Abstract
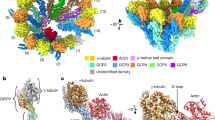
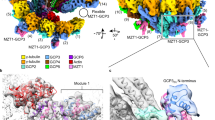
Microtubules are composed of α-tubulin and β-tubulin dimers positioned head-to-tail to form protofilaments that associate laterally in varying numbers. It is not known how cellular microtubules assemble with the canonical 13-protofilament architecture, resulting in micrometer-scale α/β-tubulin tracks for intracellular transport that align with, rather than spiral along, the long axis of the filament. We report that the human ~2.3 MDa γ-tubulin ring complex (γ-TuRC), an essential regulator of microtubule formation that contains 14 γ-tubulins, selectively nucleates 13-protofilament microtubules. Cryogenic electron microscopy reconstructions of γ-TuRC-capped microtubule minus ends reveal the extensive intra-domain and inter-domain motions of γ-TuRC subunits that accommodate luminal bridge components and establish lateral and longitudinal interactions between γ-tubulins and α-tubulins. Our structures suggest that γ-TuRC, an inefficient nucleation template owing to its splayed conformation, can transform into a compacted cap at the microtubule minus end and set the lattice architecture of cellular microtubules.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The following cryo-EM maps have been deposited at the Electron Microscopy Data Bank: free recombinant γ-TuRC (EMD-43481), overall γ-TuRC-capped microtubule (EMD-43482), refined γ-TuRC-capped microtubule (EMD-43483), symmetry expanded map of γ-tubulins and α-tubulins (EMD-43085) and γ-TuRC nucleated microtubule protofilament (EMD-43519). The rigid-body fit models for free recombinant γ-TuRC, overall γ-TuRC-capped microtubule and the refined γ-TuRC-capped microtubule have been deposited in the Protein Data Bank (PDB 8VRD, PDB 8VRJ and PDB 8VRK, respectively). The refined models for the symmetry expanded map of γ-tubulins and α-tubulins and the γ-TuRC nucleated microtubule protofilament have been deposited in PDB (PDB 8VA2 and PDB 8VT7, respectively). Source data are provided with this paper.
References
Kirschner, M. & Mitchison, T. Beyond self-assembly: from microtubules to morphogenesis. Cell 45, 329–342 (1986).
Brouhard, G. J. & Rice, L. M. Microtubule dynamics: an interplay of biochemistry and mechanics. Nat. Rev. Mol. Cell Biol. 19, 451–463 (2018).
Akhmanova, A. & Steinmetz, M. O. Control of microtubule organization and dynamics: two ends in the limelight. Nat. Rev. Mol. Cell Biol. 16, 711–726 (2015).
Tilney, L. G. et al. Microtubules: evidence for 13 protofilaments. J. Cell Biol. 59, 267–275 (1973).
Böhm, K. J., Vater, W., Fenske, H. & Unger, E. Effect of microtubule-associated proteins on the protofilament number of microtubules assembled in vitro. Biochim. Biophys. Acta 800, 119–126 (1984).
Evans, L., Mitchison, T. & Kirschner, M. Influence of the centrosome on the structure of nucleated microtubules. J. Cell Biol. 100, 1185–1191 (1985).
Chrétien, D., Metoz, F., Verde, F., Karsenti, E. & Wade, R. H. Lattice defects in microtubules: protofilament numbers vary within individual microtubules. J. Cell Biol. 117, 1031–1040 (1992).
Wieczorek, M. et al. Asymmetric molecular architecture of the human γ-tubulin ring complex. Cell 180, 165–175 (2020).
Liu, P. et al. Insights into the assembly and activation of the microtubule nucleator γ-TuRC. Nature 578, 467–471 (2020).
Consolati, T. et al. Microtubule nucleation properties of single human γTuRCs explained by their cryo-EM structure. Dev. Cell 53, 603–617 (2020).
Zimmermann, F. et al. Assembly of the asymmetric human γ-tubulin ring complex by RUVBL1–RUVBL2 AAA ATPase. Sci. Adv. 6, eabe0894 (2020).
Wieczorek, M., Huang, T.-L., Urnavicius, L., Hsia, K.-C. & Kapoor, T. M. MZT proteins form multi-faceted structural modules in the γ-tubulin ring complex. Cell Rep. 31, 107791 (2020).
Zheng, Y., Wong, M. L., Alberts, B. & Mitchison, T. Nucleation of microtubule assembly by a γ-tubulin-containing ring complex. Nature 378, 578–583 (1995).
Moritz, M., Braunfeld, M. B., Guénebaut, V., Heuser, J. & Agard, D. A. Structure of the γ-tubulin ring complex: a template for microtubule nucleation. Nat. Cell Biol. 2, 365–370 (2000).
Keating, T. J. & Borisy, G. G. Immunostructural evidence for the template mechanism of microtubule nucleation. Nat. Cell Biol. 2, 352–357 (2000).
Wiese, C. & Zheng, Y. A new function for the γ-tubulin ring complex as a microtubule minus-end cap. Nat. Cell Biol. 2, 358–364 (2000).
Kollman, J. M. et al. Ring closure activates yeast γTuRC for species-specific microtubule nucleation. Nat. Struct. Mol. Biol. 22, 132–137 (2015).
Kollman, J. M., Merdes, A., Mourey, L. & Agard, D. A. Microtubule nucleation by γ-tubulin complexes. Nat. Rev. Mol. Cell Biol. 12, 709–721 (2011).
Oakley, B. R., Paolillo, V. & Zheng, Y. γ-Tubulin complexes in microtubule nucleation and beyond. Mol. Biol. Cell 26, 2957–2962 (2015).
Moritz, M., Zheng, Y., Alberts, B. M. & Oegema, K. Recruitment of the γ-tubulin ring complex to Drosophila salt-stripped centrosome scaffolds. J. Cell Biol. 142, 775–786 (1998).
Oegema, K. et al. Characterization of two related Drosophila γ-tubulin complexes that differ in their ability to nucleate microtubules. J. Cell Biol. 144, 721–733 (1999).
Kollman, J. M. et al. The structure of the γ-tubulin small complex: implications of its architecture and flexibility for microtubule nucleation. Mol. Biol. Cell 19, 207–215 (2008).
Kollman, J. M., Polka, J. K., Zelter, A., Davis, T. N. & Agard, D. A. Microtubule nucleating γ-TuSC assembles structures with 13-fold microtubule-like symmetry. Nature 466, 879–882 (2010).
Brilot, A. F. et al. CM1-driven assembly and activation of yeast γ-tubulin small complex underlies microtubule nucleation. eLife 10, e65168 (2021).
Wieczorek, M. et al. Biochemical reconstitutions reveal principles of human γ-TuRC assembly and function. J. Cell Biol. 220, e202009146 (2021).
Berman, A. Y. et al. A nucleotide binding-independent role for γ-tubulin in microtubule capping and cell division. J. Cell Biol. 222, e202204102 (2023).
Brouhard, G. J. et al. XMAP215 is a processive microtubule polymerase. Cell 132, 79–88 (2008).
Thawani, A., Kadzik, R. S. & Petry, S. XMAP215 is a microtubule nucleation factor that functions synergistically with the γ-tubulin ring complex. Nat. Cell Biol. 20, 575–585 (2018).
Guyomar, C. et al. Changes in seam number and location induce holes within microtubules assembled from porcine brain tubulin and in Xenopus egg cytoplasmic extracts. eLife 11, e83021 (2022).
Pierson, G. B., Burton, P. R. & Himes, R. H. Alterations in number of protofilaments in microtubules assembled in vitro. J. Cell Biol. 76, 223–228 (1978).
Unger, E., Böhm, K. J. & Vater, W. Structural diversity and dynamics of microtubules and polymorphic tubulin assemblies. Electron Microsc. Rev. 3, 355–395 (1990).
Roostalu, J. et al. The speed of GTP hydrolysis determines GTP cap size and controls microtubule stability. eLife 9, e51992 (2020).
Valdez, V., Ma, M., Gouveia, B., Zhang, R. & Petry, S. HURP facilitates spindle assembly by stabilizing microtubules and working synergistically with TPX2. Preprint at https://doi.org/10.1101/2023.12.18.571906 (2023).
Alushin, G. M. et al. High-resolution microtubule structures reveal the structural transitions in αβ-tubulin upon GTP hydrolysis. Cell 157, 1117–1129 (2014).
Zhang, R., LaFrance, B. & Nogales, E. Separating the effects of nucleotide and EB binding on microtubule structure. Proc. Natl Acad. Sci. USA 115, E6191–E6200 (2018).
LaFrance, B. J. et al. Structural transitions in the GTP cap visualized by cryo-electron microscopy of catalytically inactive microtubules. Proc. Natl Acad. Sci. USA 119, e2114994119 (2022).
Chaaban, S. & Brouhard, G. J. A microtubule bestiary: structural diversity in tubulin polymers. Mol. Biol. Cell 28, 2924–2931 (2017).
Kiewisz, R. et al. Three-dimensional structure of kinetochore-fibers in human mitotic spindles. eLife 11, e75459 (2022).
Guichard, P., Chrétien, D., Marco, S. & Tassin, A.-M. Procentriole assembly revealed by cryo-electron tomography. EMBO J. 29, 1565–1572 (2010).
Pamula, M. C. et al. High-resolution imaging reveals how the spindle midzone impacts chromosome movement. J. Cell Biol. 218, 2529–2544 (2019).
Nogales, E. & Zhang, R. Visualizing microtubule structural transitions and interactions with associated proteins. Curr. Opin. Struct. Biol. 37, 90–96 (2016).
Manka, S. W. & Moores, C. A. Microtubule structure by cryo-EM: snapshots of dynamic instability. Essays Biochem. 62, 737–751 (2018).
Barford, D. et al. Structure of the native γ-tubulin ring complex capping spindle microtubules. Preprint at https://doi.org/10.21203/rs.3.rs-3481382/v1 (2023).
Xu, Y. et al. Closure of the γ-tubulin ring complex by CDK5RAP2 activates microtubule nucleation. Preprint at https://doi.org/10.1101/2023.12.14.571518 (2023).
Ti, S.-C., Alushin, G. M. & Kapoor, T. M. Human β-tubulin isotypes can regulate microtubule protofilament number and stability. Dev. Cell 47, 175–190 (2018).
Miller, M. P., Asbury, C. L. & Biggins, S. A TOG protein confers tension sensitivity to kinetochore–microtubule attachments. Cell 165, 1428–1439 (2016).
Choi, Y.-K., Liu, P., Sze, S. K., Dai, C. & Qi, R. Z. CDK5RAP2 stimulates microtubule nucleation by the γ-tubulin ring complex. J. Cell Biol. 191, 1089–1095 (2010).
Fridy, P. C. et al. A robust pipeline for rapid production of versatile nanobody repertoires. Nat. Methods 11, 1253–1260 (2014).
Abmayr, S. M., Yao, T., Parmely, T. & Workman, J. L. Preparation of nuclear and cytoplasmic extracts from mammalian cells.Curr. Protoc. Mol. Biol. 12, 1.1–1.10 (2006).
Snijder, J. et al. Vitrification after multiple rounds of sample application and blotting improves particle density on cryo-electron microscopy grids. J. Struct. Biol. 198, 38–42 (2017).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005).
Cook, A. D., Manka, S. W., Wang, S., Moores, C. A. & Atherton, J. A microtubule RELION-based pipeline for cryo-EM image processing. J. Struct. Biol. 209, 107402 (2020).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Zivanov, J. et al. A Bayesian approach to single-particle electron cryo-tomography in RELION-4.0. eLife 11, e83724 (2022).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D Struct. Biol. 75, 861–877 (2019).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Pettersen, E. F., Goddard, T. D. & Huang, C. C. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Tan, Y. Z. et al. Addressing preferred specimen orientation in single-particle cryo-EM through tilting. Nat. Methods 14, 793–796 (2017).
Acknowledgements
This work was funded by a National Institutes of Health grant (GM130234) to T.M.K. A.A. was supported in part by a Pels Family Center Postdoctoral Fellowship at The Rockefeller University. Some of this work was performed at the Simons Electron Microscopy Center (SEMC) at the New York Structural Biology Center, with major support from the Simons Foundation (SF349247). We are grateful to R. Roeder for the gift of HeLa cell extracts. We are grateful to J. Mendez, SEMC, for help with data acquisition. We are grateful to M. Ebrahim, J. Sotiris H. Ng and the Evelyn Gruss Lipper Cryo-Electron Microscopy Resource Center for cryo-EM support. We are also grateful to H. A. Pasolli at the Electron Microscopy Resource Center, both at The Rockefeller University, for EM support. We are grateful to Y. Niu and G. M. Alushin at The Rockefeller University for help with processing microtubule protofilament structure.
Author information
Authors and Affiliations
Contributions
A.A., L.U. and T.M.K. conceived the study. A.A. and L.U. performed the experiments. A.A., L.U., A.X. and K.N. analyzed the data. T.M.K. supervised the study. A.A., L.U. and T.M.K. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
T.M.K is a co-founder of and has an ownership interest in RADD Pharmaceuticals, Inc. All other authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Rui Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Katarzyna Ciazynska, in collaboration with the Nature Structural & Molecular Biology team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Note added in proof: In a manuscript by Brito et al., Science 383, 870–876 (2024), the authors arrive at many similar conclusions regarding conformational changes from free to the microtubule bound form of γ-TuRC.
Extended data
Extended Data Fig. 1 TIRF-based optimization of γ-TuRC-dependent microtubule nucleation assay with slow GTP hydrolysing mutant of ɑ-tubulin, E254D.
(a) Transmission EM micrograph of negatively stained native γ-TuRC (representative image from 441 micrographs). (b) 2D class-averages showing two orientations of native γ-TuRC particles, top view (upper panel, 1182 particles) and side-view (bottom panel, 1048 particles) from negative stain EM. (c,d) 2D class averages for native γ-TuRC-capped minus-ends (177 ends, 125 from which were used for protofilament number analysis) (c) and for uncapped ends from spontaneously nucleated microtubules (183 ends) from cryo-EM (d). (e) SDS-PAGE analysis (Coomassie stained) of recombinant γ-TuRC-GFP following sucrose density gradient centrifugation. Expected positions for the components of the complex are indicated. (f,g) Images from a TIRF-microscopy sequence showing the γ-TuRC (green), tubulin (magenta) and merge channels after 2 minutes 30 s in the presence of chTOG (100 nM) and either wild type tubulin (10 µM) (f) or E254D (TUBA1B-E254D, TUBB3) tubulin (10 µM) (g). (h) Plot of the percentage (mean ± s.d.) of microtubules associated with a γ-TuRC-GFP puncta or not associated with a γ-TuRC-GFP puncta at 50 s post start of nucleation in the presence of E254D tubulin (10 µM) and chTOG (100 nM), total 95 microtubules, n = 3 independent experiments for each condition. (i) Plot of the cumulative number of microtubules (mean ± s.d.) nucleated by γ-TuRC in the presence of chTOG (100 nM) and either wild type tubulin or E254D tubulin (10 µM) over time, is shown. Data were fitted using linear regression, E254D tubulin (red line, total 193 microtubules) and wild type tubulin (blue line, no microtubules were detected). n = 4 independent experiments for each condition.
Extended Data Fig. 2 Cryo-EM data processing workflow for γ-TuRC-capped microtubule minus-end and free γ-TuRC.
A multi-step workflow is schematized.
Extended Data Fig. 3 Estimates of resolution and projection angle distribution for the overall γ-TuRC-capped microtubule minus-end and symmetry expanded α-α and γ-γ-tubulin density maps.
(a-d) Two views showing the Euler angle distribution (a,b), gold-standard Fourier shell correlation (FSC) curve (c) and directional FSC as estimated using the 3DFSC server62 (d) for the overall γ-TuRC-capped microtubule end density map presented in Figs. 1 and 2. (e-h) Two views showing the Euler angle distribution (e,f), gold-standard Fourier shell correlation (FSC) curve (g) and directional FSC as estimated using the 3DFSC server62 (h) for the symmetry expanded ɑ-ɑ and γ-γ-tubulin density map presented in Fig. 4.
Extended Data Fig. 4 Cryo-EM reconstruction of γ-TuRC-nucleated microtubule protofilament reveals a compacted lattice.
(a) Schematic for γ-TuRC-nucleated microtubule used for cryo-EM reconstruction (b, c) Cryo-EM density map for γ-TuRC-nucleated microtubule protofilament showing a view of the ɑ/β-tubulin dimer from the outside (b) and from within the lumen (c) (ɑ-tubulin: lighter green; β-tubulin: darker green). (d-e) Densities (mesh) for the nucleotide bound to ɑ-tubulin (GTP) (d) and β-tubulin (GDP) (e). (f-g) Models for 2 tubulin heterodimers from a γ-TuRC-nucleated microtubule protofilament and a protofilament from GDP-microtubule (PDB ID:6DPV) (f) or GMPCPP-microtubule (PDB ID:6DPU) (g) aligned at the first ɑ-tubulin. Distances between Cɑ atoms for 10 residues across 2 successive ɑ-tubulins were measured and averaged. (h) Four tubulin models rigid-body fitted into γ-TuRC-nucleated microtubule map at position 7 were used to measure distances between Cɑ atoms for 10 residues across two successive ɑ-tubulins.
Extended Data Fig. 5 Cryo-EM data processing workflow for a γ-TuRC-nucleated microtubule protofilament.
A multi-step workflow is schematized.
Extended Data Fig. 7 Estimates of resolution and projection angle distribution for the refined γ-TuRC-capped microtubule minus-end map and free γ-TuRC map.
(a-d) Two views showing the Euler angle distribution (a,b), gold-standard Fourier shell correlation (FSC) curve (c) and directional FSC as estimated by the 3DFSC server62 (d) for the refined γ-TuRC-capped microtubule end density map presented in Fig. 3. (e-h) Two views showing the Euler angle distribution (e,f), gold-standard Fourier shell correlation (FSC) curve (g) and directional FSC as estimated using the 3DFSC server62 (h) for the free recombinant γ-TuRC density map, corresponding to Extended Data Figure 8.
Extended Data Fig. 8 Comparison of γ-TuRC in its free and microtubule bound form and analyses of lateral α-α and β-β tubulin interactions.
(a, b) Comparison of the rigid-body fitted models of native γ-TuRC (gray) and recombinant γ-TuRC (colored, GCPs in pink, γ-tubulins in purple) showing top (a) and side views (b). (c, d) Comparison of the rigid-body fitted models of recombinant γ-TuRC (gray) and microtubule bound recombinant γ-TuRC (colored, GCPs in pink, γ-tubulins in purple) showing top (c) and side views (d). (e-h) Lateral interactions at the ɑ-tubulin-ɑ-tubulin interface (e,g) and β-tubulin-β-tubulin interface (ribbons) (f,h).
Supplementary information
Source data
Source Data Fig. 1
Plot source data
Source Data Extended Data Fig. 1
Plot source data
Source Data Extended Data Fig. 1
Unprocessed gel image.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Aher, A., Urnavicius, L., Xue, A. et al. Structure of the γ-tubulin ring complex-capped microtubule. Nat Struct Mol Biol (2024). https://doi.org/10.1038/s41594-024-01264-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41594-024-01264-z