Abstract
The extent to which unconventional forms of antigen presentation drive T cell immunity is unknown. By convention, CD8 T cells recognize viral peptides, or epitopes, in association with classical major histocompatibility complex (MHC) class I, or MHC-Ia, but immune surveillance can, in some cases, be directed against peptides presented by nonclassical MHC-Ib, in particular the MHC-E proteins (Qa-1 in mice and HLA-E in humans); however, the overall importance of nonclassical responses in antiviral immunity remains unclear. Similarly uncertain is the importance of ‘cryptic’ viral epitopes, defined as those undetectable by conventional mapping techniques. Here we used an immunopeptidomic approach to search for unconventional epitopes that drive T cell responses in mice infected with influenza virus A/Puerto Rico/8/1934. We identified a nine amino acid epitope, termed M-SL9, that drives a co-immunodominant, cytolytic CD8 T cell response that is unconventional in two major ways: first, it is presented by Qa-1, and second, it has a cryptic origin, mapping to an unannotated alternative reading frame product of the influenza matrix gene segment. Presentation and immunogenicity of M-SL9 are dependent on the second AUG codon of the positive sense matrix RNA segment, suggesting translation initiation by leaky ribosomal scanning. During influenza virus A/Puerto Rico/8/1934 infection, M-SL9-specific T cells exhibit a low level of egress from the lungs and strong differentiation into tissue-resident memory cells. Importantly, we show that M-SL9/Qa-1-specific T cells can be strongly induced by messenger RNA vaccination and that they can mediate antigen-specific cytolysis in vivo. Our results demonstrate that noncanonical translation products can account for an important fraction of the T cell repertoire and add to a growing body of evidence that MHC-E-restricted T cells could have substantial therapeutic value.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Raw mass spectrometry data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the identifiers PXD045025 and https://doi.org/10.6019/PXD045025. Data supporting TCR sequence analysis are deposited in GitHub at the URL https://github.com/mhogan240/NatImmuno2023. Data supporting M-SL9 sequence variation analysis are publicly available from BV-BRC. Other data and reagents that support the findings of this study are available from the corresponding authors Michael J. Hogan and Laurence C. Eisenlohr upon request. Source data are provided with this paper.
Code availability
Code supporting analysis of TCR sequences and M-SL9 variant sequences is deposited in GitHub at the URL https://github.com/mhogan240/NatImmuno2023.
References
Hansen, S. G. et al. Immune clearance of highly pathogenic SIV infection. Nature 502, 100–104 (2013).
Hansen, S. G. et al. A live-attenuated RhCMV/SIV vaccine shows long-term efficacy against heterologous SIV challenge. Sci. Transl. Med. 11, eaaw2607 (2019).
Malouli, D. et al. Cytomegaloviral determinants of CD8+ T cell programming and RhCMV/SIV vaccine efficacy. Sci. Immunol. 6, eabg5413 (2021).
Hansen, S. G. et al. Myeloid cell tropism enables MHC-E–restricted CD8+ T cell priming and vaccine efficacy by the RhCMV/SIV vaccine. Sci. Immunol. 7, eabn9301 (2022).
Anderson, C. K., Reilly, E. C., Lee, A. Y. & Brossay, L. Qa-1-restricted CD8+ T cells can compensate for the absence of conventional T cells during viral infection. Cell Rep. 27, 537–548 (2019).
Chen, L., Jay, D. C., Fairbanks, J. D., He, X. & Jensen, P. E. An MHC class Ib-restricted CD8+ T cell response to lymphocytic choriomeningitis virus. J. Immunol. 187, 6463–6472 (2011).
Hansen, S. G. et al. Prevention of tuberculosis in rhesus macaques by a cytomegalovirus-based vaccine. Nat. Med. 24, 130–143 (2018).
Rodgers, J. R. & Cook, R. G. MHC class Ib molecules bridge innate and acquired immunity. Nat. Rev. Immunol. 5, 459–471 (2005).
Braud, V. M. et al. HLA-E binds to natural killer cell receptors CD94/NKG2A, B and C. Nature 391, 795–799 (1998).
Vance, R. E., Kraft, J. R., Altman, J. D., Jensen, P. E. & Raulet, D. Mouse CD94/NKG2A is a natural killer cell receptor for the nonclassical MHC class I molecule Qa-1b. J. Exp. Med. 188, 1841–1848 (1998).
Anderson, C. K. & Brossay, L. The role of MHC class Ib-restricted T cells during infection. Immunogenetics 68, 677–691 (2016).
Chen, X.-R. et al. A signal peptide derived from Hsp60 induces protective cytotoxic T lymphocyte immunity against lymphoid malignancies independently of TAP and classical MHC-I. Cancer Lett. 494, 47–57 (2020).
Malouli, D. et al. Cytomegalovirus-vaccine-induced unconventional T cell priming and control of SIV replication is conserved between primate species. Cell Host Microbe 30, 1207–1218.e7 (2022).
Grifoni, A. et al. SARS-CoV-2 human T cell epitopes: adaptive immune response against COVID-19. Cell Host Microbe 29, 1076–1092 (2021).
Mahajan, S. et al. Immunodominant T-cell epitopes from the SARS-CoV-2 spike antigen reveal robust pre-existing T-cell immunity in unexposed individuals. Sci. Rep. 11, 13164 (2021).
Saini, S. K. et al. SARS-CoV-2 genome-wide T cell epitope mapping reveals immunodominance and substantial CD8+ T cell activation in COVID-19 patients. Sci. Immunol. 6, eabf7550 (2021).
Bullock, T. N. & Eisenlohr, L. C. Ribosomal scanning past the primary initiation codon as a mechanism for expression of CTL epitopes encoded in alternative reading frames. J. Exp. Med. 184, 1319–1329 (1996).
Bullock, T. N. J., Patterson, A. E., Franlin, L. L., Notidis, E. & Eisenlohr, L. C. Initiation codon scanthrough versus termination codon readthrough demonstrates strong potential for major histocompatibility complex class I–restricted cryptic epitope expression. J. Exp. Med. 186, 1051–1058 (1997).
Schwab, S. R., Li, K. C., Kang, C. & Shastri, N. Constitutive display of cryptic translation products by MHC class I molecules. Science 301, 1367–1371 (2003).
Zook, M. B., Howard, M. T., Sinnathamby, G., Atkins, J. F. & Eisenlohr, L. C. Epitopes derived by incidental translational frameshifting give rise to a protective CTL response. J. Immunol. 176, 6928–6934 (2006).
Starck, S. R. et al. Leucine-tRNA initiates at CUG start codons for protein synthesis and presentation by MHC class I. Science 336, 1719–1723 (2012).
Apcher, S. et al. Translation of pre-spliced RNAs in the nuclear compartment generates peptides for the MHC class I pathway. Proc. Natl Acad. Sci. USA 110, 17951–17956 (2013).
Goodenough, E. et al. Cryptic MHC class I-binding peptides are revealed by aminoglycoside-induced stop codon read-through into the 3′ UTR. Proc. Natl Acad. Sci. USA 111, 5670–5675 (2014).
Yang, N. et al. Defining viral defective ribosomal products: standard and alternative translation initiation events generate a common peptide from influenza A virus M2 and M1 mRNAs. J. Immunol. 196, 3608–3617 (2016).
Sanz, M. A., Almela, E. G., García-Moreno, M., Marina, A. I. & Carrasco, L. A viral RNA motif involved in signaling the initiation of translation on non-AUG codons. RNA 25, 431–452 (2019).
Zanker, D. J. et al. Influenza A virus infection induces viral and cellular defective ribosomal products encoded by alternative reading frames. J. Immunol. 202, 3370–3380 (2019).
Hanada, K., Yewdell, J. W. & Yang, J. C. Immune recognition of a human renal cancer antigen through post-translational protein splicing. Nature 427, 252–256 (2004).
Delong, T. et al. Pathogenic CD4 T cells in type 1 diabetes recognize epitopes formed by peptide fusion. Science 351, 711–714 (2016).
Paes, W. et al. Contribution of proteasome-catalyzed peptide cis-splicing to viral targeting by CD8+ T cells in HIV-1 infection. Proc. Natl Acad. Sci. USA 116, 24748–24759 (2019).
Tran, M. T. et al. T cell receptor recognition of hybrid insulin peptides bound to HLA-DQ8. Nat. Commun. 12, 5110 (2021).
Purcell, A. W. Is the immunopeptidome getting darker?: a commentary on the discussion around Mishto et al., 2019. Front. Immunol. 12, 720811 (2021).
Miller, M. A., Ganesan, A. P. V., Luckashenak, N., Mendonca, M. & Eisenlohr, L. C. Endogenous antigen processing drives the primary CD4+ T cell response to influenza. Nat. Med. 21, 1216–1222 (2015).
Tewari, M. K., Sinnathamby, G., Rajagopal, D. & Eisenlohr, L. C. A cytosolic pathway for MHC class II-restricted antigen processing that is proteasome and TAP dependent. Nat. Immunol. 6, 287–294 (2005).
Reynisson, B. et al. Improved prediction of MHC II antigen presentation through integration and motif deconvolution of mass spectrometry MHC eluted ligand data. J. Proteome Res. 19, 2304–2315 (2020).
Parker, R. et al. Mapping the SARS-CoV-2 spike glycoprotein-derived peptidome presented by HLA class II on dendritic cells. Cell Rep. 35, 109179 (2021).
Partridge, T. et al. Discrimination between human leukocyte antigen class I-bound and co-purified HIV-derived peptides in immunopeptidomics workflows. Front. Immunol. 9, 912 (2018).
Martínez-Sobrido, L. & García-Sastre, A. Generation of recombinant influenza virus from plasmid DNA. J. Vis. Exp. https://doi.org/10.3791/2057 (2010).
Ljunggren, H.-G. et al. Empty MHC class I molecules come out in the cold. Nature 346, 476–480 (1990).
Kraft, J. R. et al. Analysis of Qa-1b peptide binding specificity and the capacity of CD94/NKG2A to discriminate between Qa-1–peptide complexes. J. Exp. Med. 192, 613–624 (2000).
Ying, G., Wang, J., Kumar, V. & Zajonc, D. M. Crystal structure of Qa-1a with bound Qa-1 determinant modifier peptide. PLoS ONE 12, e0182296 (2017).
Davies, A. et al. Infection-induced expansion of a MHC class Ib-dependent intestinal intraepithelial γδ T cell subset. J. Immunol. 172, 6828–6837 (2004).
Tang, X. et al. Regulation of immunity by a novel population of Qa-1-restricted CD8αα+ TCRαβ+ T cells. J. Immunol. 177, 7645–7655 (2006).
Niederlova, V., Tsyklauri, O., Chadimova, T. & Stepanek, O. CD8+ Tregs revisited: a heterogeneous population with different phenotypes and properties. Eur. J. Immunol. 51, 512–530 (2021).
Miller, J. D. et al. CD94/NKG2 expression does not inhibit cytotoxic function of lymphocytic choriomeningitis virus-specific CD8+ T cells. J. Immunol. 169, 693–701 (2002).
Borst, L. et al. NKG2A is a late immune checkpoint on CD8 T cells and marks repeated stimulation and cell division. Int. J. Cancer 150, 688–704 (2022).
Van Montfoort, N. et al. NKG2A blockade potentiates CD8 T cell immunity induced by cancer vaccines. Cell 175, 1744–1755.e15 (2018).
Borst, L., Van Der Burg, S. H. & Van Hall, T. The NKG2A–HLA-E axis as a novel checkpoint in the tumor microenvironment. Clin. Cancer Res. 26, 5549–5556 (2020).
Lepore, M. et al. Parallel T-cell cloning and deep sequencing of human MAIT cells reveal stable oligoclonal TCRβ repertoire. Nat. Commun. 5, 3866 (2014).
Yuan, R. et al. The roles of tissue-resident memory T cells in lung diseases. Front. Immunol. 12, 710375 (2021).
Schön, M. P. et al. Mucosal T lymphocyte numbers are selectively reduced in integrin alpha E (CD103)-deficient mice. J. Immunol. 162, 6641–6649 (1999).
Takamura, S. et al. Specific niches for lung-resident memory CD8+ T cells at the site of tissue regeneration enable CD69-independent maintenance. J. Exp. Med. 213, 3057–3073 (2016).
Walsh, D. A. et al. The functional requirement for CD69 in establishment of resident memory CD8+ T cells varies with tissue location. J. Immunol. 203, 946–955 (2019).
Laidlaw, B. J. et al. CD4+ T cell help guides formation of CD103+ lung-resident memory CD8+ T cells during influenza viral infection. Immunity 41, 633–645 (2014).
Zens, K. D., Chen, J. K. & Farber, D. L. Vaccine-generated lung tissue-resident memory T cells provide heterosubtypic protection to influenza infection. JCI Insight 1, e85832 (2016).
Belz, G. T., Xie, W., Altman, J. D. & Doherty, P. C. A previously unrecognized H-2Db-restricted peptide prominent in the primary influenza A virus-specific CD8+ T-cell response is much less apparent following secondary challenge. J. Virol. 74, 3486–3493 (2000).
Machkovech, H. M., Bloom, J. D. & Subramaniam, A. R. Comprehensive profiling of translation initiation in influenza virus infected cells. PLoS Pathog. 15, e1007518 (2019).
Wise, H. M. et al. Identification of a novel splice variant form of the influenza A virus M2 ion channel with an antigenically distinct ectodomain. PLoS Pathog. 8, e1002998 (2012).
Pardi, N., Hogan, M. J. & Weissman, D. Recent advances in mRNA vaccine technology. Curr. Opin. Immunol. 65, 14–20 (2020).
Liu, X. et al. MARCH8 inhibits influenza A virus infection by targeting viral M2 protein for ubiquitination-dependent degradation in lysosomes. Nat. Commun. 12, 4427 (2021).
Kozak, M. Adherence to the first-AUG rule when a second AUG codon follows closely upon the first. Proc. Natl Acad. Sci. USA 92, 2662–2666 (1995).
Hogan, M. J. & Pardi, N. mRNA vaccines in the COVID-19 pandemic and beyond. Annu. Rev. Med. 73, 17–39 (2022).
Knudson, C. J., Hartwig, S. M., Meyerholz, D. K. & Varga, S. M. RSV vaccine-enhanced disease is orchestrated by the combined actions of distinct CD4 T cell subsets. PLoS Pathog. 11, 1–23 (2015).
Wei, J. & Yewdell, J. W. Flu DRiPs in MHC class I immunosurveillance. Virol. Sin. 34, 162–167 (2019).
Lodha, M., Erhard, F., Dölken, L. & Prusty, B. K. The hidden enemy within: non-canonical peptides in virus-induced autoimmunity. Front. Microbiol. 13, 840911 (2022).
Kracht, M. J. L. et al. Autoimmunity against a defective ribosomal insulin gene product in type 1 diabetes. Nat. Med. 23, 501–507 (2017).
Marcu, A. et al. Natural and cryptic peptides dominate the immunopeptidome of atypical teratoid rhabdoid tumors. J. Immunother. Cancer 9, e003404 (2021).
Chong, C. et al. Integrated proteogenomic deep sequencing and analytics accurately identify non-canonical peptides in tumor immunopeptidomes. Nat. Commun. 11, 1293 (2020).
Ruiz Cuevas, M. V. et al. Most non-canonical proteins uniquely populate the proteome or immunopeptidome. Cell Rep. 34, 108815 (2021).
Croft, N. P. et al. Kinetics of antigen expression and epitope presentation during virus infection. PLoS Pathog. 9, e1003129 (2013).
Wu, T. et al. Quantification of epitope abundance reveals the effect of direct and cross-presentation on influenza CTL responses. Nat. Commun. 10, 2846 (2019).
Yewdell, J. W., Dersh, D. & Fåhraeus, R. Peptide channeling: the key to MHC class I immunosurveillance? Trends Cell Biol. 29, 929–939 (2019).
Rutigliano, J. A. et al. Highly pathological influenza A virus infection is associated with augmented expression of PD-1 by functionally compromised virus-specific CD8+ T cells. J. Virol. 88, 1636–1651 (2014).
Vogel, A. J., Harris, S., Marsteller, N., Condon, S. A. & Brown, D. M. Early cytokine dysregulation and viral replication are associated with mortality during lethal influenza infection. Viral Immunol. 27, 214–224 (2014).
Seaman, M. S., Wang, C.-R. & Forman, J. MHC class Ib-restricted CTL provide protection against primary and secondary Listeria monocytogenes infection. J. Immunol. 165, 5192–5201 (2000).
Laidlaw, B. J. et al. Cooperativity between CD8+ T cells, non-neutralizing antibodies, and alveolar macrophages is important for heterosubtypic influenza virus immunity. PLoS Pathog. 9, e1003207 (2013).
LaMere, M. W. et al. Contributions of antinucleoprotein IgG to heterosubtypic immunity against influenza virus. J. Immunol. 186, 4331–4339 (2011).
Kanevskiy, L. et al. Dimorphism of HLA-E and its disease association. Int. J. Mol. Sci. 20, 5496 (2019).
Sahin, U. et al. Personalized RNA mutanome vaccines mobilize poly-specific therapeutic immunity against cancer. Nature 547, 222–226 (2017).
Rojas, L. A. et al. Personalized RNA neoantigen vaccines stimulate T cells in pancreatic cancer. Nature 618, 144–150 (2023).
Voogd, L., Ruibal, P., Ottenhoff, T. H. M. & Joosten, S. A. Antigen presentation by MHC-E: a putative target for vaccination? Trends Immunol. 43, 355–365 (2022).
Sinnathamby, G., Maric, M., Cresswell, P. & Eisenlohr, L. C. Differential requirements for endosomal reduction in the presentation of two H2-Ed-restricted epitopes from influenza hemagglutinin. J. Immunol. 172, 6607–6614 (2004).
Sanderson, S. & Shastri, N. LacZ inducible, antigen/MHC-specific T cell hybrids. Int. Immunol. 6, 369–376 (1994).
Chen, L. et al. Expression of the mouse MHC class Ib H2-T11 gene product, a paralog of H2-T23 (Qa-1) with shared peptide-binding specificity. J. Immunol. 193, 1427–1439 (2014).
Purcell, A. W., Ramarathinam, S. H. & Ternette, N. Mass spectrometry–based identification of MHC-bound peptides for immunopeptidomics. Nat. Protoc. 14, 1687–1707 (2019).
Pardi, N., Muramatsu, H., Weissman, D. & Karikó, K. In vitro transcription of long RNA containing modified nucleosides. Methods Mol. Biol. 969, 29–42 (2013).
Laczkó, D. et al. A single immunization with nucleoside-modified mRNA vaccines elicits strong cellular and humoral immune responses against SARS-CoV-2 in mice. Immunity 53, 724–732.e7 (2020).
Baiersdörfer, M. et al. A facile method for the removal of dsRNA contaminant from in vitro-transcribed mRNA. Mol. Ther. Nucleic Acids 15, 26–35 (2019).
Freyn, A. W. et al. A multi-targeting, nucleoside-modified mRNA influenza virus vaccine provides broad protection in mice. Mol. Ther. 28, 1569–1584 (2020).
Thompson, M. G. et al. Co-regulatory activity of hnRNP K and NS1-BP in influenza and human mRNA splicing. Nat. Commun. 9, 2407 (2018).
Shu, Y. & McCauley, J. GISAID: global initiative on sharing all influenza data—from vision to reality. Eurosurveillance 22, 30494 (2017).
Wagih, O. ggseqlogo: a versatile R package for drawing sequence logos. Bioinformatics 33, 3645–3647 (2017).
Acknowledgements
We thank F. Tuluc, J. Murray and J. Lora of the Children’s Hospital of Philadelphia Flow Cytometry Core Facility for technical advice and services; L. Spruce, H. Fazelinia and S. Seeholzer (formerly) of the Children’s Hospital of Philadelphia Proteomics Core Facility for technical guidance and services; the NIH Tetramer Core Facility for providing tetramers for this study; J. R. Melamed and D. Weissman for technical advice on LNP generation; R. Serafin for providing related data; J. J. Rim for assistance with manuscript preparation; and D. F. Jenkins for data management support. We gratefully acknowledge the contributors to the Influenza Research Database, BV-BRC and the GISAID database, including the laboratories and authors responsible for obtaining specimens, generating genetic sequences and sharing data via the GISAID Initiative. M.J.H. was supported by the Cancer Research Institute as a Cancer Research Institute Irvington Fellow and by the Roberts Family–Katalin Karikó Fellowship in Vaccine Development from the Aileen K. and Brian L. Roberts Family Foundation via the University of Pennsylvania Institute for Immunology & Immune Health (I3H). N.M. was supported by the Roy and Diana Vagelos Molecular Life Sciences Program and by a College Alumni Society Research Grant from the University of Pennsylvania. N.P. was supported by NIH R01AI146101 and R01AI153064. S.P.R. is supported by research supplement 3R01AI046709-18S1 to promote diversity and L.B. is supported by NIH R01AI046709. K.W.L. and B.E.B. are supported by R01AI125524. L.C.E. and N.T. were supported by NIH R21AI153978. This work was funded in part by contract #75N93021C00015 from NIH NIAID. BioRender.com was used to create panels in Figs. 1, 2 and 8.
Author information
Authors and Affiliations
Contributions
M.J.H. conceived the project, designed the studies, performed experiments, analyzed and interpreted the data and wrote the paper. N.M. co-conceived the project, co-designed studies, co-performed the immunoprecipitation, ELISpot and antigen presentation experiments and analyzed and interpreted data. B.E.B. performed primer extension assays and provided interpretation together with K.W.L. A.N. and N.T. performed LC–MS2, analyzed data and provided interpretation. E.J.H. provided essential support for animal studies. H.M. and N.P. contributed mRNA reagents and related expertise. M.A.M. isolated the B6.23 hybridoma. S.P.R. and L.B. provided reagents and expertise regarding MHC-Ia and Qa-1b−/− bone marrow. L.C.E. co-conceived and advised the project and interpreted the data. All authors provided critical scientific feedback, aided in the preparation of the manuscript and agree with the conclusions.
Corresponding authors
Ethics declarations
Competing interests
N.T. is or has been a paid consultant to Roche Pharma, Enara Bio, Grey Wolf Therapeutics, T-Cypher Bio and Infinitopes on the topic of cancer antigen discovery. All other authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks Katherine Kedzierska and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: S. Houston, in collaboration with the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Peptide analysis by mass spectrometry (MS).
a, b, Histograms of peptide lengths of unique peptide species identified from (a) any origin and (b) IAV origin. c–f, NetMHCIIpan 4.0 was used to predict peptide:MHC-II affinity (KD) values for core epitope sequences of 9 amino acids from MS-identified peptides of all lengths. Sequence logo diagrams were prepared using unique core epitopes predicted to bind to (c,d) I-Ab or (e,f) I-Ed with a KD < 2,000 nM for peptides of (c,e) any origin or KD < 10,000 nM for peptides of (d,f) IAV origin. These sequence logos clearly exhibit the expected MHC-II ligand sequence motifs (based on ref. 34). g, The sequence logo diagram for all unique 9-mer peptide identifications (appearing as a small local peak in panel ‘a’) shows strong similarity to the H2-Kb and H2-Db sequence motifs, but not to the Qa-1 sequence motif (motifs available from NetMHCpan 4.0 at ref. 82). In sequence logo diagrams, ‘bit’ is a unit of relative amino acid frequency that is inversely related to the Shannon entropy of each position.
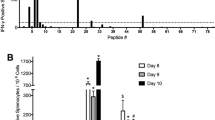
Extended Data Fig. 2 Identification of M-SL9 by mass spectrometry and validation of immunogenicity in C57Bl/6 mice.
a, MS2 spectrum resulting in M-SL9 identification, with b and y ion fragments indicated, along with mass/charge (m/z), retention time (RT), and P-value. b, c, Spleens were recovered from C57Bl/6 (N = 3) or BALB/c (N = 4) mice 9 days after infection with IAV PR8, and either bulk spleen cells or isolated spleen CD4 T cells were stimulated overnight with synthetic M-SL9 peptide, positive control peptides, or DMSO vehicle, and secreted IFN-γ was detected by ELISpot. b, For C57Bl/6 mice, the MHC-Ia control peptide was NS2109-121; the MHC-II control peptides were NP264-280 and NP306-322; and DC2.4 cells were used as the APC to stimulate CD4 T cells. c, For BALB/c mice, the MHC-II control peptides were NP55-71 and HA121-137, and A20 cells were used as the APC to stimulate CD4 T cells. Data points represent individual mice in one independent experiment; bars are the mean +/− s.e.m.
Extended Data Fig. 3 Annotation of M-SL9 coding sequence within the full IAV PR8 matrix gene segment.
The positive-sense RNA sequence is shown on top. Primary and secondary AUG codons are underlined and labeled, M-SL9 amino acids are highlighted in yellow, the M1 protein sequence is in light blue, M-MG16 nucleotides including stop codon are colored red, and relevant splice sites are labeled. The first nucleotide of each codon is aligned with the single-letter amino acid code and the first digit of the nucleotide number. This sequence was used to generate PR8 for this study and matches the sequence in GenBank accession AF389121.
Extended Data Fig. 4 Intracellular cytokines, cytolytic markers, and FoxP3 expression in individual mice.
a–d, Representative gating strategy for intracellular cytokine and cytolytic marker staining (lower boxes from two different experiments/stains). b–e, Lymphocytes were isolated from naïve (N = 21) or PR8 flu-infected (N = 34) C57Bl/6 mouse lungs, stimulated with indicated peptides, and stained for the indicated markers. b–d, Data for individual mice are shown in the same order for each epitope. c, Comparison of intracellular cytokine responses following infection with 40 FFU PR8 (N = 7), 160 FFU PR8 (N = 7), or no virus (naïve; N = 5), showing more consistent M-SL9 responses to 160 FFU. Female and male mice are indicated by purple and orange bars underneath the graphs. e, CD8 T cells (both total unstimulated as well as peptide-restimulated IFN-γ+ cells) from PR8-infected mice (N = 11) stain do not upregulate FoxP3 expression relative to the naïve (N = 4) mouse baseline. Gray events are all CD3+ cells; blue events and blue percentages represent CD3+ CD8+ cells. Bars show the mean +/- s.d. and P-values of interest are shown from a two-way ANOVA with Sidak’s multiple comparisons test comparing naïve and PR8-infected conditions. GzmB: granzyme B.
Extended Data Fig. 5 Hybridoma clone B6.23 recognizes two forms of M-SL9 present in isolates of PR8.
Amino acid sequences are shown for the originally identified M-SL9, present in pDZ PR8, and M-SL9-P, present in other PR8 isolates (for example GenBank V01099). B6-CIITA fibroblasts served as APCs and were co-cultured overnight with B6.23 cells in the presence of the indicated peptide concentrations. A sigmoidal curve was fit to the data points above (mean +/− s.d.), representative of three independent experiments. The geometric mean half-maximal effective concentration (EC50) values across all three experiments were computed as 940 ng/ml for M-SL9 and 51 ng/ml for M-SL9-P.
Extended Data Fig. 6 Evidence supporting Qa-1 restriction of M-SL9.
a, The MHC-Ia molecules H2-Db and H2-Kb are not stabilized on RMA-S cells by M-SL9 peptide. RMA-S cells bearing unstable empty MHC-I molecules (due to TAP deficiency) were incubated in the presence of the indicated synthetic peptides, and surface expression of H2-Db and H2-Kb was measured by flow cytometry. Mean fluorescence intensities of each stain were normalized to the negative control condition using HA91-107, an I-Ab-binding epitope with no known binding to H2-Db or H2-Kb, and shown as averages +/- s.d from 3 independent experiments. H2-Db-binding NP366-374 and H2-Kb-binding SIINFEKL were used as positive controls. b-e, Validation of HeLa cell lines and BMDCs showing Qa-1 restriction of M-SL9. b, The sufficiency of Qa-1b expression for M-SL9 presentation to its cognate T hybridoma was confirmed using a HeLa cell line transduced with full-length, wild-type Qa-1b and an untransduced parental HeLa cell line as a control. Bars are mean +/- s.e.m. from triplicate technical replicates, representative of 3 independent experiments, and P-values were calculated by Welch’s t-test (two-tailed). c, Qa-1 expression on cell lines used in b was validated by flow cytometry. d, The expected staining pattern was confirmed for HeLa cell lines used in Fig. 2; these lines were transduced with retroviruses encoding chimeric MHC-Ib molecules containing the α3 domain (D3) from H2-Db to allow efficient staining with the H2-Db D3-specific mAb 28-14-8. e, The expected staining pattern was also confirmed for BMDCs used in Fig. 2.
Extended Data Fig. 7 Qdm/Qa-1b tetramer co-stains with NKG2A/C/E.
a, Lung lymphocytes from naïve C57Bl/6 mice were stained with an anti-NKG2A/C/E mAb and Qdm/Qa-1b, M-SL9/Qa-1b, and control NP366-374/Db tetramers at 37 °C and gated on CD3− CD19− cells to interrogate natural killer (NK) cells. NK cells expressing NKG2A/C/E (the natural receptors for Qdm/Qa-1b) were the only population that stained with Qdm/Qa-1b tetramer, but neither these nor other NK cells stained with M-SL9/Qa-1b tetramer. b, C57Bl/6 mice were intranasally infected with 160 FFU of PR8 and 9 days later lung lymphocytes were stained with anti-NKG2A/C/E and the indicated tetramers at 37 °C. Qdm/Qa-1b tetramer generally stained PR8-induced CD8 T cells in a manner that was dependent on NKG2A/C/E but independent of TCR specificity. Flow plots are representative and show the gating strategy used, and bars show mean +/- s.e.m. for (a) N = 4 mice and (b) N = 5 to 6 mice per group across 2 independent experiments each. P-values are shown from two-way ANOVA with Dunnett’s multiple comparisons test.
Extended Data Fig. 8 Analysis of TCRβ V and J gene usage in sorted CD8 T cell populations.
a-d, CD8 T cell populations were sorted by FACS into three populations: naïve (CD44− CD62L+), M-SL9-specific (CD44+ M-SL9/Qa-1b tetramer+), and NP366-374-specific (CD44+ NP366-374/H2-Db tetramer+). Genomic DNA was isolated, the VDJ region of recombined TCRβ-coding genes was sequenced, and gene usage was analyzed by (a,b) the immunoSEQ Analyzer and (c,d) Immunarch. a-b, The frequencies of the top 10 most-used (a) V genes and (b) J genes, on average across all mice, are shown as stacked bar graphs, where each bar represents one mouse. c, Principal component analysis (PCA) of the Tcrb V and J gene usage showing clustering by T cell population. d, Pearson correlation analysis of M-SL9- and NP366-374-specific T cells showing greater correlation between mice within each T cell specificity rather than between specificities within each mouse. N = 6 mice, half males and half females.
Extended Data Fig. 9 Tracking CD69 and CD103 expression in PR8-infected mice.
a-d, C57Bl/6 mice were intranasally infected with 160 FFU of PR8 and were euthanized at day 6 (N = 7), 9 (N = 5-6), 14, 31 (N = 7), or 56 (N = 9) to collect the indicated tissues/fluids. Uninfected mice were used as day 0 controls (N = 7-11). a, Gating strategy. b, Frequency of all CD3+ T cells in lung and BALF over time, showing the lack of T cell infiltration in uninfected mice (plotted as day 0). Data points were omitted when there were <20 live singlet CD3+ CD8+ T cells collected in total. P-values are calculated from Brown-Forsythe and Welch one-way ANOVA with Dunnett T3 multiple comparisons test comparing each condition to day 0 controls. c, Frequency of CD103+ and CD69+ CD8 T cells (analyzed separately) in lung and BALF starting from the approximate peak of the T cell response on day 9. d, Frequency of CD103+ CD69+ double positive TRM cells in lung at 9 days after PR8 only, X31 only, or PR8 prime and X31 boost. c, d, P-values were calculated by two-way ANOVA with Tukey’s multiple comparisons test.
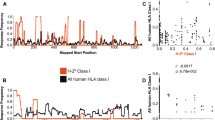
Extended Data Fig. 10 Sequence evolution and variation of the M-SL9 epitope and open reading frame in IAV strains over time since 1980.
a–c, Sequence logo diagrams were produced from M-SL9-homologous sequences from (a) human H1N1 isolates, (b) human H3N2 isolates, and (c) H5N1 isolates from all avian species, downloaded between April and June 2023 for the indicated sample collection time periods. The BV-BRC database was used for sequences from 1980-1999, while the GISAID database was used for all others. Diagrams were created using the ggseqlogo package in R, and the y-axis units are the probability of each amino acid from 0 to 1. H1N1 sequences after 2009 correspond to the swine-origin pandemic H1N1 lineage, while H1N1 sequences prior to 2009 are from the earlier seasonal H1N1 lineage; sequences from 2009 were omitted to avoid ambiguity. Amino acids are numbered so that position 1 corresponds to the initial serine residue of the M-SL9 epitope, and the preceding residue was designated as position −1 and shown to assess the presence of an initiation codon. The two forms of M-SL9 encoded by PR8 isolates are shown at bottom; note that the avian H5N1 consensus sequence exactly matches the M-SL9-P amino acid sequence.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2.
Source data
Source Data Fig. 7
Source data for Fig. 7d (full unprocessed image for IAV matrix RNA primer extension gel).
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hogan, M.J., Maheshwari, N., Begg, B.E. et al. Cryptic MHC-E epitope from influenza elicits a potent cytolytic T cell response. Nat Immunol 24, 1933–1946 (2023). https://doi.org/10.1038/s41590-023-01644-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-023-01644-5
This article is cited by
-
Unconventionally presenting an unconventional viral peptide
Nature Immunology (2023)