Abstract
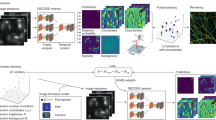
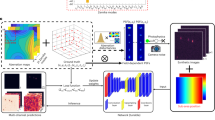
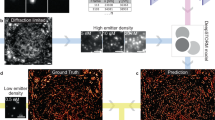
An outstanding challenge in single-molecule localization microscopy is the accurate and precise localization of individual point emitters in three dimensions in densely labeled samples. One established approach for three-dimensional single-molecule localization is point-spread-function (PSF) engineering, in which the PSF is engineered to vary distinctively with emitter depth using additional optical elements. However, images of dense emitters, which are desirable for improving temporal resolution, pose a challenge for algorithmic localization of engineered PSFs, due to lateral overlap of the emitter PSFs. Here we train a neural network to localize multiple emitters with densely overlapping Tetrapod PSFs over a large axial range. We then use the network to design the optimal PSF for the multi-emitter case. We demonstrate our approach experimentally with super-resolution reconstructions of mitochondria and volumetric imaging of fluorescently labeled telomeres in cells. Our approach, DeepSTORM3D, enables the study of biological processes in whole cells at timescales that are rarely explored in localization microscopy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Code availability
Code is made publicly available at https://github.com/EliasNehme/DeepSTORM3D.
Change history
26 June 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41592-020-0910-0
References
Katayama, Y. et al. Real-time nanomicroscopy via three-dimensional single-particle tracking. Chem. Phys. Chem. 10, 2458–2464 (2009).
Manzo, C. & Garcia-Parajo, M. F. A review of progress in single particle tracking: from methods to biophysical insights. Rep. Prog. Phys. 78, 124601 (2015).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Hess, S. T., Girirajan, T. P. & Mason, M. D. Ultra-high resolution imaging by fluorescence photoactivation localization microscopy. Biophysical J. 91, 4258–4272 (2006).
Rust, M. J., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (storm). Nat. Methods 3, 793–796 (2006).
Sahl, S. J. & Moerner, W. Super-resolution fluorescence imaging with single molecules. Curr. Opin. Struct. Biol. 23, 778–787 (2013).
von Diezmann, A., Shechtman, Y. & Moerner, W. Three-dimensional localization of single molecules for super-resolution imaging and single-particle tracking. Chem. Rev. 117, 7244–7275 (2017).
Pavani, S. R. P. et al. Three-dimensional, single-molecule fluorescence imaging beyond the diffraction limit by using a Double-Helix point spread function. Proc. Natl Acad. Sci. USA 106, 2995–2999 (2009).
Huang, B., Wang, W., Bates, M. & Zhuang, X. Three-dimensional super-resolution imaging by stochastic optical reconstruction microscopy. Science 319, 810–813 (2008).
Shechtman, Y., Sahl, S. J., Backer, A. S. & Moerner, W. Optimal point spread function design for 3D imaging. Phys. Rev. Lett. 113, 133902 (2014).
Backer, A. S. & Moerner, W. Extending single-molecule microscopy using optical Fourier processing. J. Phys. Chem. B 118, 8313–8329 (2014).
Liu, S., Kromann, E. B., Krueger, W. D., Bewersdorf, J. & Lidke, K. A. Three-dimensional single-molecule localization using a phase retrieved pupil function. Opt. express 21, 29462–29487 (2013).
Babcock, H. P. & Zhuang, X. Analyzing single molecule localization microscopy data using cubic splines. Sci. Rep. 7, 552 (2017).
Li, Y. et al. Real-time 3D single-molecule localization using experimental point-spread functions. Nat. Methods 15, 367 (2018).
Aristov, A., Lelandais, B., Rensen, E. & Zimmer, C. Zola-3D allows flexible 3D localization microscopy over an adjustable axial range. Nat. Commun. 9, 2409 (2018).
Ferdman, B. et al. VIPR: vectorial implementation of phase retrieval for fast and accurate microscopic pixel-wise pupil estimation. Opt. Express 28, 10179–10198 (2020).
Shechtman, Y., Weiss, L. E., Backer, A. S., Sahl, S. J. & Moerner, W. Precise three-dimensional scan-free multiple-particle tracking over large axial ranges with tetrapod point spread functions. Nano Lett. 15, 4194–4199 (2015).
Min, J. et al. Falcon: fast and unbiased reconstruction of high-density super-resolution microscopy data. Sci. Rep. 4, 4577 (2014).
Boyd, N., Schiebinger, G. & Recht, B. The alternating descent conditional gradient method for sparse inverse problems. SIAM J. Optim. 27, 616–639 (2017).
Nehme, E., Weiss, L. E., Michaeli, T. & Shechtman, Y. Deep-storm: super-resolution single-molecule microscopy by deep learning. Optica 5, 458–464 (2018).
Sage, D. et al. Super-resolution fight club: assessment of 2D and 3D single-molecule localization microscopy software. Nat. Methods 16, 387 (2019).
Rivenson, Y., Zhang, Y., Günaydın, H., Teng, D. & Ozcan, A. Phase recovery and holographic image reconstruction using deep learning in neural networks. Light-Sci. Appl. 7, 17141 (2018).
Nguyen, T., Xue, Y., Li, Y., Tian, L. & Nehmetallah, G. Deep learning approach for Fourier ptychography microscopy. Opt. Express 26, 26470–26484 (2018).
Weigert, M. et al. Content-aware image restoration: pushing the limits of fluorescence microscopy. Nat. Methods 15, 1090 (2018).
Ounkomol, C., Seshamani, S., Maleckar, M. M., Collman, F. & Johnson, G. R. Label-free prediction of three-dimensional fluorescence images from transmitted-light microscopy. Nat. Methods 15, 917–920 (2018).
Krull, A., Buchholz, T.-O. & Jug, F. Noise2void-learning denoising from single noisy images. in Proc. IEEE Conference on Computer Vision and Pattern Recognition (eds. Davis, L., Torr, P. & Zhu, S. C.) 2129–2137 (2019).
Falk, T. et al. U-net: deep learning for cell counting, detection, and morphometry. Nat. Methods 16, 67–70 (2019).
Rivenson, Y. et al. Phasestain: the digital staining of label-free quantitative phase microscopy images using deep learning. Light-Sci. Appl. 8, 23 (2019).
Liu, T. et al. Deep learning-based super-resolution in coherent imaging systems. Sci. Rep. 9, 3926 (2019).
Smith, J. T. et al. Fast fit-free analysis of complex fluorescence lifetime imaging via deep learning. Proc. Natl Acad. Sci. USA 116, 24019–24030 (2019).
Boyd, N., Jonas, E., Babcock, H. P. & Recht, B. DeepLoco: fast 3D localization microscopy using neural networks. Preprint at bioRxiv https://doi.org/10.1101/267096 (2018).
Ouyang, W., Aristov, A., Lelek, M., Hao, X. & Zimmer, C. Deep learning massively accelerates super-resolution localization microscopy. Nat. Biotechnol. 36, 460–468 (2018).
Diederic, B., Then, P., Jügler, A., Förster, R. & Heintzmann, R. cellSTORM: cost-effective super-resolution on a cellphone using dSTORM. PloS ONE 14, e0209827 (2019).
Newby, J. M., Schaefer, A. M., Lee, P. T., Forest, M. G. & Lai, S. K. Convolutional neural networks automate detection for tracking of submicron-scale particles in 2D and 3D. Proc. Natl Acad. Sci. USA 115, 9026–9031 (2018).
Zelger, P. et al. Three-dimensional localization microscopy using deep learning. Opt. Express 26, 33166–33179 (2018).
Liu, K. et al. Fast 3D cell tracking with wide-field fluorescence microscopy through deep learning. Preprint at https://arXiv.org/abs/1805.05139 (2018).
Hershko, E., Weiss, L. E., Michaeli, T. & Shechtman, Y. Multicolor localization microscopy and point-spread-function engineering by deep learning. Opt. Express 27, 6158–6183 (2019).
Speiser, A., Turaga, S. C. & Macke, J. H. Teaching deep neural networks to localize sources in super-resolution microscopy by combining simulation-based learning and unsupervised learning. Preprint at https://arXiv.org/abs/1907.00770 (2019).
Zhang, P. et al. Analyzing complex single-molecule emission patterns with deep learning. Nat. methods 15, 913 (2018).
Chakrabarti, A. Learning sensor multiplexing design through back-propagation. in Advances in Neural Information Processing Systems (eds. Lee, D. D. et al.) 3081–3089 (Curran Associates, 2016).
Horstmeyer, R., Chen, R. Y., Kappes, B. & Judkewitz, B. Convolutional neural networks that teach microscopes how to image. Preprint at https://arXiv.org/abs/1709.07223 (2017).
Turpin, A., Vishniakou, I. & D Seelig, J. Light-scattering control in transmission and reflection with neural networks. Opt. Express 26, 30911–30929 (2018).
Haim, H., Elmalem, S., Giryes, R., Bronstein, A. M. & Marom, E. Depth estimation from a single image using deep learned phase coded mask. IEEE Trans. Comput. Imaging 4, 298–310 (2018).
He, L., Wang, G. & Hu, Z. Learning depth from single images with deep neural network embedding focal length. IEEE Trans. Image Process. 27, 4676–4689 (2018).
Sitzmann, V. et al. End-to-end optimization of optics and image processing for achromatic extended depth of field and super-resolution imaging. ACM Trans. Graph. 37, 114 (2018).
Chang, J. & Wetzstein, G. Deep optics for monocular depth estimation and 3D object detection. in Proc. IEEE International Conference on Computer Vision (eds. Lee, K. M. et al.) 10193–10202 (2019).
Wu, Y., Boominathan, V., Chen, H., Sankaranarayanan, A. & Veeraraghavan, A. Phasecam3D: learning phase masks for passive single view depth estimation. in IEEE International Conference on Computational Photography (ed. Nedevschi, S.) 1–12 (2019).
Shechtman, Y., Weiss, L. E., Backer, A. S., Lee, M. Y. & Moerner, W. Multicolour localization microscopy by point-spread-function engineering. Nat. Photonics 10, 590 (2016).
Bickel, P. J. & Doksum, K. A. Mathematical Statistics: Basic Ideas and Selected Topics, Volumes I-II Package (Chapman and Hall/CRC, 2015).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Bronshtein, I. et al. Loss of lamin A function increases chromatin dynamics in the nuclear interior. Nat. Commun. 6, 8044 (2015).
Nahidiazar, L., Agronskaia, A. V., Broertjes, J., van den Broek, B. & Jalink, K. Optimizing imaging conditions for demanding multi-color super resolution localization microscopy. PLoS ONE 11, e0158884 (2016).
Ovesný, M., Křížek, P., Borkovec, J., Švindrych, Z. & Hagen, G. M. ThunderSTORM: a comprehensive ImageJ plug-in for PALM and STORM data analysis and super-resolution imaging. Bioinformatics 30, 2389–2390 (2014).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676 (2012).
Yu, F. & Koltun, V. Multi-scale context aggregation by dilated convolutions. Preprint at https://arXiv.org/abs/1511.07122v3 (2016).
Acknowledgements
We thank the Garini laboratory (Bar-Ilan University) for the U2OS cells, lmna−/− MEFs and the plasmid encoding for DsRed-hTRF1. We thank J. Ries for help with the application of SMAP-2018 to Tetrapod PSFs. We gratefully acknowledge the support of the NVIDIA Corporation with the donation of the Titan V GPU used for this research. We thank the staff of the Micro-Nano-Fabrication and Printing Unit at the Technion for their assistance with the phase mask fabrication. We thank Google for the research cloud units provided to accelerate this research. E.N., O.A., B.F. and R.O. are supported by H2020 European Research Council Horizon 2020 (802567); T.M. is supported by the Israel Science Foundation (grant no. 852/17) and by the Ollendorff Foundation; R.G. and O.A. are supported by the Israel Science Foundation (grant no. 450/18); Y.S. is supported by the Technion-Israel Institute of Technology Career Advancement Chairship; L.E.W. and Y.S. are supported by the Zuckerman Foundation. D.F. is supported by Google.
Author information
Authors and Affiliations
Contributions
E.N., D.F., T.M. and Y.S. conceived the approach. E.N. performed the simulations and analyzed the data with contributions from all authors. E.N., R.G., B.F., L.E.W., O.A. and T.N. collected the data. R.O. fabricated the physical phase mask. T.N. prepared MEF cells. E.N., D.F., L.E.W., T.M. and Y.S. wrote the paper with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Rita Strack was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Notes 1–14.
Supplementary Video 1
Localizations overlaid on experimental frames. This movie shows 70 representative experimental frames followed by an overlay of their regenerated images using the recovered 3D positions by the CNN (Fig. 3a in the main text). Note that the experimental frames are shown before and after the regenerated images for ease of visualization. The STORM experiment was repeated three times, twice analyzing 20,000 frames of two different cells from the same cell culture and once analyzing 10,000 frames of a cell from a different cell culture all leading to similar performance. Scale bar, 5 μm.
Supplementary Video 2
Rotating 3D rendering of the recovered mitochondria. This movie shows a 3D rendering of the super-resolved mitochondria spanning a 4-μm axial range (Fig. 3b in the main text). The z range is rendered with a scaling factor of 2 to ease axial visualization. Scale bar, 5 μm.
Supplementary Video 3
Sweep through the axial slices of the recovered mitochondria. This movie shows a sweep through 33 nm axial slices of the rendered 3D histogram for the mitochondria data (Fig. 3b in the main text). Scale bar, 5 μm.
Supplementary Video 4
Phase mask learning via backpropagation. This movie shows the phase mask (left) and the corresponding PSF (right) being learned over training iterations (Fig. 4c in the main text). Note that the phase mask is initialized to zero modulation, meaning the standard microscope PSF. Scale bar, 2 μm.
Supplementary Video 5
Rotating Telomere z stack without a mask. This movie shows a 3D rendering of the telomere data z stack without the application of a phase mask (Fig. 5b in the main text). As clearly shown in the rendered PSFs, the telomeres exhibit different sizes and intensities. The experiment was repeated independently for n = 10 U2OS cells all showing similar characteristics. Scale bar, 5 μm.
Supplementary Video 6
Super-critical angle fluorescence (SAF) light effect on the learned PSF. This movie shows the effect of the SAF light on the experimental PSF with the learned phase mask. Upper panel shows the experimental PSF (left), the result of VIPR45, the vectorial model assuming dipole emission, and the scalar model assuming isotropic emission. The lower panel shows the difference from the experimental measurement for each model. The SAF light effect is indicated in the middle of the axial range with a red arrow. Scale bar, 2 μm.
Supplementary Video 7
Volumetric tracking of telomeres in MEF cells. This movie shows 3D tracking of telomeres in live MEF cells over a period of 50 s using the learned phase mask (Fig. 6a in the main text). White sticks point to the emitter being tracked. Time is encoded in color. The results indicate that individual telomeres exhibit different types of movements. The experiment was repeated independently for n = 10 MEF cells all showing similar characteristics and performance. Scale bar, 2 μm.
Rights and permissions
About this article
Cite this article
Nehme, E., Freedman, D., Gordon, R. et al. DeepSTORM3D: dense 3D localization microscopy and PSF design by deep learning. Nat Methods 17, 734–740 (2020). https://doi.org/10.1038/s41592-020-0853-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-0853-5
This article is cited by
-
Temporal analysis of relative distances (TARDIS) is a robust, parameter-free alternative to single-particle tracking
Nature Methods (2024)
-
DeepDOF-SE: affordable deep-learning microscopy platform for slide-free histology
Nature Communications (2024)
-
Depth-enhanced high-throughput microscopy by compact PSF engineering
Nature Communications (2024)
-
High-density volumetric super-resolution microscopy
Nature Communications (2024)
-
Computational coherent Raman scattering imaging: breaking physical barriers by fusion of advanced instrumentation and data science
eLight (2023)