Abstract
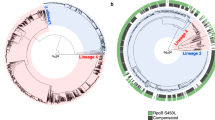
Multidrug-resistant tuberculosis (MDR-TB) accounts for one third of the annual deaths due to antimicrobial resistance1. Drug resistance-conferring mutations frequently cause fitness costs in bacteria2,3,4,5. Experimental work indicates that these drug resistance-related fitness costs might be mitigated by compensatory mutations6,7,8,9,10. However, the clinical relevance of compensatory evolution remains poorly understood. Here we show that, in the country of Georgia, during a 6-year nationwide study, 63% of MDR-TB was due to patient-to-patient transmission. Compensatory mutations and patient incarceration were independently associated with transmission. Furthermore, compensatory mutations were overrepresented among isolates from incarcerated individuals that also frequently spilled over into the non-incarcerated population. As a result, up to 31% of MDR-TB in Georgia was directly or indirectly linked to prisons. We conclude that prisons fuel the epidemic of MDR-TB in Georgia by acting as ecological drivers of fitness-compensated strains with high transmission potential.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The raw sequences were deposited at the European Nucleotide Archive under BioProject ID PRJEB39561. Accession numbers are listed in Supplementary Table 2. Metadata associated with the genomes are provided in Supplementary Table 2.
Change history
01 June 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41591-021-01417-3
References
O’Neill, J. Antimicrobial Resistance: Tackling a Crisis for the Health and Wealth of Nations. http://www.jpiamr.eu/wp-content/uploads/2014/12/AMR-Review-Paper-Tackling-a-crisis-for-the-health-and-wealth-of-nations_1-2.pdf (Joint Programming Initiative on Antimicrobial Resistance, 2014).
Andersson, D. I. & Levin, B. R. The biological cost of antibiotic resistance. Curr. Opin. Microbiol. 2, 489–493 (1999).
Andersson, D. I. & Hughes, D. Antibiotic resistance and its cost: is it possible to reverse resistance? Nat. Rev. Microbiol. 8, 260–271 (2010).
Gygli, S. M., Borrell, S., Trauner, A. & Gagneux, S. Antimicrobial resistance in Mycobacterium tuberculosis: mechanistic and evolutionary perspectives. FEMS Microbiol. Rev. 41, 354–373 (2017).
Gagneux, S. et al. The competitive cost of antibiotic resistance in Mycobacterium tuberculosis. Science 312, 1944–1946 (2006).
Reynolds, M. G. Compensatory evolution in rifampin-resistant Escherichia coli. Genetics 156, 1471–1481 (2000).
Song, T. et al. Fitness costs of rifampicin resistance in Mycobacterium tuberculosis are amplified under conditions of nutrient starvation and compensated by mutation in the β′ subunit of RNA polymerase. Mol. Microbiol. 91, 1106–1119 (2014).
Maisnier-Patin, S., Berg, O. G., Liljas, L. & Andersson, D. I. Compensatory adaptation to the deleterious effect of antibiotic resistance in Salmonella typhimurium. Mol. Microbiol. 46, 355–366 (2002).
Qi, Q., Preston, G. M. & MacLean, R. C. Linking system-wide impacts of RNA polymerase mutations to the fitness cost of rifampin resistance in Pseudomonas aeruginosa. mBio 5, 1–12 (2014).
Comas, I. et al. Whole-genome sequencing of rifampicin-resistant Mycobacterium tuberculosis strains identifies compensatory mutations in RNA polymerase genes. Nat. Genet. 44, 106–110 (2011).
World Health Organization. Global Tuberculosis Report 2020. https://www.who.int/publications/i/item/9789240013131 (World Health Organization, 2020).
Dye, C. Doomsday postponed? Preventing and reversing epidemics of drug-resistant tuberculosis. Nat. Rev. Microbiol. 7, 81–87 (2009).
Dye, C. & Espinal, M. A. Will tuberculosis become resistant to all antibiotics? Proc. R. Soc. B Biol. Sci. 268, 45–52 (2001).
Dye, C., Williams, B. G., Espinal, M. A. & Raviglione, M. C. Erasing the world’s slow stain: strategies to beat multidrug-resistant tuberculosis. Science 295, 2042–2046 (2002).
World Health Organization. Global Tuberculosis Report 2019. https://www.who.int/teams/global-tuberculosis-programme/tb-reports/global-report-2019 (World Health Organization, 2019).
Casali, N. et al. Evolution and transmission of drug-resistant tuberculosis in a Russian population. Nat. Genet. 46, 279–286 (2014).
Eldholm, V. et al. Four decades of transmission of a multidrug-resistant Mycobacterium tuberculosis outbreak strain. Nat. Commun. 6, 7119 (2015).
Gagneux, S. et al. Impact of bacterial genetics on the transmission of isoniazid-resistant Mycobacterium tuberculosis. PLoS Pathog. 2, e61 (2006).
Yang, C. et al. Transmission of multidrug-resistant Mycobacterium tuberculosis in Shanghai, China: a retrospective observational study using whole-genome sequencing and epidemiological investigation. Lancet Infect. Dis. 3099, 1–10 (2016).
Brandis, G., Wrande, M., Liljas, L. & Hughes, D. Fitness-compensatory mutations in rifampicin-resistant RNA polymerase. Mol. Microbiol. 85, 142–151 (2012).
Merker, M. et al. Compensatory evolution drives multidrug-resistant tuberculosis in Central Asia. eLife 7, 1–31 (2018).
Liu, Q. et al. Have compensatory mutations facilitated the current epidemic of multidrug-resistant tuberculosis? Emerg. Microbes Infect. 7, 98 (2018).
de Vos, M. et al. Putative compensatory mutations in the rpoC gene of rifampin-resistant Mycobacterium tuberculosis are associated with ongoing transmission. Antimicrob. Agents Chemother. 57, 827–832 (2013).
Klinkenberg, D., Backer, J. A., Didelot, X., Colijn, C. & Wallinga, J. Simultaneous inference of phylogenetic and transmission trees in infectious disease outbreaks. PLoS Comput. Biol. 13, e1005495 (2017).
Ahmad, N. et al. Treatment correlates of successful outcomes in pulmonary multidrug-resistant tuberculosis: an individual patient data meta-analysis. Lancet 392, 821–834 (2018).
Lomtadze, N. et al. Prevalence and risk factors for multidrug-resistant tuberculosis in the Republic of Georgia: a population-based study. Int. J. Tuberc. Lung Dis. 13, 68–73 (2009).
World Health Organization. Companion Handbook to the WHO Guidelines for the Programmatic Management of Drug-resistant Tuberculosis. https://www.who.int/tb/publications/pmdt_companionhandbook/en/ (World Health Organization, 2014).
Ismail, N. A. et al. Defining bedaquiline susceptibility, resistance, cross-resistance and associated genetic determinants: a retrospective cohort study. EBioMedicine 28, 136–142 (2018).
World Health Organization. Extensive Review of Tuberculosis Prevention, Control and Care in Georgia 6–14 November 2014. https://www.euro.who.int/en/health-topics/communicable-diseases/tuberculosis/publications/2016/extensive-review-of-tuberculosis-prevention,-control-and-care-in-georgia-614-november-2014 (World Health Organization, 2014).
Nimmo, C. et al. Population-level emergence of bedaquiline and clofazimine resistance-associated variants among patients with drug-resistant tuberculosis in southern Africa: a phenotypic and phylogenetic analysis. Lancet Microbe 1, e165–e174 (2020).
Behr, M. A., Edelstein, P. H. & Ramakrishnan, L. Revisiting the timetable of tuberculosis. BMJ 362, k2738 (2018).
Stuckler, D., Basu, S., McKee, M. & King, L. Mass incarceration can explain population increases in TB and multidrug-resistant TB in European and central Asian countries. Proc. Natl Acad. Sci. USA 105, 13280–13285 (2008).
Kenyon, G. Russia’s prisons fuel drug-resistant tuberculosis. Lancet Infect. Dis. 9, 594 (2009).
Aerts, A. et al. Pulmonary tuberculosis in prisons of the ex-USSR state Georgia: results of a nation-wide prevalence survey among sentenced inmates. Int. J. Tuberc. Lung Dis. 4, 1104–1110 (2000).
Yates, T. A. et al. The transmission of Mycobacterium tuberculosis in high burden settings. Lancet Infect. Dis. 16, 227–238 (2016).
Niemann, S. et al. Mycobacterium tuberculosis Beijing lineage favors the spread of multidrug-resistant tuberculosis in the Republic of Georgia. J. Clin. Microbiol. 48, 3544–3550 (2010).
Baussano, I. et al. Tuberculosis incidence in prisons: a systematic review. PLoS Med. 7, 1–10 (2010).
Gegia, M., Kalandadze, I., Madzgharashvili, M. & Furin, J. Developing a human rights-based program for tuberculosis control in Georgian prisons. Health Hum. Rights 13, 9 (2011).
Menardo, F. et al. Treemmer: a tool to reduce large phylogenetic datasets with minimal loss of diversity. BMC Bioinformatics 19, 164 (2018).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Koboldt, D. C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576 (2012).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff. Fly 6, 80–92 (2012).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Coll, F. et al. A robust SNP barcode for typing Mycobacterium tuberculosis complex strains. Nat. Commun. 5, 4812 (2014).
Stefan, M. A., Ugur, F. S. & Garcia, G. A. Source of the fitness defect in rifamycin-resistant Mycobacterium tuberculosis RNA polymerase and the mechanism of compensation by mutations in the β′ subunit. Antimicrob. Agents Chemother. 62, e00164–18 (2018).
Walker, T. M. et al. Whole-genome sequencing to delineate Mycobacterium tuberculosis outbreaks: a retrospective observational study. Lancet Infect. Dis. 13, 137–146 (2013).
Menardo, F., Duchêne, S., Brites, D. & Gagneux, S. The molecular clock of Mycobacterium tuberculosis. PLoS Pathog. 15, e1008067 (2019).
Lanfear, R., Hua, X. & Warren, D. L. Estimating the effective sample size of tree topologies from Bayesian phylogenetic analyses. Genome Biol. Evol. 8, 2319–2332 (2016).
Acknowledgements
Calculations were performed at the sciCORE (http://scicore.unibas.ch/) scientific computing core facility at the University of Basel. We would like to thank D. Klinkenberg (Dutch National Institute for Public Health and the Environment) for help with the phybreak analysis and D. Brites for feedback on the manuscript. We also thank J. Andrews and the two other anonymous reviewers for their excellent comments that helped us improve our manuscript. Funding: This work was supported by the Swiss National Science Foundation (grants 310030_188888, IZRJZ3_164171, IZLSZ3_170834 and CRSII5_177163, all to S.G.), the European Research Council (309540-EVODRTB and 883582-ECOEVODRTB, both to S.G.) and SystemsX.ch (to S.G. and C.B.).
Author information
Authors and Affiliations
Contributions
S.G. and L.J. conceived the idea. S.G. and Z.A. supervised the project. S.M.G. performed data curation, data analysis and wrote the first draft. C.L. performed data curation and data analysis. L.J. supervised data acquisition and curation. N.A., R.A., M.R. and N.M. carried out data acquisition. A.T. and A.R. carried out data analysis. S.B. supervised data acquisition. K.R. critically reviewed the drafts. C.B. and N.T. supervised data acquisition. All authors reviewed the draft and assisted in the manuscript preparation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Medicine thanks Jason Andrews and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Editor recognition statement: Alison Farrell was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Results; Supplementary Tables 1, 3, 5, 6, 8 and 14–19; Supplementary References; and Supplementary Figs. 1–8.
Supplementary Tables
Supplementary Tables 2, 4, 7 and 9–13
Rights and permissions
About this article
Cite this article
Gygli, S.M., Loiseau, C., Jugheli, L. et al. Prisons as ecological drivers of fitness-compensated multidrug-resistant Mycobacterium tuberculosis. Nat Med 27, 1171–1177 (2021). https://doi.org/10.1038/s41591-021-01358-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-021-01358-x
This article is cited by
-
On the onset and dispersal of a major MDR TB clone among HIV-negative patients, Tunisia
Antimicrobial Resistance & Infection Control (2024)
-
Phylogeography and transmission of Mycobacterium tuberculosis spanning prisons and surrounding communities in Paraguay
Nature Communications (2023)
-
Regulatory fine-tuning of mcr-1 increases bacterial fitness and stabilises antibiotic resistance in agricultural settings
The ISME Journal (2023)
-
The relative transmission fitness of multidrug-resistant Mycobacterium tuberculosis in a drug resistance hotspot
Nature Communications (2023)
-
Transcontinental spread and evolution of Mycobacterium tuberculosis W148 European/Russian clade toward extensively drug resistant tuberculosis
Nature Communications (2022)