Abstract
Although B cells are implicated in multiple sclerosis (MS) pathophysiology, a predictive or diagnostic autoantibody remains elusive. In this study, the Department of Defense Serum Repository (DoDSR), a cohort of over 10 million individuals, was used to generate whole-proteome autoantibody profiles of hundreds of patients with MS (PwMS) years before and subsequently after MS onset. This analysis defines a unique cluster in approximately 10% of PwMS who share an autoantibody signature against a common motif that has similarity with many human pathogens. These patients exhibit antibody reactivity years before developing MS symptoms and have higher levels of serum neurofilament light (sNfL) compared to other PwMS. Furthermore, this profile is preserved over time, providing molecular evidence for an immunologically active preclinical period years before clinical onset. This autoantibody reactivity was validated in samples from a separate incident MS cohort in both cerebrospinal fluid and serum, where it is highly specific for patients eventually diagnosed with MS. This signature is a starting point for further immunological characterization of this MS patient subset and may be clinically useful as an antigen-specific biomarker for high-risk patients with clinically or radiologically isolated neuroinflammatory syndromes.
Similar content being viewed by others
Main
Multiple sclerosis (MS) is a chronic inflammatory autoimmune disease that primarily affects the white matter of the central nervous system (CNS)1,2. Although traditionally thought to be T cell mediated3,4, the widespread success of B-cell-depleting therapies in humans has focused attention on a central role of B cells in the etiology and progression of MS5,6. MS is often disabling when untreated; it affects women more often than men; and it appears to have increased in frequency, with nearly 1 million individuals currently affected in the United States alone7,8.
In the large majority of patients with MS (PwMS), MS presents as a bout or relapse: a neurological symptom complex suggestive of demyelination, such as optic neuritis, partial myelitis, brainstem syndrome or multifocal onset1,2,4. Magnetic resonance imaging (MRI) of the CNS performed shortly after clinical onset often shows acute demyelinating plaques that enhance after intravenous contrast administration9. In addition, there are often one or more chronic demyelinating lesions9, suggesting that neuroinflammation precedes symptomatic onset4. The recognition of the ‘radiologically isolated syndrome’ (RIS) demonstrated that plaques could be identified on brain and spinal cord MRI before symptom onset when these scans were performed for other reasons10. Histopathologically, demyelinating plaques are heterogeneous and are populated by complex and varying immune cell populations, including CD4+ and CD8+ T cells, activated macrophages and B cells, and deposition of immunoglobulin and complement3,4.
Several retrospective studies that assessed clinical records found that non-specific neurologic episodes occur more frequently in individuals who received an MS diagnosis later in life, suggesting that MS prodrome involves an ongoing inflammatory process11,12. Given that MS lesions are thought to develop after demyelinating events, it may be that these neurological episodes are indicative of ongoing neuroinflammatory process in the preclinical period13. Although molecular evidence for this has been limited, the theory that neuronal tissue injury precedes clinical onset is supported by recent studies showing elevated serum neurofilament light (sNfL), indicating axonal damage several years before MS diagnosis14,15,16.
Studies in several autoimmune diseases—systemic lupus erythematosus17, type 1 diabetes mellitus18 and rheumatoid arthritis19, among others—indicate that diagnostic autoantibodies can appear years before the onset of symptoms20. In MS, by contrast, no such validated diagnostic autoantibodies exist21. Indeed, the role of autoantibodies in MS5,22, and their link to pathogenesis23,24, has proven notoriously difficult to ascertain, with studies suggesting myelin or potassium channel (KIR4.1) as putative antigenic targets22,25,26. A hallmark of MS is the presence of unique oligoclonal bands in the cerebrospinal fluid (CSF) of almost all PwMS13,24,27, which implies intrathecal antibody synthesis28. However, no clearly predictive or diagnostic autoantigen has been identified21. The need for prospectively ascertained cohorts of sufficient size to study this heterogeneous disease makes prospective autoantigen identification exceptionally challenging29,30.
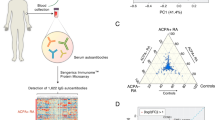
To assess autoantibody signatures, we used a large, prospective, incident MS cohort assembled during the Gulf War era (GWEMSC) from over 10 million individuals in the active duty United States military population31. From this group of cases, the Department of Defense Serum Repository (DoDSR) staff retrieved the earliest serum aliquot from entry into active duty, an average of 5 years before their first clinical symptom (Fig. 1a,b), and then another serum sample on average 1 year after this first attack. Control samples, without an MS diagnosis, were selected and matched to cases on age, sex, race/ethnicity and dates of serum collection (year). After a whole-proteome autoantibody screen and sNfL measurements on these samples (Fig. 1c,d), autoantibody results were orthogonally validated on both serum and CSF using a gold standard, prospective, incident MS cohort at the University of California, San Francisco (ORIGINS) that enrolled patients at clinical onset.
a, Schematic of DoDSR cohort and collection. b, Age and time to symptom onset for MS cases (data are presented as median values; box edges are 1st and 3rd quartiles; and whiskers represent 1.5× interquartile range; n = 250 for each group). c,d, Molecular biomarker assays performed on DoDSR cohort of longitudinal sera. NCBI, National Center for Biotechnology Information.
Results
DoDSR and ORIGINS MS cohorts
The DoDSR cohort contained 250 PwMS 5 years before (5.0 \(\pm\), s.d. 3.3) and 1 year after (1.2 \(\pm\), s.d. 0.4) first symptom onset and 250 controls who were matched for age, sex, race/ethnicity and serum collection dates (Extended Data Table 1). The UCSF ORIGINS cohort, comprising untreated patients presenting with their first-ever attack, consisted of 103 patients who were ultimately diagnosed with relapsing-remitting multiple sclerosis (RRMS) (Extended Data Table 2). Other neurologic disease (OND) controls included 14 patients who enrolled in ORIGINS but were ultimately found to have a non-MS diagnosis (varicella zoster virus meningitis n = 1, neuromyelitis optica spectrum disorder (NMOSD) n = 4, vitamin B12 deficiency n = 2, Leber’s hereditary optic neuropathy n = 2, neurosarcoidosis n = 2, primary CNS vasculitis n = 1 and lymphoma n = 1) as well as nine additional OND controls (migraine n = 4, brainstem stroke n = 1, anti-phospholipid antibody syndrome n = 1, brainstem demyelinating syndrome n = 1, idiopathic spastic paraparesis n = 1 and peripheral neuropathy n = 1).
Immunologic signature present early in DODSR cohort
Molecular profiling of both autoantibody repertoire and neuronal damage were carried out on the longitudinal samples acquired from the DoDSR (Fig. 2). sNfL, a marker of axonal injury, was measured in cases and controls at both timepoints. Among the preclinical serum samples, NfL levels were higher at timepoints closer to date of diagnosis (Fig. 2b). Consistent with a recent study14, sNfL levels were significantly higher in post-onset samples compared to their pre-onset counterparts for PwMS (Fig. 2c).
a, UMAP of autoantigen enrichments in PhIP-Seq screen of DoDSR sera, showing distinct immunogenic clusters (IC1 and IC2). sNfL levels across DoDSR cohort sera with respect to time before onset (b) and grouped by timepoint for cases and controls (c) (data represented as geometric means and standard factors; P values shown above; evaluated via t-test comparing log-transformed NfL values; first timepoint: n = 236 for HC and n = 217 for MS; second timepoint: n = 234 for HC and n = 212 for MS). Sum of normalized reads (d) and individual fold enrichments (e) of top 192 peptides, with IC patientsʼ pre-onset and post-onset samples highlighted on left grouped by hierarchical clustering. f, Blowout of top 26 most enriched peptides in 27 IC patients grouped by gene and respective enrichments in their non-MS matched controls.
In patients ultimately diagnosed with MS, sNfL levels were higher even many years before their first clinical flare when compared to the matched control cohort (Fig. 2c). Significant differences in sNfL levels across timepoints were not observed in the control cohort. Together, these data provide evidence that at least some PwMS exhibit early signs of neuroaxonal injury long before onset of symptoms.
A whole-human proteome seroreactivity approach called Phage Immunoprecipitation Sequencing (PhIP-Seq)32 was employed to determine if an antigen-specific signature of future disease accompanied elevated sNfL levels. This technique uses a T7 phage display library to probe antibody–antigen interactions by immunoprecipitation from patient serum or CSF and has been used for antibody-based biomarker detection in diseases of unknown etiology32,33,34,35,36,37,38,39.
The longitudinal nature of this cohort with its well-matched control samples allows for detection of changes in the antigenic repertoire before and after clinical onset. The collection of peptides enriched by immunoprecipitation using sera from patients (the autoantibody signature) in the cohort was consistent over time (Extended Data Fig. 1). When blinded, timepoints from the same patient could be unambiguously associated with nearly 96% accuracy across the 500 individuals assayed (Extended Data Fig. 2). This consistency over time was present regardless of diagnosis. No compelling enrichment of antigens specific for PwMS after diagnosis was observed.
We then asked whether there might be a serologic signature in patients who went on to develop MS compared to those who did not. To visualize the differences between the overall PhIP-Seq results across all samples, a uniform manifold approximation and projection (UMAP) was generated using the fold change (FC) enrichment for each peptide relative to the control cohort (Methods). In this projection, a distinct immunogenic cluster (IC) emerged that included PwMS before and after disease onset (Fig. 2a). We specifically looked for previously identified putative targets of molecular mimicry, such as GlialCAM40 and anoctamin-2 (ref. 41), and identified them in rare instances (Extended Data Fig. 3).
After unsupervised clustering of this cohort of samples, serum antibodies from this group of patients enriched several of the same peptides that were preferentially found in patients who developed MS (n = 27) as opposed to controls without MS (n = 3). These cases separated into two clusters: IC1, with a more polyspecific profile, and IC2, which appeared to be reactive to a subset of IC1 peptides (Fig. 2d). These patients were grouped into an MSIC category for the purpose of further analysis compared to those who did not exhibit reactivity (MSno-IC).
Patients in these clusters were defined by a clear but disparate group of enriched peptides derived from 54 different proteins in both pre-onset and post-onset samples (Methods). Most striking about this result was its longitudinal stability (Fig. 2e). Analysis of all IC samples, both before and after disease onset, clearly shows that this class of antigens is enriched in both timepoints in the majority (17/26, 65%) of cases (one case did not meet sequencing depth cutoff in respective pre-onset sample).
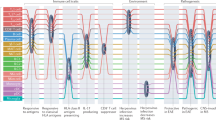
Alignment of peptides preferentially enriched by sera from the IC subset of individuals revealed a characteristic protein motif (‘IC motif’) described by the regular expression P-(SA)-x-(SGA)-R-(SN)-(LRKH), with the initial proline being the most conserved (Fig. 3a). This motif is broadly represented in the human proteome (Fig. 3a). Although the motif-bearing peptides derive from diverse coding sequences, proteins containing splicing activation and mRNA-binding domains, due to their characteristic arginine–serine repeat sequences42,43,44—that is, SRSF4, SRRM3 and CLASRP—were highly represented in the enriched set.
a, Protein alignment of top 45 peptides within human proteome library that contain the IC regular expression, in order of patient prevalence that exceeded cutoffs in either pre-onset or post-onset samples. Bottom is alignment of regular expression to selected pathogens that infect humans (via PROSITE scan). b, Normalized reads assigned to regex-containing peptides in PwMS within either IC cluster as compared to those without and non-MS controls (data represented as means and s.d.; one-way ANOVA with Tukey’s multiple comparisons adjustment; P values shown above; n = 490 for HC, n = 433 for MSno-IC and n = 44 for MSIC). c, Serum levels of NfL in patients with MS within IC clusters as compared to those without and controls (data represented as geometric means and standard factors; P values shown above; evaluated via t-test with Bonferroni correction comparing log-transformed NfL values; n = 470 for HC, n = 395 for MSno-IC and n = 34 for MSIC). d, Serum NfL levels modeled over time for each group with respect to time to onset.
Searching the PROSITE database45 using the IC motif regular expression and not simply a most-probable collapsed sequence disclosed the motif’s presence across many phyla. In addition to humans, the IC motif is present in several human pathogens, including Epstein–Barr virus (EBV) (both in the BRRF2 tegument protein and the envelope glycoprotein M), hepatitis C virus (NS3 protease), Pseudomonas aeruginosa (NosR), Escherichia coli (HTH-type transcriptional repressor) and Aspergillus fumigatus (RNA-binding protein vts1) (Fig. 3a).
Neither patient demographic nor clinical features (for example, time to symptom onset, MS subtype and gender) distinguished these IC reactive patients from the larger cohort (Extended Data Table 3). Nevertheless, patients with enriched signal across peptides described by the IC motif had significantly higher sNfL levels than PwMS not assigned to this cluster as well as the controls (Fig. 3b–d). The difference in sNfL was preserved at both pre and post timepoints (Extended Data Fig. 4).
IC signature present in both CSF and serum in the ORIGINS cohort
Although the longitudinal stability of this signature is important in the context of surveillance, we sought to replicate this finding in patients with incident MS or clinically isolated syndrome (high risk for MS) within 30 d of their first symptomatic episode, where its diagnostic value would be more apparent. Here, 126 paired CSF and serum samples were analyzed from a completely independent incident MS cohort of treatment-naive patients (UCSF ORIGINS study) of which most (n = 104, 83%) were eventually diagnosed with MS after radiologic and clinical follow-up. A similar fraction of patients with confirmed MS (8/104) expressed the same signature as those of IC reactive patients in the DoDSR longitudinal cohort, with characteristic enrichment of peptides containing the IC motif (Fig. 4a,b). These patients similarly split into IC1 and IC2 type profiles, clustered together via UMAP (Fig. 4d) and demonstrated enrichment of these peptides in both compartments (Fig. 4c). Interestingly, this signature was highly specific for MS cases, where only one patient was diagnosed with NMOSD, having low-level enrichment in their serum but not in CSF, out of 22 confirmed OND controls. Additional OND CSF controls (n = 20) without matching serum also lacked this signature (Extended Data Fig. 5).
Sum of normalized reads for previously identified top 26 peptides (a) and PhIP-Seq fold enrichments of ORIGINS patient sera and CSF for peptides mapping to same regions of ICs from the DoDSR cohort, relative to controls (b). c, Blowup of IC signature patients with all OND controls with CSF and serum for each patient grouped by column. d, UMAP of PhIP-Seq enrichments for all ORIGINS patients, shaded by sum RPK levels from a. Arrows indicate where IC clusters lie on UMAP.
This result was orthogonally validated using a bead-based (Luminex) multiplexed indirect immunofluorescence assay featuring six distinct peptides that were highly enriched across the MSIC patient samples (Fig. 5a). Relevant peptides from proteins RIMS2, KRT75, SRSF4, CLASRP, SRRM3 and TRIO were included, each of which contained variations of the IC motif (Table 1). After testing all ORIGINS paired CSF and sera using this assay, only patients exceeding the cutoff were ultimately diagnosed with MS (Fig. 5b–e). Similar to the PhIP-Seq results, some degree of heterogeneity was observed across the peptide combinations, but samples from the same group of patients yielded enrichment of the IC motif (Fig. 5b–e and Extended Data Fig. 6). A metric of the summed mean fluorescence intensity (MFI) (Fig. 5f) across these peptides confirmed and extended the PhIP-Seq findings. Specifically, seroreactivity to peptides bearing the IC motif was highly specific for PwMS at the time of symptom onset, were not found in OND controls and were nearly always present in both CSF and serum in each case.
a, Schematic for barcoded immunofluorescence assay against top peptides from IC motif. Normalized median fluorescence intensity for each patient, matched by colored line, across all ORIGINS patientsʼ CSF (b) and sera (c) (n = 104) as well as OND controls (n = 22 serum and n = 42 CSF) (d,e). f, Same patient population shown with sum of MFI across peptides shown against all four groups, with those exceeding cutoff (normalized logMFI > 3) in CSF highlighted and matched by patient.
We also investigated whether peptides from sequence-aligned pathogens (as seen in Fig. 3a) had cross-reactivity in patient sera and CSF in PwMS with and without this signature (Table 1). We saw that the IC1 patients had the highest cross-reactivity across the cohort to two key peptides from EBV BRRF2 and HFM NS2PC (Extended Data Fig. 7).
To evaluate whether human leukocyte antigen (HLA) variation may contribute to the unique phenotype observed in MSIC patients, the classical HLA genes HLA-A, HLA-B, HLA-C, HLA-DPB1, HLA-DRB1 and HLA-DQB1 were fully sequenced and genotyped (Supplementary Table 1). Due to limited sample size, formal analysis to determine whether any HLA locus or allele exhibited a significant association with MSIC status was not possible. Qualitative assessment of allele frequencies shows that some HLA alleles were found at higher frequencies in the MSIC group relative to the remaining ORIGINS MS cohort; however, it must be noted that many alleles in the MSIC group were present as only single copies (Supplementary Table 2). Interestingly, all but one of the MSIC individuals carry the HLA-DPB1*04:01 allele (Supplementary Table 1). This high carrier frequency of 83%, relative to the 67% observed in the remaining cohort, suggests a possible enrichment for the HLA-DPB1*04:01 allele in the MSIC subset. However, these observations must be considered strictly speculative, as a much larger sample will be required to determine whether any observed differences in HLA frequency distributions between these groups is significant.
Discussion
In this study, an MS-specific autoantibody signature was identified in a subset of patients years before their first clinical MS attack. This same signature was validated using two different methods in an independent, prospective cohort of patient samples (serum and CSF) collected near the time of their first MS relapse (UCSF ORIGINS cohort).
This study presents some of the first prospective autoantigen-specific biomarkers found in the preclinical phase of MS, which is consistent with prior clinical analyses that MS pathophysiology may begin years before symptom onset and diagnosis. The evidence includes numerous studies that thoroughly investigated clinical and indirect markers of this phase via chart review of healthcare claims11,29,46. Documented symptoms range from fatigue and headache to mental health dysfunction, including anxiety and depression, and prescriptions for antispasmodics, all of which have elevated odds ratios in patients who went on to develop MS. Evidence from a similar prospective military cohort found that poor cognitive performance was found at a higher rate in patients who went on to receive an MS diagnosis in the following 2 years compared to controls47.
This work validates and adds to prior evidence of neuro-axonal injury occurring in patients during the MS preclinical phase14, as sNfL is a robust biomarker that is likely specific to neuro-axonal damage and is elevated in the serum of PwMS compared to healthy controls (HCs)16,48,49,50,51. Elevated levels are associated with faster progression to Expanded Disability Status Scale (EDSS) scores higher than 4, and reduction in sNfL concentration is associated with initiation of disease-modifying therapy52. In the present study, significantly elevated sNfL levels in the pre-onset group of patients compared to matched controls were observed, especially in the distinct immunogenic cluster that we identified, supporting the notion that neurodegeneration is already occurring as a result of underlying immune-mediated neuroaxonal pathology in the preclinical phase of MS. We cannot completely exclude the impact of other CNS insults on the reported sNfL concentrations (for example, trauma and infection). Although data regarding these factors were not available for correction, our control group included randomly selected age-matched and sex-matched participants from the same population. Therefore, it is unlikely that trauma or other similar events was more common in the MSIC group compared to the MSno-IC group or HCs.
The role of autoantibodies in MS remains unclear; many candidate autoantigens have not survived validation studies21. However, other rare demyelinating diseases of the CNS, such as NMOSD or myelin oligodendrocyte glycoprotein antibody-associated disease (MOGAD), were originally part of the MS spectrum and were definitively separated only after the identification of disease-specific autoantibodies53, findings that also informed subsequent therapeutic development54,55.
Remarkably, prior studies screening smaller numbers of MS patient serum or CSF samples on human and viral antigen libraries uncovered a similar motif as described here in MSIC patients37,56. The identification, corroboration and orthogonal confirmation of this autoantibody signature in two distinct, large patient cohorts across three different disease epochs (that is, preclinical, initial flare and after diagnosis) strongly suggests that the MSIC signature has clinical diagnostic potential, especially in the context of early MS detection. Given its specificity for MS both before and after diagnosis, an autoantibody serology test against the MSIC peptides could be implemented in a surveillance setting for patients with high probability of developing MS or, crucially, at a first clinically isolated neurologic episode.
The similarity of this motif to domains contained in a broad array of infectious agents, including two EBV proteins37,56, as well as limited cross-reactivity within the IC1 patient cluster, suggests that it is possible that infection by one or several of these agents contributes to an autoreactive response and disease pathogenesis, perhaps via molecular mimicry40,57. The role of infection as the temporally initiating event in MS etiology is the focus of current work15,58 and is crucial to the differentiation of causal versus spuriously associated features of the MS prodrome. However, this remains challenging given near-ubiquitous seropositivity for many of the implicated viruses, including EBV, whereas only a small fraction of individuals infected develop MS. Nonetheless, EBV infection, and infectious mononucleosis in particular, represents the most compelling epidemiologic link to MS59,60, including from the DoDSR studied here15. Given the degree of polyspecificity of this motif in the human proteome, it will be important to elucidate the precise origins of the MSIC signature, with special attention to past exposures, genetic risk factors and temporal dynamics of disease manifestation.
This study has several limitations. The small number of patients identified in this cluster makes genetic or other associations difficult. Although similarity exists with this motif and those found in other human infectious agents, such as EBV, this study measured antibodies to only a limited set of antiviral antigens. Two MSIC patients identified by PhIP-Seq were not identified by our Luminex validation panel. Sensitivity would likely be improved by adding more candidates to the multiplexed assay, as the validation panel used in the Luminex assay consists of only six peptides that encompass the motif.
Conclusion
This work identifies a longitudinally stable autoantibody profile that is present before, during and after the time of first symptom onset for a subset of PwMS. Taken together with elevated sNfL levels, this hallmark patient cluster is an attractive target for further immunological and clinicopathologic study. This study, along with other evidence of ongoing neurodegeneration during MS prodrome, suggests that additional evaluations of these patients beyond sNfL and autoantibodies might bear additional insight into underlying immunological processes during this crucial disease phase and stratify patients with different immunological features.
Methods
Patient cohort details
Informed consent
Where applicable, all necessary participant consent was obtained. To maximize privacy, samples were de-identified and not tied to sensitive metadata.
DoDSR cohort
The incident Gulf War Era MS cohort (GWEMSC, n = 2,691) was used as the source for identifying US military MS cases in this study61. The GWEMSC cohort was drawn from the broader US military population that served during the Gulf War era (1990–2007), with relevant demographic and clinical data abstracted from Department of Defense (DoD) and Department of Veterans Affairs (VA) records62. This cohort is population based within the US military and, thereby, has a male preponderance and race and ethnicity subgroups that mirror the US Census. All veterans in this cohort are service connected for MS, which requires evidence of clinical signs upon examination attributable to MS during or within 7 years after active duty military service. Additionally, all cases were adjudicated by the VA MS Center of Excellence study team and met the McDonald MS criteria63.
The DoDSR MS cohort (Veterans Affairs Medical Center, institutional review board (IRB) no. 1624644-4) was created using the clinical information on cases from the GWEMSC and linking serum aliquots before and after first symptom onset from the DoDSR. To build the DoDSR MS cohort, a stratified, population-based sample was created within the GWEMSC. Based on pre-specified demographic strata, the DoDSR staff identified a group of MS cases (n = 250) that had at least one serum aliquot in the repository before MS onset and one aliquot after. The earliest serum sample obtained before onset of MS symptoms from each veteran with MS was identified by DoDSR staff along with a second sample within 2 years after initial MS symptoms. For each MS case, one non-MS HC (n = 250) was randomly selected and matched on year of entry to the military, age, sex, race, ethnicity and dates of blood collection (within 180 d). Although we cannot exclude the possibility that some controls went on to develop MS or another autoimmune condition later in life, we have a high degree of confidence in their health status at the time of their blood draw.
ORIGINS cohort
All patients with RRMS or clinically isolated syndrome (CIS) were participants in the University of California, San Francisco (UCSF; IRB no. 14-15278) ORIGINS study and were diagnosed according to the 2017 McDonald criteria63. Patients were not on immunomodulatory or immunosuppressive disease-modifying therapy at the time of sample collection.
Additional CSF experimental reference controls for ORIGINS PhIP-Seq analysis were generated from 22 healthy or non-MS participants enrolled in a biobanking study, ‘Immunological Studies of Neurologic Subjects’. Additional serum experimental reference controls for ORIGINS PhIP-Seq analysis were generated from 95 reference donors provided by the New York Blood Center.
Nfl measurements
NfL values were measured in duplicate from sera samples with sufficient volumes (n = 944) using the Simoa NF-light Advantage Kit (Quanterix) on an HD-X analyzer by laboratory assistants blinded to the clinical characteristics of the included patients. Samples with a coefficient of variation (%CV) > 20% or where duplicate values were not available were excluded from the final analysis (n = 14). The mean intra-run %CV of the included samples was 5.02% (±3.9%) (n = 930), and the inter-run %CV for the quality control samples were 9.42% and 9.02% for low and high concentrations, respectively.
PhIP-Seq protocol
PhIP-Seq was performed using a high-throughput protocol64. First, 96-well, 2-ml, deep-well polypropylene plates were incubated with a blocking buffer (3% BSA in TBST) overnight at 4 °C to prevent non-specific binding. Blocking buffer was then replaced with 500 µl of freshly grown phage library (at 1011 plaque-forming units per milliliter) and 1 µl of human sera or 10 µl of CSF in 1:1 storage buffer (PBS supplemented with 0.04% NaN3, 40% glycerol, 40 mM HEPES). To facilitate antibody-phage binding, the deep-well plates with library and sample were incubated overnight at 4 °C on a rocker platform in secondary containment. Then, 20 µl of each of Pierce Protein A and G Beads (Thermo Fisher Scientific, 10002D and 10004D) slurry were aliquoted per reaction and washed three times in TNP-40 (140 mM NaCl, 10 mM Tris-HCL, 0.1% NP40). After the final wash, beads were resuspended in TNP-40 in half the slurry volume (20 µl) and added to the phage-patient antibody mixture and incubated on the rocker at 4 °C for 1 h. Beads were then transferred to a vacuum manifold compatible filter plate65 and then washed five times with RIPA (140 mM NaCl, 10 mM Tris-HCl, 1% Triton X, 0.1% SDS) using an Integra VIAFLO 96 to add RIPA each time and using the vacuum to remove the supernatant. In each wash, beads were incubated for 5 min with gentle rocking at 4 °C. After the fifth wash, immunoprecipitated solution was resuspended in 150 µl of LB-Carb and then added to 0.5 ml of log-phase BL5403 E. coli for amplification (OD600 = 0.4–0.6) until lysis was complete (approximately 2 h) on an 800-r.p.m. shaker. After amplification, sterile 5 M NaCl was added to lysed E. coli to a final concentration of 0.5 M NaCl to ensure complete lysis. The lysed solution was spun at 3,220 relative centrifugal force (rcf) for 20 min, and the top 500 µl was filtered through 0.2 mM PVDF filter plates (Arctic White, AWFP-F20080) to remove remaining cell debris. Filtered solution was transferred to a new pre-blocked deep-well plate where patient sera was added and subjected to another round of immunoprecipitation and amplification. The final lysate was spun at 3,220 rcf for 30 min, with supernatant then filtered and stored at 4 °C for subsequent next-generation sequencing (NGS) library prep.
Phage DNA from each sample was barcoded and amplified (Phusion PCR, 18 rounds) and subjected to NGS on an Illumina NovaSeq instrument at an average read depth of 1 million reads per sample. Two post-onset MS samples and one second timepoint control sample library did not pass sequencing quality control and were excluded from analysis.
Luminex assay
Peptides containing the relevant motif from TRIO, KRT75, RIMS2 and SRSF4 (Table 1) were synthesized by LifeTein and conjugated to BSA via SMCC coupling to cysteines on BSA at a 1:1 final mass ratio. Spectrally distinct Luminex beads were conjugated in separate 1.5-ml protein LoBind tubes to their corresponding BSA-peptide antigen. A control bead population was made by conjugating BSA only (Sigma-Aldrich, 10735094001). Each bead conjugation was performed at a concentration of 5 μg of protein per 1 million beads in a 0.5-ml reaction volume. Conjugation was done via an EDC/sulfo–N-hydroxysuccinimide coupling strategy to free amines using an Antibody Coupling Kit following the manufacturer’s instructions (Luminex, 40-50016)64.
All serological analyses were performed in duplicate, and beads were pooled on the day of use. Thawed serum samples were diluted in PBS + 0.05% Tween 20 (PBST) containing 2% non-fat milk and mixed with pooled protein-coated beads (2,000–2,500 beads per protein) at a final serum dilution of 1:250 or final CSF dilution of 1:20. Samples were incubated for 1 h at room temperature at 250 .r.p.m, washed three times with PBST and then stained with 1:2,000 phycoerythrin-conjugated anti-human IgG Fc antibody (BioLegend, 637310) in PBST for 30 min at room temperature. Beads were washed twice with PBST and analyzed in a 96-well plate format on a Luminex LX 200 cytometer. The net MFI for each peptide antigen was calculated for each sample by dividing by the MFI of that sample’s corresponding intra-assay BSA negative control and averaging across duplicate wells. A cutoff of log(MFI) > 3 for each CSF sample was chosen to maximize the specificity of the assay.
Sequencing and HLA genotyping
For each sample, 100 ng of high-quality DNA was fragmented with the Library Preparation Enzymatic Fragmentation Kit 2.0 (Twist Bioscience). After fragmentation, DNA was repaired, and poly(A) tails were attached and ligated to Illumina-compatible dual index adapters with unique barcodes. After ligating, fragments were purified with 0.8× ratio AMPure XP magnetic beads (Beckman Coulter). Double size selection was performed (0.42× and 0.15× ratios), and libraries of approximately 800 bp were selected, at which point libraries were amplified and purified using magnetic beads.
After fluorometric quantification, each sample was pooled (30 ng per sample) via ultrasonic acoustic energy. A Twist Target Enrichment Kit (Twist Bioscience) was then used to perform target capture on pooled samples. Sample volumes were then reduced using magnetic beads, and DNA libraries were bound to 1,394 biotinylated probes. Probes were designed specifically to target all exons, introns and regulatory regions of the classical HLA loci, including HLA-A, HLA-B, HLA-C, HLA-DPB1, HLA-DRB1 and HLA-DQB1. Then, streptavidin magnetic beads were used to capture fragments targeted by the probes. Captured fragments were then amplified and purified. Bioanalyzer (Agilent) was then used to analyze the enriched libraries. After evaluation, enriched libraries were sequenced using a paired-end 150-bp sequencing protocol on the NovaSeq platform (Illumina). After sequencing, HLA genotypes were predicted using HLA Explorer (Omixon).
Statistics and analysis
sNfL analysis
Independent samples Studentʼs t-test (equal variance not assumed) was applied to compare log-transformed NfL concentrations between samples collected from PwMS and age-matched and gender-matched HCs at each timepoint separately. To compare between longitudinal NfL concentration in MSIC and MSNO-IC groups, mixed linear models assessed log-transformed NfL concentrations as dependent variables in the three groups (HC, MSIC and MSNO-IC) correcting for age at sampling (fixed effects). All reported P values were Bonferroni adjusted.
Predicted NfL concentrations used to plot their dynamics in the three groups over the duration between the first and second samples were fitted using a linear mixed model of NfL concentration (log-transformed) accounting for the group (HC, MSIC and MSNO-IC) and duration between samples in relation to documented MS onset, with an additional interaction term (groups × duration). The statistical analysis was conducted with SPSS (version 28) and JMP Pro (version 16) software.
PhIP-Seq
Raw sequencing reads were aligned to a reference library using Bowtie 2 (ref. 64). To correct for differences in sequencing depth, read counts for each peptide within a sample were normalized to the total reads expressed as reads per 100,000 (RPK). FC for each peptide was generated from the mean RPK of the controls, and a z-score was calculated from the background distribution.
A custom bioinformatics pipeline was used to generate a list of candidate peptides enriched in the MS group. For each peptide, FC values for all MS samples were calculated by dividing the RPK value by the mean RPK of all matched HC samples. FC values were then used to calculate z-score values. The first round of filters identified enriched peptides with a minimum RPK ≥ 1 and an FC ≥ 10. Among enriched peptides, more stringent cutoffs were applied to identify candidate peptides. This included FC ≥ 100 and z-score ≥ 10. Mutually enriched peptides from the same gene were also added to the candidate peptide list if they had FC ≥ 100 and z-score ≥ 3. Additionally, peptides from given genes were included in the candidate list only if they had a sum RPK ≥ 50. A seven-amino-acid (7AA) k-mer analysis was performed on all enriched peptides. Enriched peptides that shared an identical 7AA sequence with at least one other peptide were kept for further analysis.
Because a bona fide autoantibody would likely target the same epitope on multiple enriched peptides, the candidate list was further narrowed by filtering out peptides without k-mer overlaps. Given the size of the cohort and the tendency of individuals to enrich highly personalized epitopes, we further narrowed the candidate list requiring peptides to have an FC > 10 in at least 15 of 250 MS cases34.
Defining patient clusters for the DoD cohort
When we performed hierarchical clustering of the resulting candidate peptides, we found that a subset of PwMS and two HCs (n = 18) enriched several peptides bearing a nearly identical epitope. Unsupervised clustering with UMAP was used to further define patient clusters among enriched candidate peptides. Among samples in the DoD cohort, all MS and HC samples were clustered using only 192 enriched peptides from 151 proteins that were a product of the cutoffs stated above. Separately, a smaller panel of enriched peptides was used to further define subclusters. To assemble this panel of peptides, for each peptide both the sum RPK across all PwMS and sum RPK across all HC samples was calculated. These values were used to calculate a sum RPK ratio of MS to HC. Peptides with a sum RPK > 800 across all samples and an MS-to-HC ratio greater than 6 were kept in our smaller target panel of 13 peptides (nine proteins). For each sample, both the sum RPK of our target panel as well as the sum RPK for all 119 peptides bearing the characterized motif were used.
Unsupervised clustering with UMAP (n_neighbors = 20, n_components = 3, learning_rate = 1, init = ‘random’, n_epochs = 100,000) revealed one outlier cluster of 49 samples that contained increased expression in both of our target peptide panels. This subset of samples was re-clustered under the same parameters, which created two patient clusters. Cluster 1 contained 30 samples (n = 18) that showed higher sum RPK values among our target panel relative to the 19 samples in cluster 2 (n = 12).
Defining patient clusters for the ORIGINS cohort
The same custom bioinformatics pipeline was used to identify enriched peptides among CSF and serum samples within the ORIGINS cohort. To identify the top enriched peptides in the CSF and serum, the sum FC of samples with FC > 10 was calculated. In total, this filtered out 2,307 peptides in the CSF (sum FC > 10) and 289 peptides (sum FC > 12) in the serum. A panel of 26 peptides that were enriched in nine or more DoD samples (Fig. 1b) was used to identify patient clusters. The sum RPK of this panel was calculated across all samples. Hierarchical clustering was used to identify a subset of 16 samples (n = 8) with the highest sum RPK. Only samples with a sum RPK > 200 in our panel were kept. This subset of samples was divided into two clusters due to the difference in the FC expression profile among the panel of 26 peptides. Cluster 1 (n = 4) showed more pronounced expression in RIMS2, TRA2B, KRT75, SRRM3 and TRIO. Cluster 2 (n = 4) showed more pronounced expression in CLASRP and SRSF4.
We used the following cutoffs in our analysis code (package, PairSeq) to generate the candidate list of peptides:
MIN_RPK = 1
ZSCORE_THRESH1 = 3
ZSCORE_THRESH2 = 10
FC_THRESH1 = 10
FC_THRESH2 = 100
SUM_RPK_THRESH = 50
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The PROSITE database used for this study is freely available (https://prosite.expasy.org/). All data produced in the present study are available online at https://github.com/UCSF-Wilson-Lab/MS_DoD_and_ORIGINS_study_data.
Code availability
All analysis code can be found in the following publicly available repository: https://github.com/UCSF-Wilson-Lab/MS_DoD_and_ORIGINS_study_data. PairSeq package source code can be found in the following publicly available repository: https://github.com/cmbartley/PairSeq.
References
Reynolds, R. et al. The neuropathological basis of clinical progression in multiple sclerosis. Acta Neuropathol. 122, 155–170 (2011).
Freedman, M. S. Multiple Sclerosis and Demyelinating Diseases (Lippincott Williams & Wilkins, 2006).
Sospedra, M. & Martin, R. Immunology of multiple sclerosis. Annu. Rev. Immunol. 23, 683–747 (2005).
Lassmann, H., Brück, W. & Lucchinetti, C. F. The immunopathology of multiple sclerosis: an overview. Brain Pathol. 17, 210–218 (2007).
Li, R., Patterson, K. R. & Bar-Or, A. Reassessing B cell contributions in multiple sclerosis. Nat. Immunol. 19, 696–707 (2018).
Sabatino, J. J., Pröbstel, A.-K. & Zamvil, S. S. B cells in autoimmune and neurodegenerative central nervous system diseases. Nat. Rev. Neurosci. 20, 728–745 (2019).
Walton, C. et al. Rising prevalence of multiple sclerosis worldwide: insights from the Atlas of MS, third edition. Mult. Scler. 26, 1816–1821 (2020).
Wallin, M. T. et al. The prevalence of MS in the United States: a population-based estimate using health claims data. Neurology 92, e1029–e1040 (2019).
Rovira, A. et al. A single, early magnetic resonance imaging study in the diagnosis of multiple sclerosis. Arch. Neurol. 66, 587–592 (2009).
Lebrun-Frénay, C. et al. Risk factors and time to clinical symptoms of multiple sclerosis among patients with radiologically isolated syndrome. JAMA Netw. Open 4, e2128271 (2021).
Högg, T. et al. Mining healthcare data for markers of the multiple sclerosis prodrome. Mult. Scler. Relat. Disord. 25, 232–240 (2018).
Disanto, G. et al. Prodromal symptoms of multiple sclerosis in primary care. Ann. Neurol. 83, 1162–1173 (2018).
Hauser, S. L. & Oksenberg, J. R. The neurobiology of multiple sclerosis: genes, inflammation, and neurodegeneration. Neuron 52, 61–76 (2006).
Bjornevik, K. et al. Serum neurofilament light chain levels in patients with presymptomatic multiple sclerosis. JAMA Neurol. 77, 58–64 (2020).
Bjornevik, K. et al. Longitudinal analysis reveals high prevalence of Epstein–Barr virus associated with multiple sclerosis. Science 375, 296–301 (2022).
Ning, L. & Wang, B. Neurofilament light chain in blood as a diagnostic and predictive biomarker for multiple sclerosis: a systematic review and meta-analysis. PLoS ONE 17, e0274565 (2022).
Arbuckle, M. R. et al. Development of autoantibodies before the clinical onset of systemic lupus erythematosus. N. Engl. J. Med. 349, 1526–1533 (2003).
Primavera, M., Giannini, C. & Chiarelli, F. Prediction and prevention of type 1 diabetes. Front. Endocrinol. (Lausanne) 11, 248 (2020).
Alpizar-Rodriguez, D. & Finckh, A. Is the prevention of rheumatoid arthritis possible? Clin. Rheumatol. 39, 1383–1389 (2020).
Hundt, J. E., Hoffmann, M. H., Amber, K. T. & Ludwig, R. J. Editorial: Autoimmune pre-disease. Front. Immunol. 14, 1159396 (2023).
Höftberger, R., Lassmann, H., Berger, T. & Reindl, M. Pathogenic autoantibodies in multiple sclerosis—from a simple idea to a complex concept. Nat. Rev. Neurol. 18, 681–688 (2022).
Kuerten, S. et al. Autoantibodies against central nervous system antigens in a subset of B cell–dominant multiple sclerosis patients. Proc. Natl Acad. Sci. USA 117, 21512–21518 (2020).
McLaughlin, K. A. & Wucherpfennig, K. W. B cells and autoantibodies in the pathogenesis of multiple sclerosis and related inflammatory demyelinating diseases. Adv. Immunol. 98, 121–149 (2008).
Ramesh, A. et al. A pathogenic and clonally expanded B cell transcriptome in active multiple sclerosis. Proc. Natl Acad. Sci. USA 117, 22932–22943 (2020).
Willis, S. N. et al. Investigating the antigen specificity of multiple sclerosis central nervous system-derived immunoglobulins. Front. Immunol. 6, 600 (2015).
Elliott, C. et al. Functional identification of pathogenic autoantibody responses in patients with multiple sclerosis. Brain 135, 1819–1833 (2012).
Greenfield, A. L. et al. Longitudinally persistent cerebrospinal fluid B-cells can resist treatment in multiple sclerosis. JCI Insight 4, e126599 (2019).
Petzold, A. Intrathecal oligoclonal IgG synthesis in multiple sclerosis. J. Neuroimmunol. 262, 1–10 (2013).
Wijnands, J. M. A. et al. Prodrome in relapsing-remitting and primary progressive multiple sclerosis. Eur. J. Neurol. 26, 1032–1036 (2019).
Tremlett, H. & Marrie, R. A. The multiple sclerosis prodrome: emerging evidence, challenges, and opportunities. Mult. Scler. 27, 6–12 (2021).
Pavlin, J. A. & Welch, R. A. Ethics, human use, and the Department of Defense Serum Repository. Mil. Med. 180, 49–56 (2015).
Larman, H. B. et al. Autoantigen discovery with a synthetic human peptidome. Nat. Biotechnol. 29, 535–541 (2011).
Mandel-Brehm, C. et al. Kelch-like protein 11 antibodies in seminoma-associated paraneoplastic encephalitis. N. Engl. J. Med. 381, 47–54 (2019).
Vazquez, S. E. et al. Identification of novel, clinically correlated autoantigens in the monogenic autoimmune syndrome APS1 by proteome-wide PhIP-Seq. eLife 9, e55053 (2020).
Mandel-Brehm, C. et al. Autoantibodies to perilipin-1 define a subset of acquired generalized lipodystrophy. Diabetes 72, 59–70 (2022). db211172.
Mandel-Brehm, C. et al. ZSCAN1 autoantibodies are associated with pediatric paraneoplastic ROHHAD. Ann. Neurol. 92, 279–291 (2022).
Larman, H. B. et al. PhIP-Seq characterization of autoantibodies from patients with multiple sclerosis, type 1 diabetes and rheumatoid arthritis. J. Autoimmun. 43, 1–9 (2013).
Rasquinha, M. T. et al. PhIP-Seq reveals autoantibodies for ubiquitously expressed antigens in viral myocarditis. Biology 11, 1055 (2022).
Rasquinha, M. T., Lasrado, N., Larman, B. H. & Reddy, J. PhIP-Seq analysis reveals autoantibodies for novel antigens in the mouse model of Coxsackievirus B3 infection. J. Immunol. 206, 21.19 (2021).
Lanz, T. V. et al. Clonally expanded B cells in multiple sclerosis bind EBV EBNA1 and GlialCAM. Nature 603, 321–327 (2022).
Ayoglu, B. et al. Anoctamin 2 identified as an autoimmune target in multiple sclerosis. Proc. Natl Acad. Sci. USA 113, 2188–2193 (2016).
Tatsuno, T. & Ishigaki, Y. C-terminal short arginine/serine repeat sequence-dependent regulation of Y14 (RBM8A) localization. Sci. Rep. 8, 612 (2018).
Philipps, D., Celotto, A. M., Wang, Q.-Q., Tarng, R. S. & Graveley, B. R. Arginine/serine repeats are sufficient to constitute a splicing activation domain. Nucleic Acids Res. 31, 6502–6508 (2003).
Richardson, D. N. et al. Comparative analysis of serine/arginine-rich proteins across 27 eukaryotes: insights into sub-family classification and extent of alternative splicing. PLoS ONE 6, e24542 (2011).
Sigrist, C. J. A. et al. New and continuing developments at PROSITE. Nucleic Acids Res. 41, D344–D347 (2013).
Wijnands, J. M. A. et al. Health-care use before a first demyelinating event suggestive of a multiple sclerosis prodrome: a matched cohort study. Lancet Neurol. 16, 445–451 (2017).
Cortese, M. et al. Preclinical disease activity in multiple sclerosis: a prospective study of cognitive performance prior to first symptom. Ann. Neurol. 80, 616–624 (2016).
Khalil, M. et al. Neurofilaments as biomarkers in neurological disorders. Nat. Rev. Neurol. 14, 577–589 (2018).
Disanto, G. et al. Serum neurofilament light: a biomarker of neuronal damage in multiple sclerosis. Ann. Neurol. 81, 857–870 (2017).
Bose, G. et al. Early neurofilament light and glial fibrillary acidic protein levels improve predictive models of multiple sclerosis outcomes. Mult. Scler. Relat. Disord. 74, 104695 (2023).
Jons, D. et al. Axonal injury in asymptomatic individuals preceding onset of multiple sclerosis. Ann. Clin. Transl. Neurol. 9, 882–887 (2022).
Benkert, P. et al. Serum neurofilament light chain for individual prognostication of disease activity in people with multiple sclerosis: a retrospective modelling and validation study. Lancet Neurol. 21, 246–257 (2022).
Häusser-Kinzel, S. & Weber, M. S. The role of B cells and antibodies in multiple sclerosis, neuromyelitis optica, and related disorders. Front. Immunol. 10, 201 (2019).
Sechi, E. et al. Myelin oligodendrocyte glycoprotein antibody-associated disease (MOGAD): a review of clinical and MRI features, diagnosis, and management. Front. Neurol. 13, 885218 (2022).
Wingerchuk, D. M. & Lucchinetti, C. F. Neuromyelitis optica spectrum disorder. N. Engl. J. Med. 387, 631–639 (2022).
Cepok, S. et al. Identification of Epstein–Barr virus proteins as putative targets of the immune response in multiple sclerosis. J. Clin. Invest. 115, 1352–1360 (2005).
Riedhammer, C. & Weissert, R. Antigen presentation, autoantigens, and immune regulation in multiple sclerosis and other autoimmune diseases. Front. Immunol. 6, 322 (2015).
Comabella, M. et al. Increased cytomegalovirus immune responses at disease onset are protective in the long-term prognosis of patients with multiple sclerosis. J. Neurol. Neurosurg. Psychiatry 94, 173–180 (2022).
Bar-Or, A. et al. Epstein–Barr virus in multiple sclerosis: theory and emerging immunotherapies. Trends Mol. Med. 26, 296–310 (2020).
Soldan, S. S. & Lieberman, P. M. Epstein–Barr virus and multiple sclerosis. Nat. Rev. Microbiol. 21, 51–64 (2023).
Wallin, M. T. et al. The Gulf War era multiple sclerosis cohort: age and incidence rates by race, sex and service. Brain 135, 1778–1785 (2012).
Wallin, M. T., Culpepper, W. J., Maloni, H. & Kurtzke, J. F. The Gulf War era multiple sclerosis cohort: 3. Early clinical features. Acta Neurol. Scand. 137, 76–84 (2018).
Thompson, A. J. et al. Diagnosis of multiple sclerosis: 2017 revisions of the McDonald criteria. Lancet Neurol. 17, 162–173 (2018).
Zamecnik, C. R. et al. ReScan, a multiplex diagnostic pipeline, pans human sera for SARS-CoV-2 antigens. Cell Rep. Med. 1, 100123 (2020).
Vazquez, S. E. et al. Autoantibody discovery across monogenic, acquired, and COVID-19-associated autoimmunity with scalable PhIP-seq. eLife 11, e78550 (2022).
Acknowledgements
This work was supported by the Valhalla Foundation (S.L.H., M.R.W., J.R.O. and B.A.C.); the Weill Neurohub (C.R.Z.); the Westridge Foundation (M.R.W., H.-C.v.B. and A.J.G.); National Institute of Neurological Disorders and Stroke R35NS122073 (S.L.H., M.R.W. and R.D.); National Institute of Allergy and Infectious Diseases R01AI170863-01A1 (S.S.Z.); National Multiple Sclerosis Society RFA-2104-37504 (M.R.W., M.T.W., J.H., B.A.C., C.R.Z., R.D. and R.D.B.); National Multiple Sclerosis Society RFA-2104-3747 (J.R.O.); the Department of Defense (A.A.); the German Society of Multiple Sclerosis (A.A.); the Water Cove Charitable Foundation (M.R.W., M.T.W., J.H., B.A.C. and C.R.Z.); Tim and Laura O’Shaughnessy (M.R.W.); the Littera Family (M.R.W.); the School of Medicine Dean’s Yearlong Fellowship, supported by residual funds from the Howard Hughes Medical Institute Medical Fellows at UCSF (G.M.S.); the Chan Zuckerberg Biohub San Francisco (J.L.D. and S.A.M.); the John A. Watson Scholar Program at UCSF (C.M.B.); the Hanna H. Gray Fellowship, Howard Hughes Medical Institute (C.M.B.); National Institutes of Health R01AI158861 (J.A.H. and K.J.W.); and the University of California President’s Postdoctoral Fellowship Program (C.M.B.). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript. The authors would like to acknowledge the Chan Zuckerberg Biohub San Francisco and its sequencing team for assistance. The following members of the Walter Reed National Military Medical Center Neurology Department were collaborators on this project: A. Lewis and N. Tagg. Data were obtained from the Defense Medical Surveillance System, Armed Forces Health Surveillance Branch, Defense Health Agency (data from 1987–2007, released 2010). Figures 1 and 5a were created with BioRender. The authors would also like to thank the patients and their families for their contribution to this study.
Author information
Authors and Affiliations
Contributions
C.R.Z., G.M.S., M.R.W., M.T.W. and J.L.D. designed the experiments. C.R.Z. performed all PhIP-Seq and Luminex assays. R.D.B. and C.M.C. performed the Luminex assays, and G.M.S. and S.A.M. performed PhIP-Seq immunoprecipitations and NGS library preparation. S.A.M., C.F. and A.E.W. assisted with DoDSR sample preparation. A.A., N.J. and K.A. performed the sNfL studies, and A.A. analyzed the data. R.D. and C.M.B. built the analysis code and pipeline for PhIP-Seq data and analyzed the PhIP-Seq data, with G.M.S., as well as generated figures. K.C.Z., A.T., C.F., J.A., C.G., E.B.T., H.-C.v.B., R.P.L., R.G. and E.L.E. curated and collected patient samples and clinical data for the UCSF ORIGINS cohort. K.J.W., J.R.O., D.G.A., K.K. and J.A.H. performed the HLA sequencing, genotyping and association. M.T.W. generated the DoDSR cohort and epidemiologic analysis, and A.J.G. contributed valuable sNfL analysis interpretation. S.L.H., B.A.C.C. and J.R.O., together with the UCSF MS-EPIC Team and J.M.G., C.-Y.G. and R.C., led the recruitment and phenotyping of the UCSF ORIGINS cohort. S.L.H., B.A.C.C. and J.L.D. provided valuable experimental guidance and manuscript editing. All authors contributed to study design. C.R.Z., G.M.S. and M.R.W. wrote the manuscript. All authors provided editorial comments.
Corresponding author
Ethics declarations
Competing interests
M.R.W. receives unrelated research grant funding from Roche/Genentech and Novartis and has received speaking honoraria from Genentech, Takeda, WebMD and Novartis. M.R.W. and J.L.D. receive licensing fees from CDI Labs. C.M.B. serves as a paid consultant for the Neuroimmune Foundation. J.J.S. has unrelated research grant funding from Roche/Genentech and Novartis and advisory board honoraria from IgM Biosciences. C.-Y.G. has received financial compensation from serving on advisory boards for Genentech and Horizon. R.G.H. has unrelated research funding from Roche/Genentech and Atara Bio; consulting fees from Roche/Genentech, Novartis and QIA Consulting; and discussion leader fees from Sanofi/Genzyme. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Medicine thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Jerome Staal, in collaboration with the Nature Medicine team
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1
Heatmap of logRPK values from whole proteome. PhIP-Seq data on MS patient sera from DoDSR cohort. Top peptides shown with peptide enrichment. blowout to demonstrate similarity between pre and post onset sample enrichments over time.
Extended Data Fig. 2
We investigated the similarity between different timepoints from the from the DoDSR cohort (n = 500). (Top Left) Unfiltered normalized read counts (rpK) were log transformed, and a correlation matrix was computed using the Euclidean distance measure. The similarity between any two samples was defined by their n-dimensional Euclidean distance, where each peptide in the library was treated as a dimension. The rows and columns of this matrix are ordered such that the early timepoints are followed by the late timepoints. The central diagonal stripe in the heatmap represents a distance of zero where the distance was computed between the same sample. However, the diagonal stripes represent distances calculated between different timepoints of the same sample. (Top Right) Box plots of the Euclidean distance between timepoints, with a trend towards increasing distance at longer intervals. However, patterns of self-reactivity remain stable for more than 15 years. Notches approximate 95% confidence intervals. (Bottom) A boxplot demonstrates the significant difference (two-sided t-test) in Euclidean distance between two timepoints of the same sample versus the distance between two unrelated samples. Notches approximate 95% confidence intervals.
Extended Data Fig. 3
A subset of the library capturing relevant motifs from previously identified putative targets of molecular mimicry such as GlialCAM and anoctamin-2. (Top) MS patient samples are ordered so as to preserve the IC clusters. Although individual samples can be seen to enrich peptides of interest, they did not meet the predefined cutoffs for this analysis. (Bottom) HC samples demonstrating background levels of enrichment.
Extended Data Fig. 4
Time point specific NfL data from DoD cohort. (1st time point: HC n = 236, MSno-IC n = 204, MSIC n = 13. 2nd time point: HC n = 234, MSno-IC n = 191, MSIC n = 21. Data represented as geometric mean and standard factors for each box and whisker.).
Extended Data Fig. 5
Additional OND CSF controls (n = 20), lacking enrichment of the IC motif.
Extended Data Fig. 6
Correlation of sum of MFI from all six peptides of interest in Table 1 from Luminex assay with their sum of respective normalized read counts from PhIP-Seq data in both CSF and serum samples from ORIGINS cohort. Shaded area represents 95% confidence intervals.
Extended Data Fig. 7
Luminex assay for antibodies against listed pathogen peptides in ORIGINS cohort a) MS patient CSF (n = 104), b) MS patient serum (n = 104), c) OND patient CSF(n = 42) and d) OND patient serum (n = 22). e) Same patient populations respresnted as sum of MFI combining signal across pathogen peptides with highlighted patients having normalized sumMFI values > 3 in CSF.
Supplementary information
Supplementary Tables 1 and 2
Supplementary Table 1: HLA data for the ORIGINS cohort; Supplementary Table 2: HLA allele frequency assessment.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zamecnik, C.R., Sowa, G.M., Abdelhak, A. et al. An autoantibody signature predictive for multiple sclerosis. Nat Med (2024). https://doi.org/10.1038/s41591-024-02938-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41591-024-02938-3