Abstract
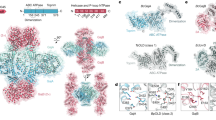
As one of the most prevalent anti-phage defense systems in prokaryotes, Gabija consists of a Gabija protein A (GajA) and a Gabija protein B (GajB). The assembly and function of the Gabija system remain unclear. Here we present cryo-EM structures of Bacillus cereus GajA and GajAB complex, revealing tetrameric and octameric assemblies, respectively. In the center of the complex, GajA assembles into a tetramer, which recruits two sets of GajB dimer at opposite sides of the complex, resulting in a 4:4 GajAB supramolecular complex for anti-phage defense. Further biochemical analysis showed that GajA alone is sufficient to cut double-stranded DNA and plasmid DNA, which can be inhibited by ATP. Unexpectedly, the GajAB displays enhanced activity for plasmid DNA, suggesting a role of substrate selection by GajB. Together, our study defines a framework for understanding anti-phage immune defense by the GajAB complex.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Accession numbers for Gabija A tetramer (4A), Gabija AB complex 1 (4A:4B, C1 symmetry) and Gabija AB complex 2 (4A:4B, D2 symmetry) are as follows: coordinates of atomic models, 8TK0, 8TK1 and 8TJY, deposited to Protein Data Bank, and density map. EMD-41319, EMD-41321 and EMD-41314, deposited to Electron Microscopy Data Bank. All data needed to evaluate the conclusions are present in the paper. Source data are provided with this paper.
References
Tal, N. & Sorek, R. SnapShot: bacterial immunity. Cell 185, 578–578 e571 (2022).
Koonin, E. V., Makarova, K. S. & Zhang, F. Diversity, classification and evolution of CRISPR–Cas systems. Curr. Opin. Microbiol. 37, 67–78 (2017).
Duncan-Lowey, B. & Kranzusch, P. J. CBASS phage defense and evolution of antiviral nucleotide signaling. Curr. Opin. Immunol. 74, 156–163 (2022).
Doron, S. et al. Systematic discovery of antiphage defense systems in the microbial pangenome. Science https://doi.org/10.1126/science.aar4120 (2018).
Gao, L. et al. Diverse enzymatic activities mediate antiviral immunity in prokaryotes. Science 369, 1077–1084 (2020).
Millman, A. et al. An expanded arsenal of immune systems that protect bacteria from phages. Cell Host Microbe 30, 1556–1569 e1555 (2022).
Tesson, F. et al. Systematic and quantitative view of the antiviral arsenal of prokaryotes. Nat. Commun. 13, 2561 (2022).
Cheng, R. et al. A nucleotide-sensing endonuclease from the Gabija bacterial defense system. Nucleic Acids Res. 49, 5216–5229 (2021).
Cheng, R. et al. Prokaryotic Gabija complex senses and executes nucleotide depletion and DNA cleavage for antiviral defense. Cell Host Microbe https://doi.org/10.1016/j.chom.2023.06.014 (2023).
Liu, Y. et al. ATP-dependent DNA binding, unwinding, and resection by the Mre11/Rad50 complex. EMBO J. 35, 743–758 (2016).
Hopfner, K. P. et al. Structural biology of Rad50 ATPase: ATP-driven conformational control in DNA double-strand break repair and the ABC-ATPase superfamily. Cell 101, 789–800 (2000).
Aravind, L., Leipe, D. D. & Koonin, E. V. Toprim—a conserved catalytic domain in type IA and II topoisomerases, DnaG-type primases, OLD family nucleases and RecR proteins. Nucleic Acids Res. 26, 4205–4213 (1998).
Schiltz, C. J., Lee, A., Partlow, E. A., Hosford, C. J. & Chappie, J. S. Structural characterization of Class 2 OLD family nucleases supports a two-metal catalysis mechanism for cleavage. Nucleic Acids Res. 47, 9448–9463 (2019).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Antine, S. P. et al. Structural basis of Gabija anti-phage defence and viral immune evasion. Nature https://doi.org/10.1038/s41586-023-06855-2 (2023).
Schiltz, C. J., Adams, M. C. & Chappie, J. S. The full-length structure of Thermus scotoductus OLD defines the ATP hydrolysis properties and catalytic mechanism of Class 1 OLD family nucleases. Nucleic Acids Res. 48, 2762–2776 (2020).
Baek, M. et al. Accurate prediction of protein–nucleic acid complexes using RoseTTAFoldNA. Nat. Methods https://doi.org/10.1038/s41592-023-02086-5 (2023).
Korolev, S., Hsieh, J., Gauss, G. H., Lohman, T. M. & Waksman, G. Major domain swiveling revealed by the crystal structures of complexes of E. coli Rep helicase bound to single-stranded DNA and ADP. Cell 90, 635–647 (1997).
Velankar, S. S., Soultanas, P., Dillingham, M. S., Subramanya, H. S. & Wigley, D. B. Crystal structures of complexes of PcrA DNA helicase with a DNA substrate indicate an inchworm mechanism. Cell 97, 75–84 (1999).
Lee, J. Y. & Yang, W. UvrD helicase unwinds DNA one base pair at a time by a two-part power stroke. Cell 127, 1349–1360 (2006).
Duncan-Lowey, B. et al. Cryo-EM structure of the RADAR supramolecular anti-phage defense complex. Cell 186, 987–998 e915 (2023).
Gao, Y. et al. Molecular basis of RADAR anti-phage supramolecular assemblies. Cell 186, 999–1012 e1020 (2023).
Gao, L. A. et al. Prokaryotic innate immunity through pattern recognition of conserved viral proteins. Science 377, eabm4096 (2022).
Shen, Z., Lin, Q., Yang, X. Y., Fosuah, E. & Fu, T. M. Assembly-mediated activation of the SIR2-HerA supramolecular complex for anti-phage defense. Mol. Cell https://doi.org/10.1016/j.molcel.2023.11.007 (2023).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Williams, C. J. et al. MolProbity: more and better reference data for improved all-atom structure validation. Protein Sci. 27, 293–315 (2018).
The PyMOL Molecular Graphics System, Version 3.0 (Schrödinger, 2022)
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Mazzocco, A., Waddell, T. E., Lingohr, E. & Johnson, R. P. Enumeration of bacteriophages using the small drop plaque assay system. Methods Mol. Biol. 501, 81–85 (2009).
Kropinski, A. M., Mazzocco, A., Waddell, T. E., Lingohr, E. & Johnson, R. P. Enumeration of bacteriophages by double agar overlay plaque assay. Methods Mol. Biol. 501, 69–76 (2009).
Marty, M. T. et al. Bayesian deconvolution of mass and ion mobility spectra: from binary interactions to polydisperse ensembles. Anal. Chem. 87, 4370–4376 (2015).
Acknowledgements
Cryo-EM data of GajA were collected at OSU CEMAS with the assistance of G. Grandinetti and Y. Narui. Cryo-EM data of GajAB were collected with the assistance of A. D. Wier, T. J. Edwards, T. Fox and J. Wang at the National Cancer Institute Cryo-Electron Microscopy Center supported by grants from the NIH National Institute of General Medical Sciences (GM103310). T.-M.F. is supported by an R35 grant from National Institute of General Medical Sciences (1R35GM147465-01).
Author information
Authors and Affiliations
Contributions
T.-M.F. conceived the project. X.-Y.Y., Z.S. and J.X. performed molecular cloning, biochemical purification, ATPase assay and plaque assays and determined nuclease activity. Z.S. prepared EM grids and determined the cryo-EM structures. Q.L. helped on structural reconstruction. Z.S. and X.-Y.Y. built the models. W.X. predicted the structure of the GajA–dsDNA complex. J.G. and I.M. performed native mass spectrometry analysis under the supervision of V.H.W. Z.S., X.-Y.Y. and T.-M.F. analyzed all the data together. T.-M.F. wrote the manuscript with inputs from all the authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Qian Yin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Sara Osman and Dimitris Typas were the primary editors on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cryo-EM reconstruction of GajA in thin ice.
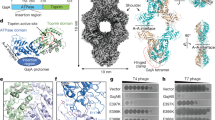
a, b, Gel filtration profile (a) and SDS-PAGE gel (b) of GajA purification. The experiment was replicated at least three times. c, A representative cryo-EM image of GajA in thin ice. Thousands of images were collected. d, Representative 2D class averages of GajA calculated from thin-ice cryo-EM images. e, Data processing workflow for 3D reconstruction of GajA tetramer from thin-ice cryo-EM images. f, The FSC curve of reconstructed GajA tetramer from thin-ice cryo-EM images. g, Representative cryo-EM density of GajA tetramer fitted with α-helices and β-strands. The density map was shown at a contour level of 0.03.
Extended Data Fig. 2 Cryo-EM reconstruction of GajA in thicker ice.
a, A representative cryo-EM image of GajA in thicker ice. Thousands of images were collected. b, 2D class averages of GajA calculated from thick-ice cryo-EM images. c, Data processing workflow for 3D reconstruction of GajA tetramer from thick-ice cryo-EM images. d, The FSC curve of reconstructed GajA tetramer from thick-ice cryo-images. e, Local resolution of reconstructed GajA tetramer from thick-ice cryo-images. f, Cryo-EM density of GajA tetramer fit with α-helices and β-strands. The density map is shown at contour levels of 0.03.
Extended Data Fig. 3 Architecture of GajA.
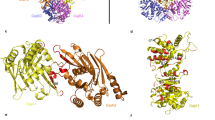
a, Ribbon diagram of GajA N-terminal ATPase domain with secondary structures indicated. b, Overlaid structures of GajA N-terminal ATPase domain (green) and Rad50 ATPase domain (PDB ID 5DNY, magenta). c, Ribbon diagram of GajA C-terminal Toprim domain with secondary structures indicated. d, The dimerization domain of GajA in complex with a phage protein Gad1 (magenata).
Extended Data Fig. 4 Interfaces in GajA tetramer.
a–c, Enlarged views of interface I (a), interface II (b), and interface III (c) in GajA tetramer. Key residues on the interfaces were highlighted in sticks. d, Superimposed structures of the active sites from GajA (green) and BpOLD (PDB ID 6NK8, gray). e, Structure of GajA in complex with dsDNA (Yellow) that was predicted by RoseTTAFoldNA. f, Electrostatic surface representation of GajA with dsDNA. The catalytic center of GajA is highlighted by a red circle. Negatively charged residues surrounding the catalytic center of GajA coordinate dsDNA. g, Key residues involved in coordinating dsDNA are highlighted in sticks.
Extended Data Fig. 5 Oligomerization state of GajB and GajAB.
a, Gel filtration profile of GajB indicates that GajB alone assembles as a monomer. b, Gel filtration profile of GajAB indicates that GajAB assembles as a tetramer of heterodimer. c, Native mass spectrometry analysis revealed that there are four copies of GajA and four copies of GajB in the GajAB complex.
Extended Data Fig. 6 Cryo-EM reconstruction of GajAB.
a, A representative cryo-EM image of GajAB complex. Thousands of images were collected. b, 2D class averages of GajAB complex. c, Data processing workflow for 3D reconstruction of GajAB complex. d, e, Local resolution (d) and FSC curve (e) of reconstructed GajAB complex without symmetry setting. f, g, Local resolution (f) and FSC curve (g) of reconstructed GajAB complex with D2 symmetry setting.
Extended Data Fig. 7 Structural comparison of GajB and UvrD.
a, Overlaid structures of GajB (magenta, pink, yellow, and orange, AlphaFold predicted structure) and UvrD (PDB ID 2IS2, cyan). b, Sequence alignment of ATP binding motifs between GajB and UvrD. c, Overlaid structures of GajB (magenta) and UvrD (cyan) showed that domain 2A of GajB is not well positioned to coordinate ATP. d, Expanded view of key residues involved in coordinating ssDNA from GajB (magenta, AlphaFold predicted structure) and UvrD (cyan). e, Overlaid structures of GajB (magenta, AlphaFold predicted structure) and UvrD (cyan) revealed that domain 2B in GajB lacks key motifs for coordinating dsDNA.
Extended Data Fig. 8 Interfaces in GajAB.
a, Key residues mediating interactions between GajB domain 1B (magenta) and GajA ATPase domain (green). b, Key residues mediating cis-interactions between GajB domain 1A (pink) and GajA ATPase domain (green). c, Key residues mediating trans-interactions between GajB 1A (pink) and GajA ATPase (blue). d, Key residues mediating interactions of two neighboring GajB protomers.
Supplementary information
Source data
Source Data Fig. 5
Unprocessed gels/unprocessed anti-phage plaque results.
Source Data Fig. 5f
Unprocessed ATPase assays.
Source Data Extended Data Fig. 1
Unprocessed sodium dodecyl sulfate polyacrylamide gel electrophoresis gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yang, XY., Shen, Z., Xie, J. et al. Molecular basis of Gabija anti-phage supramolecular assemblies. Nat Struct Mol Biol (2024). https://doi.org/10.1038/s41594-024-01283-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41594-024-01283-w