Abstract
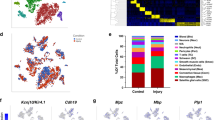
The peripheral nerve contains diverse cell types that support its proper function and maintenance. In this study, we analyzed multiple peripheral nerves using single-nuclei RNA sequencing, which allowed us to circumvent difficulties encountered in analyzing cells with complex morphologies via conventional single-cell methods. The resultant mouse peripheral nerve cell atlas highlights a diversity of cell types, including multiple subtypes of Schwann cells (SCs), immune cells and stromal cells. We identified a distinct myelinating SC subtype that expresses Cldn14, Adamtsl1 and Pmp2 and preferentially ensheathes motor axons. The number of these motor-associated Pmp2+ SCs is reduced in both an amyotrophic lateral sclerosis (ALS) SOD1G93A mouse model and human ALS nerve samples. Our findings reveal the diversity of SCs and other cell types in peripheral nerve and serve as a reference for future studies of nerve biology and disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
A total of 24 files, including 18 raw sequencing reads and six processed data such as expression matrices and RDS files for single-cell and single-nuclei atlases, have been deposited in the Gene Expression Omnibus repository under the reference series ID GSE182099. The Mpz-RiboTag and bulk RNA-seq data can be accessed through the sub-series ID GSE181858, and the single-nuclei data can be obtained through GSE182098. The glia portal can also be accessed through the link http://milbrandt.wustl.edu/glia-portal/. Source data are provided with this paper.
Code availability
The source code for data analysis and visualization in this study will be available online at https://github.com/aldrinyim/PNS-glia.
References
Fünfschilling, U. et al. Glycolytic oligodendrocytes maintain myelin and long-term axonal integrity. Nature 485, 517–522 (2012).
Nave, K. A. Myelination and support of axonal integrity by glia. Nature 468, 244–252 (2010).
Monk, K. R., Feltri, M. L. & Taveggia, C. New insights on Schwann cell development. Glia 63, 1376–1393 (2015).
Richard, L. et al. Endoneurial fibroblast-like cells. J. Neuropathol. Exp. Neurol. 71, 938–947 (2012).
Brushart, T. M. E. Preferential reinnervation of motor nerves by regenerating motor axons. J. Neurosci. 8, 1026–1031 (1988).
Madison, R. D., Sofroniew, M. V. & Robinson, G. A. Schwann cell influence on motor neuron regeneration accuracy. Neuroscience 163, 213–221 (2009).
Aguilar, S. V. et al. ImmGen at 15. Nat. Immunol. 21, 700–703 (2020).
Jesuraj, N. J. et al. Differential gene expression in motor and sensory Schwann cells in the rat femoral nerve. J. Neurosci. Res. 90, 96–104 (2012).
Carr, M. J. et al. Mesenchymal precursor cells in adult nerves contribute to mammalian tissue repair and regeneration. Cell Stem Cell 24, 240–256 (2019).
Toma, J. S. et al. Peripheral nerve single cell analysis identifies mesenchymal ligands that promote axonal growth. eNeuro 7, ENEURO.0066-20.2020 (2020).
Wolbert, J. et al. Redefining the heterogeneity of peripheral nerve cells in health and autoimmunity. Proc. Natl Acad. Sci. USA 117, 9466–9476 (2020).
Gerber, D. et al. Transcriptional profiling of mouse peripheral nerves to the single-cell level to build a sciatic nerve ATlas (SNAT). eLife 10, e58591 (2021).
Skelly, D. A. et al. Single-cell transcriptional profiling reveals cellular diversity and intercommunication in the mouse heart. Cell Rep. 22, 600–610 (2018).
Baron, M. et al. A single-cell transcriptomic map of the human and mouse pancreas reveals inter- and intra-cell population structure. Cell Syst. 3, 346–360 (2016).
Causey, G. & Barton, A. A. The cellular content of the endoneurium of peripheral nerve. Brain 82, 594–598 (1959).
Habib, N. et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955–958 (2017).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Peters, A. & Muir, A. R. The relationship between axons and Schwann cells during development of peripheral nerves in the rat. Q. J. Exp. Physiol. Cogn. Med. Sci. 44, 117–130 (1959).
Wang, Y., Teng, H. L. & Huang, Z. H. Repulsive migration of Schwann cells induced by Slit-2 through Ca2+-dependent RhoA-myosin signaling. Glia 61, 710–723 (2013).
Yoshino, J. E., Dinneen, M. P., Sprinkle, T. J. & DeVries, G. H. Localization of 2′,3′-cyclic nucleotide 3′-phosphodiesterase on cultured Schwann cells. Brain Res. 325, 199–203 (1985).
García-Villegas, R., López-Álvarez, L. E., Arni, S., Rosenbaum, T. & Morales, M. A. Identification and functional characterization of the promoter of the mouse sodium-activated sodium channel Nax gene (Scn7a). J. Neurosci. Res. 87, 2509–2519 (2009).
Ruiz-Martínez, J., Azcona, L. J., Bergareche, A., Martí-Massó, J. F. & Paisán-Ruiz, C. Whole-exome sequencing associates novel CSMD1 gene mutations with familial Parkinson disease. Neurol. Genet. 3, e177 (2017).
Friede, R. L. & Samorajski, T. Relation between the number of myelin lamellae and axon circumference in fibers of vagus and sciatic nerves of mice. J. Comp. Neurol. 130, 223–231 (1967).
Morita, K., Sasaki, H., Furuse, M. & Tsukita, S. Endothelial claudin: claudin-5/TMVCF constitutes tight junction strands in endothelial cells. J. Cell Biol. 147, 185–194 (1999).
Oliver, G. Lymphatic vasculature development. Nat. Rev. Immunol. 4, 35–45 (2004).
Wendling, O., Bornert, J. M., Chambon, P. & Metzger, D. Efficient temporally-controlled targeted mutagenesis in smooth muscle cells of the adult mouse. Genesis 47, 14–18 (2009).
Kaplan, L., Chow, B. W. & Gu, C. Neuronal regulation of the blood–brain barrier and neurovascular coupling. Nat. Rev. Neurosci. 21, 416–432 (2020).
Wang, P. L. et al. Peripheral nerve resident macrophages share tissue-specific programming and features of activated microglia. Nat. Commun. 11, 2552 (2020).
Ydens, E. et al. Profiling peripheral nerve macrophages reveals two macrophage subsets with distinct localization transcriptome and response to injury. Nat. Neurosci. 23, 676–689 (2020).
Sunderland, S. The connective tissues of peripheral nerves. Brain 88, 841–854 (1965).
Singhmar, P. et al. The fibroblast-derived protein PI16 controls neuropathic pain. Proc. Natl Acad. Sci. USA 117, 5463–5471 (2020).
Jeltsch, M. et al. CCBE1 enhances lymphangiogenesis via a disintegrin and metalloprotease with thrombospondin motifs-3–mediated vascular endothelial growth factor-C activation. Circulation 129, 1962–1971 (2014).
Hirose, T. et al. Immunohistochemical demonstration of EMA/Glut1-positive perineurial cells and CD34-positive fibroblastic cells in peripheral nerve sheath tumors. Mod. Pathol. 16, 293–298 (2003).
Richard, L., Védrenne, N., Vallat, J. M. & Funalot, B. Characterization of endoneurial fibroblast-like cells from human and rat peripheral nerves. J. Histochem. Cytochem. 62, 424–435 (2014).
Jessen, K. R. & Mirsky, R. The origin and development of glial cells in peripheral nerves. Nat. Rev. Neurosci. 6, 671–682 (2005).
Baima, J. & Krivickas, L. Evaluation and treatment of peroneal neuropathy. Curr. Rev. Musculoskelet. Med. 1, 147–153 (2008).
Herrmann, D. N., Griffin, J. W., Hauer, P., Cornblath, D. R. & McArthur, J. C. Epidermal nerve fiber density and sural nerve morphometry in peripheral neuropathies. Neurology 53, 1634–1640 (1999).
Yuan, H. & Silberstein, S. D. Vagus nerve and vagus nerve stimulation, a comprehensive review: part I. Headache 56, 71–78 (2016).
Zilic, L. et al. An anatomical study of porcine peripheral nerve and its potential use in nerve tissue engineering. J. Anat. 227, 302–314 (2015).
Velmeshev, D. et al. Single-cell genomics identifies cell type-specific molecular changes in autism. Science 364, 685–689 (2019).
Amberger, J. S., Bocchini, C. A., Schiettecatte, F., Scott, A. F. & Hamosh, A. OMIM.org: Online Mendelian Inheritance in Man (OMIM®), an online catalog of human genes and genetic disorders. Nucleic Acids Res. 43, D789–D798 (2015).
Tang, Y. et al. A genetic locus on chromosome 2q24 predicting peripheral neuropathy risk in type 2 diabetes: results from the ACCORD and BARI 2D studies. Diabetes 68, 1649–1662 (2019).
Sanders, S. J. et al. Progress in understanding and treating SCN2A-mediated disorders. Trends Neurosci. 41, 442–456 (2018).
Feldman, E. L., Nave, K. A., Jensen, T. S. & Bennett, D. L. H. New horizons in diabetic neuropathy: mechanisms, bioenergetics, and pain. Neuron 93, 1296–1313 (2017).
Vandoorne, T., de Bock, K. & van den Bosch, L. Energy metabolism in ALS: an underappreciated opportunity? Acta Neuropathol. 135, 489–509 (2018).
Bouçanova, F. et al. Disrupted function of lactate transporter MCT1, but not MCT4, in Schwann cells affects the maintenance of motor end-plate innervation. Glia 69, 124–136 (2021).
Hoke, A. Schwann cells express motor and sensory phenotypes that regulate axon regeneration. J. Neurosci. 26, 9646–9655 (2006).
Sherman, D. L., Fabrizi, C., Gillespie, C. S. & Brophy, P. J. Specific disruption of a Schwann cell dystrophin-related protein complex in a demyelinating neuropathy. Neuron 30, 677–687 (2001).
Zenker, J. et al. A role of peripheral myelin protein 2 in lipid homeostasis of myelinating Schwann cells. Glia 62, 1502–1512 (2014).
Rossi, J. et al. Melanocortin-4 receptors expressed by cholinergic neurons regulate energy balance and glucose homeostasis. Cell Metab. 13, 195–204 (2011).
Blum, J. A. et al. Single-cell transcriptomic analysis of the adult mouse spinal cord reveals molecular diversity of autonomic and skeletal motor neurons. Nat. Neurosci. 24, 572–583 (2021).
Guttenplan, K. A. et al. Knockout of reactive astrocyte activating factors slows disease progression in an ALS mouse model. Nature Commun. 11, 3753 (2020).
Hatzipetros, T. et al. A quick phenotypic neurological scoring system for evaluating disease progression in the SOD1-G93A mouse model of ALS. J. Visualized Exp. 2015, 1–6 (2015).
Gould, T. W. et al. Complete dissociation of motor neuron death from motor dysfunction by Bax deletion in a mouse model of ALS. J. Neurosci. 26, 8774–8786 (2006).
Hong, Y. Bin et al. A mutation in PMP2 causes dominant demyelinating Charcot-Marie-Tooth neuropathy. PLoS Genet. 12, e1005829 (2016).
Motley, W. W. et al. De novo PMP2 mutations in families with type 1 Charcot-Marie-Tooth disease. Brain 139, 1649–1656 (2016).
Ruskamo, S. et al. Cryo-EM, X-ray diffraction, and atomistic simulations reveal determinants for the formation of a supramolecular myelin-like proteolipid lattice. J. Biol. Chem. 295, 8692–8705 (2020).
Seetharaman, A. et al. MADD-4 is a secreted cue required for midline-oriented guidance in Caenorhabditis elegans. Dev. Cell 21, 669–680 (2011).
Domènech-Estévez, E. et al. Distribution of monocarboxylate transporters in the peripheral nervous system suggests putative roles in lactate shuttling and myelination. J. Neurosci. 35, 4151–4156 (2015).
Madison, R. D., Archibald, S. J. & Brushart, T. M. Reinnervation accuracy of the rat femoral nerve by motor and sensory neurons. J. Neurosci. 16, 5698–5703 (1996).
Llewellyn, M. E., Thompson, K. R., Deisseroth, K. & Delp, S. L. Orderly recruitment of motor units under optical control in vivo. Nat. Med. 16, 1161–1165 (2010).
Nijssen, J., Comley, L. H. & Hedlund, E. Motor neuron vulnerability and resistance in amyotrophic lateral sclerosis. Acta Neuropathol. 133, 863–885 (2017).
Hegedus, J., Putman, C. T. & Gordon, T. Time course of preferential motor unit loss in the SOD1G93A mouse model of amyotrophic lateral sclerosis. Neurobiol. Dis. 28, 154–164 (2007).
Frey, D. et al. Early and selective loss of neuromuscular synapse subtypes with low sprouting competence in motoneuron diseases. J. Neurosci. 20, 2534–2542 (2000).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Arisdakessian, C., Poirion, O., Yunits, B., Zhu, X. & Garmire, L. X. DeepImpute: an accurate, fast, and scalable deep neural network method to impute single-cell RNA-seq data. Genome Biol. 20, 211 (2019).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Kanehisa, M., Goto, S., Sato, Y., Furumichi, M. & Tanabe, M. KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res. 40, D109–D114 (2012).
Sanz, E. et al. Cell-type-specific isolation of ribosome-associated mRNA from complex tissues. Proc. Natl Acad. Sci. USA 106, 13939–13944 (2009).
Ximerakis, M. et al. Single-cell transcriptomic profiling of the aging mouse brain. Nat. Neurosci. 22, 1696–1708 (2019).
Acknowledgements
The authors thank all our collaborators at the Washington University School of Medicine for their advice and discussion. We thank R. Head, R. Barve and P. Cliften at GTAC@MGI and the McDonnell Genome Institute for discussion and assistance with RNA-seq. The authors are grateful for assistance from G. Randolph’s laboratory and B. Zinselmeyer for confocal imaging and use of equipment (supported by National Institutes of Health (NIH) grants R01DK119147 and R37AI049653 to G.R.). The authors thank M. Shabsovich, T. Shen and C. Kreple for ALS SOD1G93A mice, M. Ireland for curating patient samples (supported by NIH grant R01NS078398 to T.M.) and R. Schmidt for curating clinical pathology reports and many helpful discussions. The authors thank R. McClarney and C. Menendez for experimental assistance and members of the Milbrandt and DiAntonio laboratories for helpful comments and discussion. This project was supported by NIH grants RF1MH117070 and RO1GM123203 to R.D.M. and R01NS105645 and R01AG013730 to J.M.
Author information
Authors and Affiliations
Contributions
A.Y. designed and performed both the animal and sequencing experiments, derived the nuclei isolation protocol, analyzed both sequencing and imaging data and wrote the manuscript. P.L.W. designed and performed the animal experiments, analyzed the imaging data and wrote the manuscript. J.R.B. and A.S. performed the RiboTag experiments; A.S. helped with the animal experiments; and J.R.B. helped with RNA FISH experiments. A.H. helped with the animal experiments. C.L. and T.M. prepared the ALS animal and human samples. R.M. and J.M. designed and supervised the experiments, acquired funding and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
J.M. is a cofounder, consultant and shareholder of Disarm Therapeutics, a wholly owned subsidiary of Eli Lilly & Company. All other authors declare no competing financial interests.
Peer review
Peer review information
Nature Neuroscience thanks Goncalo Castelo-Branco and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Generation and analysis of sciatic nerve and immune cell atlases.
(a) Gating strategy for sorting GFP+ nuclei from Sun1-GFP/+:Act-Cre/+ sciatic nerves. Nuclei were isolated from single cell suspensions containing pooled sciatic nerves from at least 5 Sun1-GFP/+:Act-Cre/+ mice. Purified nuclei were selected based on GFP and Hoechst expression. (b-f) Intronic reads contain biological information. Single nuclei atlases were generated by using either exonic or intronic reads, and CCA analysis was performed to align the two data sets. (b) The two atlases showed high level of concordance. c-f, Contribution of intronic reads to signature genes in nmSCs. (c) Csmd1 and (d) Scn7a were found to mark nmSCs. (e) Over 90% of the UMIs were from intronic reads for Csmd1. (f) Approximately 60% of the UMIs were from intronic reads for Scn7a. (g) Gating strategy for sorting CD45+ cells from sciatic nerves. Sciatic nerves were pooled from 20 Cx3cr1-GFP/+ mice and prepared for FACS sorting. Cells were stained with Propidium Iodide (PI) to select for viable cells, and with CD45 antibody to select for CD45+ population. (h) Expression of lymphangiogenesis-related genes in epineurial fibroblasts. Adamts3, Adamts14, Ccbe1, Pdgfra and Adamts12 expression in the ActB-Sun1 sciatic nerve atlas.
Extended Data Fig. 2 Generation and analysis of Mpz-enriched single nuclei atlas.
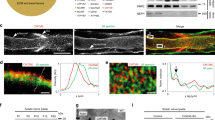
(a) Gating strategy for sorting GFP+ nuclei from Sun1-GFP/+:Mpz-Cre/+ mice. Nuclei were isolated from single cell suspensions containing pooled sciatic nerves from at least 5 Sun1-GFP/+:Act-Cre/+ mice. Purified nuclei were selected based on GFP and Hoechst expression. (b) tSNE plot with 3,047 nuclei from Sun1-GFP/+;Mpz-Cre/+ defines three GFP+ cell types – endoneurial fibroblasts, nmSCs and mSCs. (c) Differential expression analysis from Mpz-Sun1 atlas shows signature gene expression that defines endoneurial fibroblasts, including Col15a1, Lama2 and Egfr. Each column represents a gene and each row represents a nucleus, genes with high expression level are shown in yellow and low expression in purple. (d) GO analysis showing that endoneurial fibroblast markers (122 genes) are enriched for components related to collagen-containing extracellular matrix, extracellular matrix and formation of axon, neurons and synapse, percentage of GO recovery is plotted against the false discovery rate (FDR) and selected GO terms are highlighted in color. (e,f) RNA-FISH analysis of Prx, Ngfr, and Cd34 expression. (scale bar=50um) e, Representative expression of Prx and Ngfr, indicating that myelinating Schwann cells do not express Ngfr. f, Representative expression of Prx and Cd34, indicating that myelinating Schwann cells do not express Cd34. (g) RNA-FISH analysis of Csmd1 and Scn7a. Representative overlapping expression of Csmd1 (green) and Scn7a (red) in adult sciatic nerves (scale bar=50um).
Extended Data Fig. 3 Pathway-based transcriptional module analysis.
(a) Expression of all genes in the axon guidance pathway (map04360) defined by the KEGG database are shown in here, with red indicating high expression and blue indicate no expression. Many of the genes are more abundantly expressed in the Pmp2+ mSCs. (b) tSNE plot showing the expression of genes in the axon guidance pathway. (c) tSNE plot showing the expression of genes in fatty acid biosynthesis pathway (map00061). (d) tSNE plot showing the expression of genes in the sphingolipid signaling pathway (map04071).
Extended Data Fig. 4 Pmp2+ Schwann cells preferentially ensheath Chat+ axons.
(a) Validation of ChAT-Cre-tdT expression specificity. Representative expression of PMP2 (yellow) and ChAT-Ab (white) in ChAT-Cre-tdT (red) sciatic nerves. The two panels on the right have shown almost complete overlap between the antibody (white) and tdT reporter (red) for ChAT (scale bar = 50um, n = 3 biologically independent mice). (b) ChAT+ axons are mostly ensheathed by Pmp2+ SCs. Representative imaging showing the expression of MBP (blue) and PMP2 (green) in ChAT-tdT (red) sciatic nerve. (scale bar = 50um, n = 3 biologically independent mice). (c) Quantification of percentage of ChAT+ axons ensheathed by a Pmp2+ SCs in three different nerve types. Two-way ANOVA with multiple comparisons. Sciatic vs Sural: P = 0.0159, Sciatic vs Femoral: P = 0.0456, Sural vs Femoral: P = 0.0009. Data are mean+/− SD (n = 3 biologically independent mice).
Extended Data Fig. 5 SOD1G93A mice show reduced PMP2 expression in SCs.
Representative imaging of PMP2+ SCs (yellow), b-TUB+ axons (cyan) and MBP+ SCs (gray) in sciatic, femoral, and sural nerves in SOD1G93A mice. Note that femoral nerve is severely degenerated as shown by MBP staining. Sciatic nerve remains relatively intact with depletion of PMP2, while sural nerve has intact axons with MBP+ SCs ensheathing them (scale bar = 50um, n = 3 biologically independent mice).
Supplementary information
Supplementary Information
Supplementary Figs. 1–8 and Supplementary Table 3
Supplementary Table 1
SC-enriched atlas. Differential expression analysis
Supplementary Table 2
KEGG expression module analysis
Supplementary Table 4
Hereditary peripheral neuropathic genes
Supplementary Table 5
List of RNA FISH probes
Source data
Source Data Fig. 5
Statistical Source Data
Source Data Fig. 6
Statistical Source Data
Source Data Fig. 7
Statistical Source Data
Source Data Extended Data Fig. 4
Statistical Source Data
Rights and permissions
About this article
Cite this article
Yim, A.K.Y., Wang, P.L., Bermingham, J.R. et al. Disentangling glial diversity in peripheral nerves at single-nuclei resolution. Nat Neurosci 25, 238–251 (2022). https://doi.org/10.1038/s41593-021-01005-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-021-01005-1
This article is cited by
-
Deletion of CD47 from Schwann cells and macrophages hastens myelin disruption/dismantling and scavenging in Schwann cells and augments myelin debris phagocytosis in macrophages
Journal of Neuroinflammation (2023)
-
Integration of single-nuclei RNA-sequencing, spatial transcriptomics and histochemistry defines the complex microenvironment of NF1-associated plexiform neurofibromas
Acta Neuropathologica Communications (2023)
-
Tissue dissociation for single-cell and single-nuclei RNA sequencing for low amounts of input material
Frontiers in Zoology (2022)