Abstract
Single-molecule localization microscopy is a powerful tool for visualizing subcellular structures, interactions and protein functions in biological research. However, inhomogeneous refractive indices inside cells and tissues distort the fluorescent signal emitted from single-molecule probes, which rapidly degrades resolution with increasing depth. We propose a method that enables the construction of an in situ 3D response of single emitters directly from single-molecule blinking datasets, and therefore allows their locations to be pinpointed with precision that achieves the Cramér-Rao lower bound and uncompromised fidelity. We demonstrate this method, named in situ PSF retrieval (INSPR), across a range of cellular and tissue architectures, from mitochondrial networks and nuclear pores in mammalian cells to amyloid-β plaques and dendrites in brain tissues and elastic fibers in developing cartilage of mice. This advancement expands the routine applicability of super-resolution microscopy from selected cellular targets near coverslips to intra- and extracellular targets deep inside tissues.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding authors upon request. Example data are available in software packages. 3D point clouds resolved by INSPR for Supplementary Videos 4–9 are provided from figshare (https://doi.org/10.6084/m9.figshare.11962764).
Code availability
The INSPR toolbox for in situ model estimation and 3D localization is available as Supplementary Software. INSPR works for commonly used biplane and astigmatism configurations. Further updates will be made freely available at https://github.com/HuanglabPurdue/INSPR. The software package features an easy-to-use user interface including all steps of 3D single-molecule localization from INSPR model generation, pupil-based 3D localization (including both CPU and GPU versions), drift correction and volume alignment to super-resolution image reconstruction.
References
Hell, S. W. & Wichmann, J. Breaking the diffraction resolution limit by stimulated emission: stimulated-emission-depletion fluorescence microscopy. Opt. Lett. 19, 780–782 (1994).
Wu, Y. & Shroff, H. Faster, sharper, and deeper: structured illumination microscopy for biological imaging. Nat. Methods 15, 1011–1019 (2018).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Hess, S. T., Girirajan, T. P. K. & Mason, M. D. Ultra-high resolution imaging by fluorescence photoactivation localization microscopy. Biophys. J. 91, 4258–4272 (2006).
Rust, M. J., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–796 (2006).
Huang, F. et al. Ultra-high resolution 3D imaging of whole cells. Cell 166, 1028–1040 (2016).
Liu, T.-L. et al. Observing the cell in its native state: imaging subcellular dynamics in multicellular organisms. Science 360, eaaq1392 (2018).
Guo, Y. et al. Visualizing intracellular organelle and crytoskeletal interactions at nanoscale resolution on millisecond timescales. Cell 175, 1430–1442 (2018).
Schlichthaerle, T. et al. Direct visualization of single nuclear pore complex proteins using genetically-encoded probes for DNA-PAINT. Angew. Chem. Int. Ed. 58, 1–6 (2019).
Juette, M. F. et al. Three-dimensional sub-100 nm resolution fluorescence microscopy of thick samples. Nat. Methods 5, 527–529 (2008).
Huang, B., Wang, W., Bates, M. & Zhuang, X. Three-dimensional super-resolution imaging by stochastic optical reconstruction microscopy. Science 319, 810–813 (2008).
Pavani, S. R. P. et al. Three-dimensional, single-molecule fluorescence imaging beyond the diffraction limit by using a double-helix point spread function. Proc. Natl Acad. Sci. USA 106, 2995–2999 (2009).
Aquino, D. et al. Two-color nanoscopy of three-dimensional volumes by 4Pi detection of stochastically switched fluorophores. Nat. Methods 8, 353–359 (2011).
Bon, P. et al. Self-interference 3D super-resolution microscopy for deep tissue investigations. Nat. Methods 15, 449–454 (2018).
Sage, D. et al. Super-resolution fight club: assessment of 2D and 3D single-molecule localization microscopy software. Nat. Methods 16, 387–395 (2019).
Small, A. & Stahlheber, S. Fluorophore localization algorithms for super-resolution microscopy. Nat. Methods 11, 267–279 (2014).
Nehme, E., Weiss, L. E., Michaeli, T. & Shechtman, Y. Deep-STORM: super-resolution single-molecule microscopy by deep learning. Optica 5, 458–464 (2018).
Zhang, P. et al. Analyzing complex single-molecule emission patterns with deep learning. Nat. Methods 15, 913–916 (2018).
Liu, S., Kromann, E. B., Krueger, W. D., Bewersdorf, J. & Lidke, K. A. Three dimensional single molecule localization using a phase retrieved pupil function. Opt. Express 21, 29462–29487 (2013).
Petrov, P. N., Shechtman, Y. & Moerner, W. E. Measurement-based estimation of global pupil functions in 3D localization microscopy. Opt. Express 25, 7945–7959 (2017).
Li, Y. et al. Real-time 3D single-molecule localization using experimental point spread functions. Nat. Methods 15, 367–369 (2018).
Li, Y., Wu, Y.-L., Hoess, P., Mund, M. & Ries, J. Depth-dependent PSF calibration and aberration correction for 3D single-molecule localization. Biomed. Opt. Express 10, 2708–2718 (2019).
Cabriel, C., Bourg, N., Dupuis, G. & Lévêque-Fort, S. Aberration-accounting calibration for 3D single-molecule localization microscopy. Opt. Lett. 43, 174–177 (2018).
Aristov, A., Lelandais, B., Rensen, E. & Zimmer, C. ZOLA-3D allows flexible 3D localization microscopy over an adjustable axial range. Nat. Commun. 9, 2409 (2018).
Booth, M. J., Andrade, D., Burke, D., Patton, B. & Zurauskas, M. Aberrations and adaptive optics in super-resolution microscopy. Microscopy 64, 251–261 (2015).
Dempster, A. P., Laird, N. M. & Rubin, D. B. Maximum likelihood from incomplete data via the EM algorithm. J. R. Stat. Soc. Ser. B Methodol. 39, 1–38 (1977).
Hartigan, J. A. & Wong, M. A. A k-means clustering algorithm. J. R. Stat. Soc. Ser. C. Appl. Stat. 28, 100–108 (1979).
Hanser, B. M., Gustafsson, M. G. L., Agard, D. A. & Sedat, J. W. Phase-retrieved pupil functions in wide-field fluorescence microscopy. J. Microsc. 216, 32–48 (2004).
Prabhat, P., Ram, S., Ward, E. S. & Ober, R. J. Simultaneous imaging of different focal planes in fluorescence microscopy for the study of cellular dynamics in three dimensions. IEEE Trans. Nanobiosci. 3, 237–242 (2004).
Huang, F. et al. Video-rate nanoscopy using sCMOS camera-specific single-molecule localization algorithms. Nat. Methods 10, 653–658 (2013).
Mlodzianoski, M. J. et al. Active PSF shaping and adaptive optics enable volumetric localization microscopy through brain sections. Nat. Methods 15, 583–586 (2018).
Jungmann, R. et al. Multiplexed 3D cellular super-resolution imaging with DNA-PAINT and Exchange-PAINT. Nat. Methods 11, 313–318 (2014).
Oakley, H. et al. Intraneuronal β-amyloid aggregates, neurodegeneration, and neuron loss in transgenic mice with five familial Alzheimer’s disease mutations: potential factors in amyloid plaque formation. J. Neurosci. 26, 10129–10140 (2006).
Jay, T. R. et al. Disease progression-dependent effects of TREM2 deficiency in a mouse model of Alzheimer’s disease. J. Neurosci. 37, 637–647 (2017).
Hruska, M., Henderson, N., Marchand, S. J. L., Jafri, H. & Dalva, M. B. Synaptic nanomodules underlie the organization and plasticity of spine synapses. Nat. Neurosci. 21, 671–682 (2018).
Madisen, L. et al. A toolbox of Cre-dependent optogenetic transgenic mice for light-induced activation and silencing. Nat. Neurosci. 15, 793–802 (2012).
Pasquali-Ronchetti, I. & Baccarani-Contri, M. Elastic fiber during development and aging. Microsc. Res. Tech. 38, 428–435 (1997).
Burke, D., Patton, B., Huang, F., Bewersdorf, J. & Booth, M. J. Adaptive optics correction of specimen-induced aberrations in single-molecule switching microscopy. Optica 2, 177–185 (2015).
Ji, N. Adaptive optical fluorescence microscopy. Nat. Methods 14, 374–380 (2017).
Power, R. M. & Huisken, J. A guide to light-sheet fluorescence microscopy for multiscale imaging. Nat. Methods 14, 360–373 (2017).
Legant, W. R. et al. High-density three-dimensional localization microscopy across large volumes. Nat. Methods 13, 359–365 (2016).
Gradinaru, V., Treweek, J., Overton, K. & Deisseroth, K. Hydrogel-tissue chemistry: principles and applications. Annu. Rev. Biophys. 47, 355–376 (2018).
Chen, F., Tillberg, P. W. & Boyden, E. S. Expansion microscopy. Science 347, 543–548 (2015).
Douglass, K. M., Sieben, C., Archetti, A., Lambert, A. & Manley, S. Super-resolution imaging of multiple cells by optimized flat-field epi-illumination. Nat. Photonics 10, 705–708 (2016).
Stehr, F., Stein, J., Schueder, F., Schwille, P. & Jungmann, R. Flat-top TIRF illumination boosts DNA-PAINT imaging and quantification. Nat. Commun. 10, 1268 (2019).
Tokunaga, M., Imamoto, N. & Sakata-Sogawa, K. Highly inclined thin illumination enables clear single-molecule imaging in cells. Nat. Methods 5, 159–161 (2008).
McGorty, R., Kamiyama, D. & Huang, B. Active microscope stabilization in three dimensions using image correlation. Opt. Nanoscopy 2, 3 (2013).
Acknowledgements
We would like to thank K. Dhanasekaran and P. Lusk (Yale University) for sharing the labeling protocol of Nup98 and interpretation of the resolved Nup98 structures. We thank M. J. Mlodzianoski for initial instrument design, S. An for help in instrument alignment and sample preparation and D. A. Miller for providing labeling protocols of mitochondria and microtubules. F.X., D.M., S.L., C.B. and F.H. were supported by grants from the NIH (R35GM119785) and DARPA (D16AP00093). K.P.M. and G.E.L were supported by grants from the NIH (RF1AG051495 and RF1AG050597). Y.B. and S.C. were supported by a grant from the NIH (R01AR071359). Y.W. and P.Y. were supported by a grant from the NIH (1R01EB018659) and Harvard Medical School Dean’s Initiative Grant.
Author information
Authors and Affiliations
Contributions
F.X., D.M. and F.H. conceived the project. F.X., S.L. and F.H. developed the algorithm. F.X. wrote the software and performed simulation. D.M. developed the microscope setup and performed super-resolution experiments. F.X., D.M. and S.L. visualized experimental data. D.M., K.P.M., Y.B., Y.W., Y.T., C.B., T.K., A.A.C., P.Y., S.C., G.E.L. and F.H. designed the experiments. D.M., K.P.M., Y.B., Y.W., Y.T., C.B. and T.K. prepared biological samples. A.A.C., P.Y., S.C., G.E.L. and F.H. supervised the study. All authors wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
F.X., D.M. and F.H. are inventors on patent application submitted by Purdue University that covers basic principles of INSPR.
Additional information
Peer review information Rita Strack was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 INSPR framework, degeneracy illustration, and setup diagram.
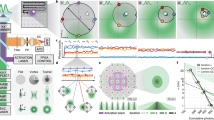
a, INSPR framework and detailed process of in situ model generation. b, Single molecules are localized by a pair of channel-specific models which share the same shape information with the corresponding sub-regions. c, Degeneracy exists in single plane configuration, where PSFs with positive and negative vertical astigmatism aberrations (Ast) are identical at opposite axial positions. d, Degeneracy is broken in biplane configuration, where PSF pairs with positive and negative vertical astigmatism aberrations are different at opposite axial positions. e, Degeneracy is absent in single plane configuration with prior knowledge of astigmatism orientation, where PSFs with additional positive and negative primary spherical aberrations (1st Sph) are different at opposite axial positions. Scale bar in (c–e): 1 μm. f, Setup diagram. M1–M8: mirrors in the excitation path; Di1–Di3: dichroic mirrors; AOTF: acousto-optic tunable filter; L1–L5: lenses in the excitation path; FM: flip mirror; MLS: mercury light source; Obj: objective lens; M1’–M11’: mirrors in the emission path; TL: tube lens; L1’–L6’: lenses in the emission path; DM: deformable mirror; BS: 50/50 non-polarizing beam splitter; SM: 90° specialty mirror; FW: filter wheel. Nominal focal lengths of lenses are, L1: 19 mm, L2: 19 mm, L3: 20 mm, L4: 125 mm, L5: 400 mm, Obj: 1.8 mm, TL: 180 mm, L1’: 88.9 mm, L2’: 250 mm, L3’: 400 mm, L4’: 150 mm, L5’: 500 mm, L6’: 250 mm. g, Definition of biplane distance. The objective lens is moved axially to make plane 1 (case 1) and plane 2 (case 2) in focus successively. The axial movement is defined as biplane distance δ.
Extended Data Fig. 2 Performance quantification of INSPR in biplane setup.
a, Similarity between the ground truth 3D PSFs and the 3D PSFs at different imaging depths when using INSPR (blue circles), Gaussian model (orange stars), and theoretical index mismatch model (IMM, yellow squares). For each depth, 3D normalized cross correlation (NCC) coefficients between the ground truth PSFs and the PSFs generated using three methods at different axial offsets are shown, with the maximum values marked (purple diamonds). b, 3D PSFs retrieved using Gaussian, IMM, and INSPR in comparison to the ground truth (GT) at different depths, when NCC reaches the maximum at each depth (purple diamonds in (a)). The defocus offset (that is, the axial shift from the actual focal plane) is obtained by finding the maximum-intensity plane of the ground truth PSFs along the axial direction. Scale bar: 1 μm. c, Root-mean-square error (RMSE) between the decomposed Zernike amplitudes of INSPR retrieved model and the ground truth amplitudes in different photon (I) and background (bg) conditions. In each condition, the amplitudes of the ground truth are randomly sampled from –1 to +1 (unit, λ/2π) for each trial (11 trials in total). d, Heat map showing the relationship between the input and phase retrieved amplitudes of 21 Zernike modes. e, Scatter plots of lateral localizations using model transformation (top) and data transformation (bottom) for PSFs with vertical astigmatism (Ast). The total photon count per emission event I is 2000, and the background count per pixel bg is 30. Plane 1 and plane 2 are related with an affine transformation including a rotation of 30 degrees. Both Poisson noise and pixel-dependent sCMOS readout noise (the variance distribution is shown in the inset) are considered. f, Localization precisions and biases in the x, y, and z dimensions for the dataset in (e).
Extended Data Fig. 3 Blind reconstruction of 3D training datasets of microtubules (MT0.N1.LD) from the SMLM challenge.
a,b, x-y and x-z overviews of the microtubules resolved by INSPR from the 3D-Biplane data. c,d, Enlarged x-y and x-z views of the areas as indicated by the magenta and blue boxed regions in (a) and (b), respectively. e,f, Intensity profiles along the y and z directions within the white boxed regions in (c,d), comparing the INSPR resolved profiles (blue solid lines) with the ground truth (red dashed lines). g, x-y views of the provided calibration PSF (3D-Biplane, top rows) and the INSPR retrieved PSF from blinking data (bottom rows). h,i, x-y and x-z overviews of the microtubules resolved by INSPR from the 3D-Astigmatism data. j,k, Enlarged x-y and x-z views of the areas as indicated by the magenta and blue boxed regions in (h) and (i), respectively. l,m, Intensity profiles along the y and z directions within the white boxed regions in (j,k), comparing the INSPR resolved profiles (blue solid lines) with the ground truth (red dashed lines). n, x-y views of the provided calibration PSF (3D-Astigmatism, top rows) and the INSPR retrieved PSF from blinking data (bottom rows). Scale bar in (g,n): 1 µm. Norm.: normalized.
Extended Data Fig. 4 3D super-resolution reconstructions of immunofluorescence-labeled TOM20 in COS-7 cells using INSPR, ZOLA-3D, and cubic spline in astigmatism-based setup.
a, x-y overview of the mitochondrial network resolved by INSPR, with a depth of 13 μm from the coverslip. b–d, x-z slices along the white dashed line in (a), reconstructed using INSPR (b), ZOLA-3D which considers PSF distortions inside the refractive index mismatched medium (c), and cubic spline from beads on the coverslip (d). The white arrows and yellow boxes highlight the differences in axial reconstructions among three methods. e–g, x-z slices along the magenta dashed line in (a), reconstructed using INSPR (e), ZOLA-3D (f), and cubic spline (g). The white arrows and orange boxes highlight the differences in axial reconstructions among three methods. h–j, Intensity profiles along the yellow dashed lines in (b–d), showing the difference in the axial width of the outer membrane contour is 10% for both ZOLA-3D and cubic spline as compared to INSPR. k–m, Intensity profiles along the orange dashed lines in (e–g), showing the differences in the axial width of the outer membrane contour are 13% and 16% for ZOLA-3D and cubic spline as compared to INSPR, respectively. The integration width of the x-z slices in (b–g) in the y direction is 200 nm. The dataset shown is representative of four datasets of mitochondria with depths of ~13 μm from the coverslip. Norm.: normalized.
Extended Data Fig. 5 3D super-resolution reconstructions of immunofluorescence-labeled TOM20 in COS-7 cells using INSPR and microsphere-calibrated Gaussian fitting in astigmatism-based setup.
a, x-y overview of the mitochondrial network resolved by INSPR on the bottom coverslip, within the expected working range of microsphere-calibrated Gaussian fitting. b–e, y-z slices along the white and magenta dashed lines in (a), reconstructed using INSPR (b,d) and microsphere-calibrated Gaussian fitting (c,e). f, Calibration curves showing σx and σy observed (solid lines) and fitted (dashed lines) obtained from the blinking data of microspheres as a function of the depth from the bottom coverslip. The crossover point of σx and σy is at the depth of 0.5 μm. g, x-y overview of the mitochondrial network resolved by INSPR with a depth of 1.5 μm from the bottom coverslip, outside the working range of microsphere-calibrated Gaussian fitting. h–k, x-z slices along the white and magenta dashed lines in (g), reconstructed using INSPR (h,j) and microsphere-calibrated Gaussian fitting (i,k). l, Calibration curves showing σx and σy observed (solid lines) and fitted (dashed lines) obtained from the blinking data of microspheres as a function of the depth from the bottom coverslip. The crossover point of σx and σy is at the depth of 2.2 μm. The integration width of the slices in (b–e, h–k) in the third dimension is 200 nm. The datasets shown are representative of four datasets of mitochondria on the coverslip and four datasets of mitochondria with depths of ~1.5 μm from the coverslip. Obs.: observed. Fit.: fitted.
Extended Data Fig. 6 3D super-resolution reconstructions of immunofluorescence-labeled TOM20 in COS-7 cells using INSPR and the in vitro method in biplane setup.
a, x-y overview of the mitochondrial network showing the positions of 25 typical outer membrane contours as indicated by the magenta and white boxed regions. b, Enlarged y’-z views of the outer membrane structures as indicated by the white boxed regions in (a), showing the reconstructed images using INSPR (left) and phase retrieval based on beads on the coverslip (in vitro (PR), right). Here the orientation of the cross section is rotated to allow projection of the 3D membrane bounded structures to the 2D image. c, x-y and x-z views of the PSFs retrieved by INSPR in different optical sections and those retrieved by in vitro PR, as well as the phase and magnitude of the corresponding pupils. Scale bar: 1 µm. d, Amplitudes of 21 Zernike modes (Wyant order, from vertical astigmatism to tertiary spherical aberration) decomposed from the pupils retrieved by INSPR and in vitro PR. e, Distribution of σy’ obtained from the intensity profiles of 25 typical outer membranes in (a) reconstructed using INSPR (blue plus signs) and in vitro PR (red circles). Sec.: optical section.
Extended Data Fig. 7 3D super-resolution reconstructions of immunofluorescence-labeled Nup98 in COS-7 cells using INSPR and the in vitro method in biplane setup.
a, x-y overview of the 3.3-µm-thick volume of the nucleus showing the positions of 40 typical Nup98 structures (yellow lines). b, x-z slice along the white dashed line in (a), showing the positions of 20 typical Nup98 structures (cyan lines). c, x-y overview of the 6.4-µm-thick entire nuclear envelope showing the positions of 40 typical Nup98 structures (yellow lines). d, x-z slice along the white dashed line in (c), showing the positions of 10 typical Nup98 structures on the top (green lines) and bottom (red lines) surfaces. e, Distribution of diameters measured from Nup98 structures in the x-y plane shown in (a,c). The diameter is 60 ± 9 nm for the 3.3-µm-thick volume (mean ± s.d., 40 measurements, red crosses), and 57 ± 11 nm for the 6.4-µm-thick volume (40 measurements, blue plus signs). f, Distribution of σy measured from Nup98 structures in the x-y plane shown in (a,c). σy is 14 ± 3 nm for the 3.3-µm-thick volume (80 measurements, black circles), and 11 ± 3 nm for the 6.4-µm-thick volume (80 measurements, magenta squares). g, Distribution of σz measured from Nup98 structures in the x-z plane shown in (b,d). For the 3.3-µm-thick volume, σz is 48 ± 9 nm (20 measurements, cyan diamonds). For the 6.4-µm-thick volume, σz is 46 ± 12 nm for the top surface (10 measurements, green upward-pointing triangles), and 36 ± 11 nm for the bottom surface (10 measurements, red downward-pointing triangles). h,i, x-z slice along the white dashed line in (c), reconstructed using INSPR (h) and in vitro phase retrieval based on beads on the coverslip (in vitro (PR), i). The integration width of the x-z slices in (b,d,h,i) in the y direction is 500 nm.
Extended Data Fig. 8 3D super-resolution reconstructions of immunofluorescence-labeled ChR2-EYFP on dendrites in 50-μm-thick brain slices using INSPR and in vitro methods in biplane setup (depth: 2 – 6.2 μm).
a, x-y overview of the super-resolution volume of immunofluorescence-labeled ChR2-EYFP on dendrites resolved by INSPR, with a depth of 2 μm from the coverslip. b–d, x-z slices along the white dashed line in (a), reconstructed using INSPR (b), phase retrieval method based on beads on the coverslip with theoretical index mismatch model (PR + IMM, c), and phase retrieval method based on beads on the coverslip (PR, d). e–j, Zoomed in x-z views of the areas as indicated by the white boxed regions in (b–d). k–m, x-z slices along the magenta dashed line in (a), reconstructed using INSPR (k), PR + IMM (l), and PR (m). n–p, Zoomed in x-z views of the areas as indicated by the white boxed regions in (k–m). q–s, Intensity profiles along the white dashed lines in (n–p), showing the differences in the axial width of the selected contour are 41% and 26% for PR + IMM and PR as compared to INSPR, respectively. The integration width of the x-z slices in (b–p) in the y direction is 200 nm. Norm.: normalized.
Extended Data Fig. 9 3D super-resolution reconstructions of immunofluorescence-labeled ChR2-EYFP on dendrites in 50-μm-thick brain slices using INSPR and phase retrieval method based on beads embedded in agarose gel in biplane setup.
a, x-y overview of the super-resolution volume of immunofluorescence-labeled ChR2-EYFP on dendrites resolved by INSPR, with a depth of 11 μm from the coverslip. b–e, x-z slices along the white and magenta dashed lines in (a), reconstructed using INSPR (b,d) and phase retrieval method based on beads embedded in agarose gel (PR in gel, c,e). f, Intensity profiles along the white dashed lines in (d,e), showing the difference between reconstructions using INSPR (red dashed lines) and PR in gel (blue solid lines). g, x-y overview of the super-resolution volume of immunofluorescence-labeled ChR2-EYFP on dendrites resolved by INSPR, with a depth of 7 μm from the coverslip. h–k, x-z slices along the white and magenta dashed lines in g, reconstructed using INSPR (h,j) and PR in gel (i,k). l, Intensity profiles along the white dashed lines in (j,k), showing the difference between reconstructions using INSPR (red dashed lines) and PR in gel (blue solid lines). The integration width of the x-z slices in (b–e, h–k) in the y direction is 200 nm. These experiments are performed twice and both of them are shown here. Norm.: normalized.
Extended Data Fig. 10 3D super-resolution reconstructions of immunofluorescence-labeled elastic fibers in developing cartilage in 20-μm-thick tissue slices using INSPR and in vitro methods in biplane setup.
a, x-y overview of the reconstructed volume of immunofluorescence-labeled elastic fibers in developing cartilage using INSPR. b, Diffraction-limited image of (a). c–k, x-z slices along the white, magenta, and yellow dashed lines in (a), reconstructed using INSPR (c,f,i), phase retrieval method based on beads on the coverslip with theoretical index mismatch model (PR + IMM, d,g,j), and phase retrieval method based on beads on the coverslip (PR, e,h,k). The integration width of the x-z slices in the y direction is 2 μm.
Supplementary information
Supplementary Information
Supplementary Figures 1–15, Supplementary Tables 1–3 and Supplementary Notes 1–3
Supplementary Video 1
Performance demonstration of INSPR on simulated single-molecule emission patterns in biplane setup.
Supplementary Video 2
Performance demonstration of INSPR on simulated single-molecule emission patterns in astigmatism-based setup.
Supplementary Video 3
INSPR estimates wavefront distortions induced by the deformable mirror from single-molecule datasets of TOM20 immune-labeled with DNA-PAINT in fixed COS-7 cells.
Supplementary Video 4
INSPR reconstruction of immunofluorescence-labeled TOM20 in fixed COS-7 cells.
Supplementary Video 5
INSPR reconstruction of immunofluorescence-labeled Nup98 on an entire nucleus in fixed COS-7 cells.
Supplementary Video 6
INSPR reconstruction of immunofluorescence-labeled amyloid-β plaque with low-density fibrils in a 30-μm-thick brain section from a 5XFAD mouse.
Supplementary Video 7
INSPR reconstruction of immunofluorescence-labeled amyloid-β plaque with high-density fibrils in a 30-μm-thick brain section from a 5XFAD mouse.
Supplementary Video 8
INSPR reconstruction of immunofluorescence-labeled ChR2–EYFP on dendrites in a 50-μm-thick brain section from mouse primary visual cortex.
Supplementary Video 9
INSPR reconstruction of immunofluorescence-labeled elastic fibers in developing cartilage from embryonic mouse forelimbs.
Supplementary Software 1
INSPR software package and user manual for biplane setup.
Supplementary Software 2
INSPR software package and user manual for astigmatism-based setup.
Rights and permissions
About this article
Cite this article
Xu, F., Ma, D., MacPherson, K.P. et al. Three-dimensional nanoscopy of whole cells and tissues with in situ point spread function retrieval. Nat Methods 17, 531–540 (2020). https://doi.org/10.1038/s41592-020-0816-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-0816-x
This article is cited by
-
Temporal analysis of relative distances (TARDIS) is a robust, parameter-free alternative to single-particle tracking
Nature Methods (2024)
-
Universal inverse modeling of point spread functions for SMLM localization and microscope characterization
Nature Methods (2024)
-
Label-free adaptive optics single-molecule localization microscopy for whole zebrafish
Nature Communications (2023)
-
Deep learning-driven adaptive optics for single-molecule localization microscopy
Nature Methods (2023)
-
Field-dependent deep learning enables high-throughput whole-cell 3D super-resolution imaging
Nature Methods (2023)