining under the Tg(RasGFP) transgenic line at 5 dpf (3D stack of images; 104 µm, 26 slices, 4 µm each slice) using this protocol. b, The liver after triple labelling of FISH-cp, 2F11, anti-GFP and with DAPI staining under the Tg(RasGFP) transgenic line at 5 dpf (2D image) using this protocol. c, The liver after triple labelling of FISH-cp, 2F11, anti-GFP and with DAPI staining under the Tg(RasGFP) transgenic line at 5 dpf (3D stack of images; 108 µm, 27 slices, 4 µm each slice) using the traditional FISH protocol combined with antibody staining. d, The liver after triple labelling of FISH-cp, 2F11, anti-GFP and with DAPI staining under the Tg(RasGFP) transgenic line at 5 dpf (2D image) using the traditional FISH protocol combined with antibody staining. Note that the 2F11 and anti-GFP are hardly detectable with the traditional protocol. e, The meningeal lymphatic and vascular cells after triple labelling of the Dig-labelled mrc1a probe, anti-Kaede and anti-CFP in the Tg(lyve1b:Kaede; kdrl:CFP−NTR) transgenic line at 5 dpf using this protocol (3D stack of images; 200 µm, 50 slices, 4 µm each slice). f, The meningeal lymphatic and vascular cells after triple labelling of Dig-labelled mrc1a probe, anti-Kaede and anti-CFP in the Tg(lyve1b:Kaede; kdrl:CFP-NTR) transgenic line at 5 dpf using traditional FISH protocol combined with antibody staining (3D stack of images; 200 µm, 50 slices, 4 µm each slice). Note that the signals of FISH-mrc1a, anti-CFP and anti-Kaede are weak in the traditional protocol compared with this protocol. Scale bar, 100 µm.
Supplementary information
Supplementary Information
Supplementary Methods, Supplementary References and Supplementary Tables 1 and 2.
Supplementary Video 1
Removing the skin of zebrafish embryo/larva. Use the tweezer in the left hand to hold the head of embryo/larva. Then, use the tweezer in the right hand to peel off the skin from the heart area. After removing the skin of the trunk from the heart to the tail fin, use tweezer in the left hand to hold the embryo/larva by the swim bladder. Then, dissect the skin from the hindbrain with the right hand, and remove all the skin from the head.
Rights and permissions
About this article
Cite this article
He, J., Mo, D., Chen, J. et al. Combined whole-mount fluorescence in situ hybridization and antibody staining in zebrafish embryos and larvae. Nat Protoc 15, 3361–3379 (2020). https://doi.org/10.1038/s41596-020-0376-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-020-0376-7
This article is cited by
-
New genes helped acorn barnacles adapt to a sessile lifestyle
Nature Genetics (2024)
-
Visualisation of gene expression within the context of tissues using an X-ray computed tomography-based multimodal approach
Scientific Reports (2024)
-
Zebrafish ELL-associated factors Eaf1/2 modulate erythropoiesis via regulating gata1a expression and WNT signaling to facilitate hypoxia tolerance
Cell Regeneration (2023)
-
hapln1a+ cells guide coronary growth during heart morphogenesis and regeneration
Nature Communications (2023)
-
Visualization of metabolites and microbes at high spatial resolution using MALDI mass spectrometry imaging and in situ fluorescence labeling
Nature Protocols (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.