Abstract
Following acute injury, stromal cells promote tissue regeneration by a diversity of mechanisms. Time-resolved single-cell RNA sequencing of muscle mesenchymal stromal cells (MmSCs) responding to acute injury identified an ‘early-responder’ subtype that spiked on day 1 and expressed a notable array of transcripts encoding immunomodulators. IL-1β, TNF-α and oncostatin M each strongly and rapidly induced MmSCs transcribing this immunomodulatory program. Macrophages amplified the program but were not strictly required for its induction. Transfer of the inflammatory MmSC subtype, tagged with a unique surface marker, into healthy hindlimb muscle induced inflammation primarily driven by neutrophils and macrophages. Among the abundant inflammatory transcripts produced by this subtype, Cxcl5 was stroma-specific and highly upregulated with injury. Depletion of this chemokine early after injury revealed a substantial impact on recruitment of neutrophils, a prolongation of inflammation to later times and an effect on tissue regeneration. Mesenchymal stromal cell subtypes expressing a comparable inflammatory program were found in a mouse model of muscular dystrophy and in several other tissues and pathologies in both mice and humans. These ‘early-responder’ mesenchymal stromal cells, already in place, permit rapid and coordinated mobilization and amplification of critical cell collaborators in response to injury.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
New data generated in this paper were deposited in Gene Expression Omnibus database under accession no. GSE205738.
Code availability
No custom code was generated for this study. Algorithms used for data processing and analysis are referenced in the Methods.
References
Robertson, T. A., Grounds, M. D., Mitchell, C. A. & Papadimitriou, J. M. Fusion between myogenic cells in vivo: an ultrastructural study in regenerating murine skeletal muscle. J. Struct. Biol. 105, 170–182 (1990).
Kuang, S., Kuroda, K., Le, G. F. & Rudnicki, M. A. Asymmetric self-renewal and commitment of satellite stem cells in muscle. Cell 129, 999–1010 (2007).
Sacco, A. et al. Self-renewal and expansion of single transplanted muscle stem cells. Nature 456, 502–506 (2008).
von Maltzahn, J., Jones, A. E., Parks, R. J. & Rudnicki, M. A. Pax7 is critical for the normal function of satellite cells in adult skeletal muscle. Proc. Natl Acad. Sci. USA 110, 16474–16479 (2013).
Sambasivan, R. et al. Pax7-expressing satellite cells are indispensable for adult skeletal muscle regeneration. Development 138, 3647–3656 (2011).
Lepper, C., Partridge, T. A. & Fan, C. M. An absolute requirement for Pax7-positive satellite cells in acute injury-induced skeletal muscle regeneration. Development 138, 3639–3646 (2011).
Murphy, M. M. et al. Satellite cells, connective tissue fibroblasts and their interactions are crucial for muscle regeneration. Development 138, 3625–3637 (2011).
Belcastro, A. N., Arthur, G. D., Albisser, T. A. & Raj, D. A. Heart, liver, and skeletal muscle myeloperoxidase activity during exercise. J. Appl. Physiol. 80, 1331–1335 (1996).
Teixeira, C. F. et al. Neutrophils do not contribute to local tissue damage, but play a key role in skeletal muscle regeneration, in mice injected with Bothrops asper snake venom. Muscle Nerve 28, 449–459 (2003).
Brigitte, M. et al. Muscle resident macrophages control the immune cell reaction in a mouse model of notexin-induced myoinjury. Arthritis Rheum. 62, 268–279 (2010).
St Pierre, B. A. & Tidball, J. G. Differential response of macrophage subpopulations to soleus muscle reloading after rat hindlimb suspension. J. Appl. Physiol. 77, 290–297 (1994).
Saclier, M. et al. Differentially activated macrophages orchestrate myogenic precursor cell fate during human skeletal muscle regeneration. Stem Cells 31, 384–396 (2013).
Tonkin, J. et al. Monocyte/macrophage-derived IGF-1 orchestrates murine skeletal muscle regeneration and modulates autocrine polarization. Mol. Ther. 23, 1189–1200 (2015).
Deng, B. et al. IL-10 triggers changes in macrophage phenotype that promote muscle growth and regeneration. J. Immunol. 189, 3669–3680 (2012).
De Micheli, A. J. et al. Single-cell analysis of the muscle stem cell hierarchy identifies heterotypic communication signals involved in skeletal muscle regeneration. Cell Rep. 30, 3583–3595 (2020).
Oprescu, S. N. et al. Temporal dynamics and heterogeneity of cell populations during skeletal muscle regeneration. iScience 23, 100993 (2020).
Arnold, L. et al. Inflammatory monocytes recruited after skeletal muscle injury switch into antiinflammatory macrophages to support myogenesis. J. Exp. Med. 204, 1057–1069 (2007).
Burzyn, D. et al. A special population of regulatory T cells potentiates muscle repair. Cell 155, 1282–1295 (2013).
Castiglioni, A. et al. FOXP3+ T cells recruited to sites of sterile skeletal muscle injury regulate the fate of satellite cells and guide effective tissue regeneration. PLoS ONE 10, e0128094 (2015).
Panduro, M., Benoist, C. & Mathis, D. Treg cells limit IFN-γ production to control macrophage accrual and phenotype during skeletal muscle regeneration. Proc. Natl Acad. Sci. USA 115, E2585–E2593 (2018).
Wosczyna, M. N. et al. Mesenchymal stromal cells are required for regeneration and homeostatic maintenance of skeletal muscle. Cell Rep. 27, 2029–2035 (2019).
Uezumi, A. et al. Mesenchymal Bmp3b expression maintains skeletal muscle integrity and decreases in age-related sarcopenia. J. Clin. Invest. 131, e139617 (2021).
Joe, A. W. et al. Muscle injury activates resident fibro/adipogenic progenitors that facilitate myogenesis. Nat. Cell Biol. 12, 153–163 (2010).
Uezumi, A. et al. Mesenchymal progenitors distinct from satellite cells contribute to ectopic fat cell formation in skeletal muscle. Nat. Cell Biol. 12, 143–152 (2010).
Uezumi, A. et al. Fibrosis and adipogenesis originate from a common mesenchymal progenitor in skeletal muscle. J. Cell Sci. 124, 3654–3664 (2011).
Oishi, T. et al. Osteogenic differentiation capacity of human skeletal muscle-derived progenitor cells. PLoS ONE 8, e56641 (2013).
Wosczyna, M. N., Biswas, A. A., Cogswell, C. A. & Goldhamer, D. J. Multipotent progenitors resident in the skeletal muscle interstitium exhibit robust BMP-dependent osteogenic activity and mediate heterotopic ossification. J. Bone Miner. Res. 27, 1004–1017 (2012).
Lemos, D. R. et al. Nilotinib reduces muscle fibrosis in chronic muscle injury by promoting TNF-mediated apoptosis of fibro/adipogenic progenitors. Nat. Med. 21, 786–794 (2015).
Kopinke, D., Roberson, E. C. & Reiter, J. F. Ciliary hedgehog signaling restricts injury-induced adipogenesis. Cell 170, 340–351 (2017).
Mozzetta, C. et al. Fibroadipogenic progenitors mediate the ability of HDAC inhibitors to promote regeneration in dystrophic muscles of young, but not old Mdx mice. EMBO Mol. Med. 5, 626–639 (2013).
Hogarth, M. W. et al. Fibroadipogenic progenitors are responsible for muscle loss in limb girdle muscular dystrophy 2B. Nat. Commun. 10, 2430 (2019).
Contreras, O. et al. Connective tissue cells expressing fibro/adipogenic progenitor markers increase under chronic damage: relevance in fibroblast-myofibroblast differentiation and skeletal muscle fibrosis. Cell Tissue Res. 364, 647–660 (2016).
Ieronimakis, N. et al. PDGFRα signalling promotes fibrogenic responses in collagen-producing cells in Duchenne muscular dystrophy. J. Pathol. 240, 410–424 (2016).
Mazala, D. A. et al. TGF-β-driven muscle degeneration and failed regeneration underlie disease onset in a DMD mouse model. JCI Insight 5, e135703 (2020).
Lukjanenko, L. et al. Aging disrupts muscle stem cell function by impairing matricellular WISP1 secretion from fibro-adipogenic progenitors. Cell Stem Cell 24, 433–446 (2019).
Iezzi, S. et al. Deacetylase inhibitors increase muscle cell size by promoting myoblast recruitment and fusion through induction of follistatin. Dev. Cell 6, 673–684 (2004).
Reggio, A. et al. Adipogenesis of skeletal muscle fibro/adipogenic progenitors is affected by the WNT5a/GSK3/β-catenin axis. Cell Death Differ. 27, 2921–2941 (2020).
Fiore, D. et al. Pharmacological blockage of fibro/adipogenic progenitor expansion and suppression of regenerative fibrogenesis is associated with impaired skeletal muscle regeneration. Stem Cell Res. 17, 161–169 (2016).
Calve, S., Odelberg, S. J. & Simon, H. G. A transitional extracellular matrix instructs cell behavior during muscle regeneration. Dev. Biol. 344, 259–271 (2010).
Pisconti, A. et al. Syndecan-3 and notch cooperate in regulating adult myogenesis. J. Cell Biol. 190, 427–441 (2010).
Silver, J. S. et al. Injury-mediated stiffening persistently activates muscle stem cells through YAP and TAZ mechanotransduction. Sci. Adv. 7, eabe4501 (2021).
Gilbert, P. M. et al. Substrate elasticity regulates skeletal muscle stem cell self-renewal in culture. Science 329, 1078–1081 (2010).
Buechler, M. B. et al. Cross-tissue organization of the fibroblast lineage. Nature 593, 575–579 (2021).
Scott, R. W. et al. Hic1 defines quiescent mesenchymal progenitor subpopulations with distinct functions and fates in skeletal muscle regeneration. Cell Stem Cell 25, 797–813 (2019).
Tidball, J. G. Regulation of muscle growth and regeneration by the immune system. Nat. Rev. Immunol. 17, 165–178 (2017).
Buechler, M. B., Fu, W. & Turley, S. J. Fibroblast-macrophage reciprocal interactions in health, fibrosis, and cancer. Immunity 54, 903–915 (2021).
Schreiber, H. A. et al. Intestinal monocytes and macrophages are required for T cell polarization in response to Citrobacter rodentium. J. Exp. Med. 210, 2025–2039 (2013).
Heink, S. et al. Trans-presentation of IL-6 by dendritic cells is required for the priming of pathogenic TH17 cells. Nat. Immunol. 18, 74–85 (2017).
Mei, J. et al. CXCL5 regulates chemokine scavenging and pulmonary host defense to bacterial infection. Immunity 33, 106–117 (2010).
Mann, A. O. et al. IL-17A-producing γδT cells promote muscle regeneration in a microbiota-dependent manner. J. Exp. Med. 219, e20211504 (2022).
Seo, B. R. et al. Skeletal muscle regeneration with robotic actuation-mediated clearance of neutrophils. Sci. Transl. Med. 13, eabe8868 (2021).
Horckmans, M. et al. Neutrophils orchestrate post-myocardial infarction healing by polarizing macrophages towards a reparative phenotype. Eur. Heart J. 38, 187–197 (2017).
Aguilar, C. A. et al. In vivo monitoring of transcriptional dynamics after lower-limb muscle injury enables quantitative classification of healing. Sci. Rep. 5, 13885 (2015).
Forte, E. et al. Dynamic interstitial cell response during myocardial infarction predicts resilience to rupture in genetically diverse mice. Cell Rep. 30, 3149–3163 (2020).
Dominguez, C. X. et al. Single-cell RNA sequencing reveals stromal evolution into LRRC15+ myofibroblasts as a determinant of patient response to cancer immunotherapy. Cancer Discov. 10, 232–253 (2020).
Croft, A. P. et al. Distinct fibroblast subsets drive inflammation and damage in arthritis. Nature 570, 246–251 (2019).
Kinchen, J. et al. Structural remodeling of the human colonic mesenchyme in inflammatory bowel disease. Cell 175, 372–386 (2018).
Zhang, F. et al. Defining inflammatory cell states in rheumatoid arthritis joint synovial tissues by integrating single-cell transcriptomics and mass cytometry. Nat. Immunol. 20, 928–942 (2019).
Warren, G. L. et al. Chemokine receptor CCR2 involvement in skeletal muscle regeneration. FASEB J. 19, 413–415 (2005).
Contreras-Shannon, V. et al. Fat accumulation with altered inflammation and regeneration in skeletal muscle of CCR2-/- mice following ischemic injury. Am. J. Physiol. Cell Physiol. 292, C953–C967 (2007).
Shireman, P. K. et al. MCP-1 deficiency causes altered inflammation with impaired skeletal muscle regeneration. J. Leukoc. Biol. 81, 775–785 (2007).
Martinez, C. O. et al. Regulation of skeletal muscle regeneration by CCR2-activating chemokines is directly related to macrophage recruitment. Am. J. Physiol. Regul. Integr. Comp. Physiol. 299, R832–R842 (2010).
Blanc, R. S. et al. Inhibition of inflammatory CCR2 signaling promotes aged muscle regeneration and strength recovery after injury. Nat. Commun. 11, 4167 (2020).
Bettelli, E. et al. Reciprocal developmental pathways for the generation of pathogenic effector TH17 and regulatory T cells. Nature 441, 235–238 (2006).
Tomura, M. et al. Monitoring cellular movement in vivo with photoconvertible fluorescence protein ‘Kaede’ transgenic mice. Proc. Natl Acad. Sci. USA 105, 10871–10876 (2008).
Wang, K. et al. Neuronal, stromal, and T-regulatory cell crosstalk in murine skeletal muscle. Proc. Natl Acad. Sci. USA 117, 5402–5408 (2020).
Liu, L., Cheung, T. H., Charville, G. W. & Rando, T. A. Isolation of skeletal muscle stem cells by fluorescence-activated cell sorting. Nat. Protoc. 10, 1612–1624 (2015).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Stoeckius, M. et al. Simultaneous epitope and transcriptome measurement in single cells. Nat. Methods 14, 865–868 (2017).
Fang, R. et al. Comprehensive analysis of single cell ATAC-seq data with snapATAC. Nat. Commun. 12, 1337 (2021).
Michelson, D. A. et al. Thymic epithelial cells co-opt lineage-defining transcription factors to eliminate autoreactive T cells. Cell 185, 2542–2558 (2022).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096–1098 (2013).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinform. 14, 128 (2013).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Xie, Z. et al. Gene set knowledge discovery with Enrichr. Curr. Protoc. 1, e90 (2021).
Acknowledgements
We thank T. Korn and S. Heink for the IL-6 reporter/depleter line; A. Muñoz-Rojas, G. Wang, T. Xiao, S. Galván-Peña, K. Hattori, A. Ortiz-Lopez, V. Piekarsa, N. Asinovski, F. Chen and S. Nepal for experimental assistance; K. Seddu, A. Baysoy, J. Lee, I. Magill and staff at the Broad Genomics Platform for RNA-seq; A. Muñoz-Rojas, L. Yang, B. Vijaykumar and N. Patel for computational help; C. Laplace for graphics; the Harvard Medical School (HMS) Immunology Flow Core; the HMS MiCRoN Core; the HMS Rodent Histopathology Core; and R.G. Spallanzani, A. Muñoz-Rojas, A. Mann, D. Owen, J. Leon and G. Wang for insightful discussions. This work was funded by National Institutes of Health (NIH) grant R01 AR070334 (D.M.), the JPB Foundation (D.M.) and NIH training grant T32GM007753 (O.K.Y., D.A.M. and T.J.). B.S.H. was partially supported by a Deutsche Forschungsgemeinschaft fellowship (HA 8510/1) and P.K.L. by F32 AG072874.
Author information
Authors and Affiliations
Contributions
O.K.Y., B.S.H. and D.M. conceptualized the study. O.K.Y., B.S.H., P.K.L., D.A.M. and M.M.R. designed and performed experiments. O.K.Y., B.S.H., P.K.L, D.A.M., T.J. and M.M.R. analyzed and interpreted data. O.K.Y., B.S.H. and D.M. wrote the paper, which all authors reviewed. D.M. and C.B. provided supervision. D.M. obtained funding.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: S. Houston, in collaboration with the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Identification of an inflammatory MmSC subtype after acute skeletal-muscle injury.
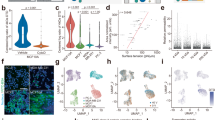
a, Gating strategy for the cytofluorimetric sorting of MmSCs from hindlimb muscles. b, Heat map of the top 20 differentially expressed genes in MmSCs from D1 following CTX-induced injury comparing each cluster vis-à-vis all other clusters. c, Violin plot of the expression of the early signature (top 100 transcripts distinguishing early time points from the rest) across all MmSCs clusters on D1 following CTX-induced injury. d, Density plot of the expression of the indicated genes and gene signatures in MmSCs from D1 following CTX-induced injury. e, Violin plots of the expression of the signatures differentiating the four muscle stromal subtypes from Scott et al.44 across all MmSCs clusters on D1 following CTX-induced injury.
Extended Data Fig. 2 Role of IL-1β, TNFα, OSM in inducing MmSC inflammatory program.
a, Population-level RNA-seq of MmSCs isolated at 2 (n = 2), 4 (n = 1) or 8 (n = 2) hrs after ip co-injection of IL-1β, TNFα, OSM and IL-17A vs after PBS injection (n = 2). Expression levels across time points of the top 20 transcripts from the 100-gene inflammatory signature most differentially expressed. Y-axes plot values in arbitrary units. Each data point represents an individual mouse. b, Violin plots of Il1b, Tnf and Osm expression across cell populations in skeletal muscle on D0.5 after CTX-induced injury. scRNA-seq dataset as per16. Cell nomenclature as per original dataset. c, Gating strategy for the cytofluorimetric analysis of diverse immunocyte populations from hindlimb muscles. AU, arbitrary units; APC, antigen-presenting cells; FAPs, fibro/adipogenic progenitors; M1, inflammatory MFs; MuSC, muscle stem cells; PBS, phosphate-buffered saline; FC, fold change.
Extended Data Fig. 3 Impact of the loss of IL-1β, TNFα and OSM on MmSC inflammatory phenotypes.
a, Population-level RNA-seq of MmSCs isolated from Il1b knock-out (KO) (n = 3) and wild-type (WT) (n = 2) littermates at D1 following CTX-induced injury. Volcano plot comparison of the different conditions. The 100-gene inflammatory signature is shown in red, with numbers at the top indicating up- and downregulated transcripts (in comparison with total transcript numbers in black). b, Same as (a) except B6 mice were treated with a combination of IL-1β, TNFα and OSM neutralizing antibodies or IgG isotype controls. P determined by Chi-squared test.
Extended Data Fig. 4 Il6 and Cxcl5 expression in MmSCs at D1 and across skeletal muscle cell populations at homeostasis and various time points after injury.
a, Violin plot of Il6 expression across all MmSCs clusters on D1 following CTX-induced injury. b, Violin plot of Il6 expression across cell populations in skeletal muscle at homeostasis, as per16. c, Violin plot of Cxcl5 expression across all MmSCs clusters on D1 following CTX-induced injury. d, Violin plot of Cxcl5 expression across cell populations in skeletal muscle at homeostasis, as per16. M2, reparative MFs; Proliferating IC, proliferating immune cells; other abbreviations as per Extended Data Fig. 2. Cell nomenclature as per original dataset.
Extended Data Fig. 5 Expression of 100-gene inflammatory signature in immunocytes and MmSCs at homeostasis and upon acute injury.
Fold change vs fold change plot of population-level RNA-seq of CD45+ cells and MmSCs at D0 and D1 following CTX-induced injury (n = 3 per group). The 100-gene inflammatory signature is shown in red. Abbreviations as per Extended Data Fig. 1.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yaghi, O.K., Hanna, B.S., Langston, P.K. et al. A discrete ‘early-responder’ stromal-cell subtype orchestrates immunocyte recruitment to injured tissue. Nat Immunol 24, 2053–2067 (2023). https://doi.org/10.1038/s41590-023-01669-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-023-01669-w