Abstract
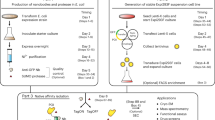
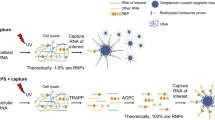
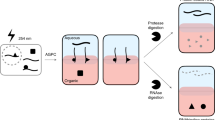
The initial interactions between incoming, pre-replicated virion RNA and host protein factors are important in infection and immunity. Yet currently there are no methods to study these crucial events. We established VIR-CLASP (VIRal Cross-Linking And Solid-phase Purification) to identify the primary viral RNA–host protein interactions. First, host cells are infected with 4-thiouridine (4SU)-labeled RNA viruses and irradiated with 365 nm light to crosslink 4SU-labeled viral genomes and interacting proteins from host or virus. The crosslinked RNA binding proteins (RBPs) are purified by solid-phase reversible immobilization (SPRI) beads with protein-denaturing buffers, and then identified by proteomics. With VIR-CLASP, only the incoming virion RNAs are labeled with 4SU, so crosslinking events specifically occur between proteins and pre-replicated virion RNA. Since solid-phase purification under protein-denaturing conditions, rather than sequence-specific nucleic acid purification, is used to pull-down total RNA and crosslinked RBPs, this method facilitates investigation of potentially all RNA viruses, regardless of RNA sequence. Preparation of 4SU-labeled virus takes ∼7 days and VIR-CLASP takes 1 day.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Code availability
All code used for proteomic analysis and figure generation can be accessed under the GNU General Public License v3.0 at https://github.com/Ascano-Lab.
References
Phillips, S. L., Soderblom, E. J., Bradrick, S. S. & Garcia-Blanco, M. A. Identification of proteins bound to dengue viral RNA in vivo reveals new host proteins important for virus replication. MBio 7, e01865–15 (2016).
Lenarcic, E. M., Landry, D. M., Greco, T. M., Cristea, I. M. & Thompson, S. R. Thiouracil cross-linking mass spectrometry: a cell-based method to identify host factors involved in viral amplification. J. Virol. 87, 8697–8712 (2013).
Garcia-Moreno, M. et al. System-wide profiling of RNA-binding proteins uncovers key regulators of virus infection. Mol. Cell 74, 196–211.e11 (2019).
Zhou, Y. & Routh, A. Mapping RNA-capsid interactions and RNA secondary structure within virus particles using next-generation sequencing. Nucleic Acids Res. 48, e12–e12 (2020).
Kim, B. et al. Discovery of widespread host protein interactions with the pre-replicated genome of CHIKV using VIR-CLASP. Mol. Cell 78, 624–640.e7 (2020).
Barrows, N. J. et al. Dual roles for the ER membrane protein complex in flavivirus infection: viral entry and protein biogenesis. Sci. Rep. 9, 9711 (2019).
Hafner, M. et al. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 141, 129–141 (2010).
Favre, A. et al. 4-Thiouridine photosensitized RNA-protein crosslinking in mammalian cells. Biochem. Biophys. Res. Commun. 141, 847–854 (1986).
DeAngelis, M. M., Wang, D. G. & Hawkins, T. L. Solid-phase reversible immobilization for the isolation of PCR products. Nucleic Acids Res. 23, 4742–4743 (1995).
Licatalosi, D. D. et al. HITS-CLIP yields genome-wide insights into brain alternative RNA processing. Nature 456, 464–469 (2008).
Van Nostrand, E. L. et al. Robust transcriptome-wide discovery of RNA-binding protein binding sites with enhanced CLIP (eCLIP). Nat. Methods 13, 508–514 (2016).
König, J. et al. iCLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Nat. Struct. Mol. Biol. 17, 909–915 (2010).
Ascano, M., Hafner, M., Cekan, P., Gerstberger, S. & Tuschl, T. Identification of RNA-protein interaction networks using PAR-CLIP. Wiley Interdiscip. Rev. RNA 3, 159–177 (2012).
Trendel, J. et al. The human RNA-binding proteome and its dynamics during translational arrest. Cell 176, 391–403.e19 (2019).
Asencio, C., Chatterjee, A. & Hentze, M. W. Silica-based solid-phase extraction of cross-linked nucleic acid-bound proteins. Life Sci. Alliance 1, e201800088 (2018).
Shchepachev, V. et al. Defining the RNA interactome by total RNA-associated protein purification. Mol. Syst. Biol. 15, e8689 (2019).
Berensmeier, S. Magnetic particles for the separation and purification of nucleic acids. Appl. Microbiol. Biotechnol. 73, 495–504 (2006).
Miller, C. et al. Dynamic transcriptome analysis measures rates of mRNA synthesis and decay in yeast. Mol. Syst. Biol. 7, 458 (2011).
Hoffmann, M. et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181, 271–280.e8 (2020).
Jiang, P., Liu, Y., Ma, H.-C., Paul, A. V. & Wimmer, E. Picornavirus morphogenesis. Microbiol. Mol. Biol. Rev. 78, 418–437 (2014).
Quail, M. A., Swerdlow, H. & Turner, D. J. Improved protocols for the illumina genome analyzer sequencing system. Curr. Protoc. Hum. Genet. Chapter 18, Unit 18, 2 (2009).
Telesnitsky, A. & Wolin, S. L. The host RNAs in retroviral particles. Viruses 8, 235 (2016).
Noda, T. et al. Importance of the 1+7 configuration of ribonucleoprotein complexes for influenza A virus genome packaging. Nat. Commun. 9, 54 (2018).
Kaur, P., Lee, R. C. H. & Chu, J. J. H. Infectious viral quantification of chikungunya virus-virus plaque assay. Methods Mol. Biol. 1426, 93–103 (2016).
Pedersen, I. R. Density gradient centrifugation studies on lymphocytic choriomeningitis virus and on viral ribonucleic acid. J. Virol. 6, 414–420 (1970).
Lim, C.-K. Virus isolation and preparation of sucrose-banded chikungunya virus samples for transmission electron microscopy. Methods Mol. Biol. 1426, 153–162 (2016).
Mortz, E., Krogh, T. N., Vorum, H. & Görg, A. Improved silver staining protocols for high sensitivity protein identification using matrix-assisted laser desorption/ionization-time of flight analysis. Proteomics 1, 1359–1363 (2001).
Link, A. J. & Labaer, J. In-gel trypsin digest of gel-fractionated proteins. Cold Spring Harb. Protoc. 2009, pdb.prot5110–pdb.prot5110 (2009).
Smyth, G. K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, Article3 (2004).
Sysoev, V. O. et al. Global changes of the RNA-bound proteome during the maternal-to-zygotic transition in Drosophila. Nat. Commun. 7, 12128–11 (2016).
Conway, J. R., Lex, A. & Gehlenborg, N. UpSetR: an R package for the visualization of intersecting sets and their properties. Bioinformatics 33, 2938–2940 (2017).
Dickson, A. M. et al. Dephosphorylation of HuR protein during alphavirus infection is associated with HuR relocalization to the cytoplasm. J. Biol. Chem. 287, 36229–36238 (2012).
Daffis, S. et al. 2′-O methylation of the viral mRNA cap evades host restriction by IFIT family members. Nature 468, 452–456 (2010).
Lex, A., Gehlenborg, N., Strobelt, H., Vuillemot, R. & Pfister, H. UpSet: visualization of intersecting sets. IEEE Trans. Vis. Comput. Graph 20, 1983–1992 (2014).
Acknowledgements
We thank K. L. Rose and W. H. McDonald at the Vanderbilt Mass Spectrometry Research Center for processing of the mass spectrometry samples; M. Albertolle for technical help in the mass spectrometry analysis; T. Voss and J. E. Crowe Jr. (Vanderbilt University Medical Center) for IAV; and T. S. Dermody (University of Pittsburgh School of Medicine) for CHIKV. Finally, we thank members of the Ascano laboratory for their support, collegiality, and critical review of the manuscript. This work was supported by the National Institutes of Health 1R35GM119569-01 (M.A.), CTSA award No.UL1TR000445 from the National Center for Advancing Translational Sciences (B.K. and M.A.), Vanderbilt University Dept. of Biochemistry start-up funds (M.A.), the Chemical Biology of Infectious Disease training grant 5T32AI11254-02 (S.A.), and the Chemistry-Biology Interface training grant 5T32GM065086-14.
Author information
Authors and Affiliations
Contributions
B.K. and M.A. designed, and B.K. optimized the VIR-CLASP technique. S.A. performed the proteomic analysis and data visualization. S.A., B.K., and M.A. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Encarnacion Martinez-Salas, Ding Shou-wei and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Kim, B. et al. Mol. Cell 78, 624–640.e7 (2020): https://doi.org/10.1016/j.molcel.2020.04.013
Barrows, N. et al. Sci. Rep. 9, 9711 (2019): https://doi.org/10.1038/s41598-019-45910-9
Supplementary information
Supplementary Information
Supplementary Fig. 1.
Rights and permissions
About this article
Cite this article
Kim, B., Arcos, S., Rothamel, K. et al. Viral crosslinking and solid-phase purification enables discovery of ribonucleoprotein complexes on incoming RNA virus genomes. Nat Protoc 16, 516–531 (2021). https://doi.org/10.1038/s41596-020-00429-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-020-00429-6
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.