Abstract
Natural killer (NK) cells are innate cytotoxic lymphocytes with adaptive immune features, including antigen specificity, clonal expansion and memory. As such, NK cells share many transcriptional and epigenetic programs with their adaptive CD8+ T cell siblings. Various signals ranging from antigen, co-stimulation and proinflammatory cytokines are required for optimal NK cell responses in mice and humans during virus infection; however, the integration of these signals remains unclear. In this study, we identified that the transcription factor IRF4 integrates signals to coordinate the NK cell response during mouse cytomegalovirus infection. Loss of IRF4 was detrimental to the expansion and differentiation of virus-specific NK cells. This defect was partially attributed to the inability of IRF4-deficient NK cells to uptake nutrients required for survival and memory generation. Altogether, these data suggest that IRF4 is a signal integrator that acts as a secondary metabolic checkpoint to orchestrate the adaptive response of NK cells during viral infection.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All published data generated and supporting the findings of this study have been deposited to NCBI’s GEO or European Genome–Phenome Archive as described. scRNA-seq and ChIP-seq data generated in this study have been deposited at GEO under accession code GSE236556. Source data are provided with this paper.
References
Lau, C. M. et al. Epigenetic control of innate and adaptive immune memory. Nat. Immunol. 19, 963–972 (2018).
Mujal, A. M., Delconte, R. B. & Sun, J. C. Natural killer cells: from innate to adaptive features. Annu. Rev. Immunol. 39, 417–447 (2021).
Sun, J. C., Beilke, J. N. & Lanier, L. L. Adaptive immune features of natural killer cells. Nature 457, 557–561 (2009).
Adams, E. J. et al. Structural elucidation of the m157 mouse cytomegalovirus ligand for Ly49 natural killer cell receptors. Proc. Natl Acad. Sci. USA 104, 10128–10133 (2007).
Arase, H., Mocarski, E. S., Campbell, A. E., Hill, A. B. & Lanier, L. L. Direct recognition of cytomegalovirus by activating and inhibitory NK cell receptors. Science 296, 1323–1326 (2002).
Nabekura, T. et al. Costimulatory molecule DNAM-1 is essential for optimal differentiation of memory natural killer cells during mouse cytomegalovirus infection. Immunity 40, 225–234 (2014).
Madera, S. et al. Type I IFN promotes NK cell expansion during viral infection by protecting NK cells against fratricide. J. Exp. Med. 213, 225–233 (2016).
Madera, S. & Sun, J. C. Cutting edge: stage-specific requirement of IL-18 for antiviral NK cell expansion. J. Immunol. 194, 1408–1412 (2015).
Sun, J. C. et al. Proinflammatory cytokine signaling required for the generation of natural killer cell memory. J. Exp. Med. 209, 947–954 (2012).
Wiedemann, G. M. et al. Divergent role for STAT5 in the adaptive responses of natural killer cells. Cell Rep. 33, 108498 (2020).
Wiedemann, G. M. et al. Deconvoluting global cytokine signaling networks in natural killer cells. Nat. Immunol. 22, 627–638 (2021).
Hammer, Q., Ruckert, T. & Romagnani, C. Natural killer cell specificity for viral infections. Nat. Immunol. 19, 800–808 (2018).
Zhao, G. N., Jiang, D. S. & Li, H. Interferon regulatory factors: at the crossroads of immunity, metabolism, and disease. Biochim. Biophys. Acta 1852, 365–378 (2015).
Huber, M. & Lohoff, M. IRF4 at the crossroads of effector T-cell fate decision. Eur. J. Immunol. 44, 1886–1895 (2014).
Mittrucker, H. W. et al. Requirement for the transcription factor LSIRF/IRF4 for mature B and T lymphocyte function. Science 275, 540–543 (1997).
Duncan, G. S., Mittrucker, H. W., Kagi, D., Matsuyama, T. & Mak, T. W. The transcription factor interferon regulatory factor-1 is essential for natural killer cell function in vivo. J. Exp. Med. 184, 2043–2048 (1996).
Lohoff, M. et al. Deficiency in the transcription factor interferon regulatory factor (IRF)-2 leads to severely compromised development of natural killer and T helper type 1 cells. J. Exp. Med. 192, 325–336 (2000).
Ogasawara, K. et al. Requirement for IRF-1 in the microenvironment supporting development of natural killer cells. Nature 391, 700–703 (1998).
Mace, E. M. et al. Biallelic mutations in IRF8 impair human NK cell maturation and function. J. Clin. Invest. 127, 306–320 (2017).
Adams, N. M. et al. Transcription factor IRF8 orchestrates the adaptive natural killer cell response. Immunity 48, 1172–1182 (2018).
Beaulieu, A. M., Zawislak, C. L., Nakayama, T. & Sun, J. C. The transcription factor Zbtb32 controls the proliferative burst of virus-specific natural killer cells responding to infection. Nat. Immunol. 15, 546–553 (2014).
Bravo Garcia-Morato, M. et al. New human combined immunodeficiency caused by interferon regulatory factor 4 (IRF4) deficiency inherited by uniparental isodisomy. J. Allergy Clin. Immunol. 141, 1924–1927 (2018).
Man, K. et al. The transcription factor IRF4 is essential for TCR affinity-mediated metabolic programming and clonal expansion of T cells. Nat. Immunol. 14, 1155–1165 (2013).
Geary, C. D. et al. Non-redundant ISGF3 components promote NK cell survival in an auto-regulatory manner during viral infection. Cell Rep. 24, 1949–19576 (2018).
Schlums, H. et al. Cytomegalovirus infection drives adaptive epigenetic diversification of NK cells with altered signaling and effector function. Immunity 42, 443–456 (2015).
Rückert, T., Lareau, C. A., Mashreghi, M. F., Ludwig, L. S. & Romagnani, C. Clonal expansion and epigenetic inheritance of long-lasting NK cell memory. Nat. Immunol. 23, 1551–1563 (2022).
Holmes, T. D. et al. The transcription factor Bcl11b promotes both canonical and adaptive NK cell differentiation. Sci. Immunol. 6, eabc9801 (2021).
Kallies, A. et al. A role for Blimp1 in the transcriptional network controlling natural killer cell maturation. Blood 117, 1869–1879 (2011).
Yao, S. et al. Interferon regulatory factor 4 sustains CD8(+) T cell expansion and effector differentiation. Immunity 39, 833–845 (2013).
Flommersfeld, S. et al. Fate mapping of single NK cells identifies a type 1 innate lymphoid-like lineage that bridges innate and adaptive recognition of viral infection. Immunity 54, 2288–2304 (2021).
Finak, G. et al. MAST: a flexible statistical framework for assessing transcriptional changes and characterizing heterogeneity in single-cell RNA sequencing data. Genome Biol. 16, 278 (2015).
Intlekofer, A. M. et al. Effector and memory CD8+ T cell fate coupled by T-bet and eomesodermin. Nat. Immunol. 6, 1236–1244 (2005).
Omilusik, K. D. et al. Transcriptional repressor ZEB2 promotes terminal differentiation of CD8+ effector and memory T cell populations during infection. J. Exp. Med. 212, 2027–2039 (2015).
van Helden, M. J. et al. Terminal NK cell maturation is controlled by concerted actions of T-bet and Zeb2 and is essential for melanoma rejection. J. Exp. Med. 212, 2015–2025 (2015).
Zook, E. C. et al. Transcription factor ID2 prevents E proteins from enforcing a naive T lymphocyte gene program during NK cell development. Sci. Immunol. 3, eaao2139 (2018).
Riggan, L. et al. The transcription factor Fli1 restricts the formation of memory precursor NK cells during viral infection. Nat. Immunol. 23, 556–567 (2022).
Milner, J. J. et al. Runx3 programs CD8(+) T cell residency in non-lymphoid tissues and tumours. Nature 552, 253–257 (2017).
Wang, D. et al. The transcription factor Runx3 establishes chromatin accessibility of cis-regulatory landscapes that drive memory cytotoxic T lymphocyte formation. Immunity 48, 659–674 (2018).
Delconte, R. B. et al. The helix-loop-helix protein ID2 governs NK cell fate by tuning their sensitivity to interleukin-15. Immunity 44, 103–115 (2016).
Loftus, R. M. et al. Amino acid-dependent cMyc expression is essential for NK cell metabolic and functional responses in mice. Nat. Commun. 9, 2341 (2018).
Frost, J. N. et al. Evaluation of perturbed iron-homeostasis in a prospective cohort of patients with COVID-19. Wellcome Open Res. 7, 173 (2022).
Frost, J. N. et al. Plasma iron controls neutrophil production and function. Sci. Adv. 8, eabq5384 (2022).
Bonadonna, M. et al. Iron regulatory protein (IRP)-mediated iron homeostasis is critical for neutrophil development and differentiation in the bone marrow. Sci. Adv. 8, eabq4469 (2022).
Li, P. et al. BATF-JUN is critical for IRF4-mediated transcription in T cells. Nature 490, 543–546 (2012).
Kaech, S. M. & Cui, W. Transcriptional control of effector and memory CD8+ T cell differentiation. Nat. Rev. Immunol. 12, 749–761 (2012).
Laidlaw, B. J. & Cyster, J. G. Transcriptional regulation of memory B cell differentiation. Nat. Rev. Immunol. 21, 209–220 (2021).
Adams, N. M. et al. Cytomegalovirus infection drives avidity selection of natural killer cells. Immunity 50, 1381–1390 (2019).
Grassmann, S. et al. Distinct surface expression of activating receptor Ly49H drives differential expansion of NK cell clones upon murine cytomegalovirus infection. Immunity 50, 1391–1400 (2019).
Gordon, S. M. et al. The transcription factors T-bet and Eomes control key checkpoints of natural killer cell maturation. Immunity 36, 55–67 (2012).
Yang, C. Y. et al. The transcriptional regulators Id2 and Id3 control the formation of distinct memory CD8+ T cell subsets. Nat. Immunol. 12, 1221–1229 (2011).
Pearce, E. L. et al. Control of effector CD8+ T cell function by the transcription factor eomesodermin. Science 302, 1041–1043 (2003).
Sheppard, S. et al. Lactate dehydrogenase A-dependent aerobic glycolysis promotes natural killer cell anti-viral and anti-tumor function. Cell Rep. 35, 109210 (2021).
Mah-Som, A. Y. et al. Reliance on Cox10 and oxidative metabolism for antigen-specific NK cell expansion. Cell Rep. 35, 109209 (2021).
Littwitz-Salomon, E. et al. Metabolic requirements of NK cells during the acute response against retroviral infection. Nat. Commun. 12, 5376 (2021).
Ganz, T. & Nemeth, E. Iron homeostasis in host defence and inflammation. Nat. Rev. Immunol. 15, 500–510 (2015).
Lane, D. J. et al. Cellular iron uptake, trafficking and metabolism: key molecules and mechanisms and their roles in disease. Biochim. Biophys. Acta 1853, 1130–1144 (2015).
Tussiwand, R. et al. Compensatory dendritic cell development mediated by BATF–IRF interactions. Nature 490, 502–507 (2012).
Ciofani, M. et al. A validated regulatory network for Th17 cell specification. Cell 151, 289–303 (2012).
Glasmacher, E. et al. A genomic regulatory element that directs assembly and function of immune-specific AP-1-IRF complexes. Science 338, 975–980 (2012).
Kim, S. et al. High amount of transcription factor IRF8 engages AP1-IRF composite elements in enhancers to direct type 1 conventional dendritic cell identity. Immunity 53, 759–774 (2020).
Consortium, I. R. F. I. et al. A multimorphic mutation in IRF4 causes human autosomal dominant combined immunodeficiency. Sci. Immunol. 8, eade7953 (2023).
Thouenon, R. et al. A neomorphic mutation in the interferon activation domain of IRF4 causes a dominant primary immunodeficiency. J. Exp. Med. 220, e20221292 (2023).
Guerin, A. et al. IRF4 haploinsufficiency in a family with Whipple’s disease. eLife 7, e32340 (2018).
Huntington, N. D., Cursons, J. & Rautela, J. The cancer-natural killer cell immunity cycle. Nat. Rev. Cancer 20, 437–454 (2020).
Bald, T., Krummel, M. F., Smyth, M. J. & Barry, K. C. The NK cell-cancer cycle: advances and new challenges in NK cell-based immunotherapies. Nat. Immunol. 21, 835–847 (2020).
Berrien-Elliott, M. M. et al. Hematopoietic cell transplantation donor-derived memory-like NK cells functionally persist after transfer into patients with leukemia. Sci. Transl. Med. 14, eabm1375 (2022).
Marin, N. D. et al. Memory-like differentiation enhances NK cell responses to melanoma. Clin. Cancer Res. 27, 4859–4869 (2021).
Gauthier, L. et al. Multifunctional natural killer cell engagers targeting NKp46 trigger protective tumor immunity. Cell 177, 1701–1713 (2019).
Andre, P. et al. Anti-NKG2A mAb is a checkpoint inhibitor that promotes anti-tumor immunity by unleashing both T and NK cells. Cell 175, 1731–1743 (2018).
Zhang, Q. et al. Blockade of the checkpoint receptor TIGIT prevents NK cell exhaustion and elicits potent anti-tumor immunity. Nat. Immunol. 19, 723–732 (2018).
Liu, E. et al. Use of CAR-transduced natural killer cells in CD19-positive lymphoid tumors. N. Engl. J. Med. 382, 545–553 (2020).
Knorr, D. A., Bachanova, V., Verneris, M. R. & Miller, J. S. Clinical utility of natural killer cells in cancer therapy and transplantation. Semin. Immunol. 26, 161–172 (2014).
Seo, H. et al. BATF and IRF4 cooperate to counter exhaustion in tumor-infiltrating CAR T cells. Nat. Immunol. 22, 983–995 (2021).
Mercier, F. E., Sykes, D. B. & Scadden, D. T. Single targeted exon mutation creates a true congenic mouse for competitive hematopoietic stem cell transplantation: the C57BL/6-CD45.1(STEM) mouse. Stem Cell Rep. 6, 985–992 (2016).
Fodil-Cornu, N. et al. Ly49h-deficient C57BL/6 mice: a new mouse cytomegalovirus-susceptible model remains resistant to unrelated pathogens controlled by the NK gene complex. J. Immunol. 181, 6394–6405 (2008).
Stoeckius, M. et al. Cell hashing with barcoded antibodies enables multiplexing and doublet detection for single cell genomics. Genome Biol. 19, 224 (2018).
Azizi, E. et al. Single-cell map of diverse immune phenotypes in the breast tumor microenvironment. Cell 174, 1293–1308 (2018).
Shekhar, K. et al. Comprehensive classification of retinal bipolar neurons by single-cell transcriptomics. Cell 166, 1308–1323 (2016).
Wolf, F. A. et al. PAGA: graph abstraction reconciles clustering with trajectory inference through a topology preserving map of single cells. Genome Biol. 20, 59 (2019).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
van Dijk, D. et al. Recovering gene interactions from single-cell data using data diffusion. Cell 174, 716–729 (2018).
Joshi, N. S. et al. Inflammation directs memory precursor and short-lived effector CD8(+) T cell fates via the graded expression of T-bet transcription factor. Immunity 27, 281–295 (2007).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Zhu, L. J. et al. ChIPpeakAnno: a Bioconductor package to annotate ChIP-seq and ChIP-chip data. BMC Bioinform. 11, 237 (2010).
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. 9, R137 (2008).
Ramirez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Hammer, Q. et al. Peptide-specific recognition of human cytomegalovirus strains controls adaptive natural killer cells. Nat. Immunol. 19, 453–463 (2018).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Lareau, C. A. et al. Massively parallel single-cell mitochondrial DNA genotyping and chromatin profiling. Nat. Biotechnol. 39, 451–461 (2021).
Stuart, T., Srivastava, A., Madad, S., Lareau, C. A. & Satija, R. Single-cell chromatin state analysis with Signac. Nat. Methods 18, 1333–1341 (2021).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Acknowledgements
We thank members of the Sun laboratory, K. Lupo and X. Chen, as well as the Single-Cell Analytics Innovation Lab at the Sloan Kettering Institute for technical support and experimental assistance, especially R. Chaligné. We also thank L. Lanier for helpful discussions. K. Murphy and T. Murphy provided mice critical to this study. T. Holmes and Y. Bryceson shared sequencing data from their study. We acknowledge the use of the Integrated Genomics Operation Core, funded by the National Cancer Institute Cancer Center Support Grant (P30CA008748), Cycle for Survival and the Marie-Josée and Henry R. Kravis Center for Molecular Oncology. J.N.F. is a Cancer Research Institute Irvington Fellow supported by the Cancer Research Institute (award CRI4186). K.C.H. and J.-B.L.L were supported by funding from the National Institutes of Health (R01 AI150999, R01HL155741, U01AI069197). J.C.S. was supported by the Ludwig Center for Cancer Immunotherapy, the American Cancer Society, the Burroughs Wellcome Fund and the National Institutes of Health (AI100874, AI130043, AI155558 and P30CA008748).
Author information
Authors and Affiliations
Contributions
E.K.S. and J.C.S. designed the study. E.K.S., H.K., J.-B.L.L., A.J.A., C.K.W., L.P. and J.N.F. performed experiments. E.K.S. and T.R. performed bioinformatics analyses. K.C.H., C.R. and J.C.S. provided critical resources and supervision for the study. E.K.S. and J.C.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks Dagmar Gotthardt and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Evidence of IRF4-mediated transcriptional control in NK cells.
a. Heatmap of IRF family members expression in NK cells throughout the course of MCMV infection represented as z-score of log2 normalized counts based on RNA-seq. b. Representative histogram of IRF4 expression in WT and Irf4−/− NK cells on day 2 PI (n = 4 biological replicates). c. Tracks (top) show chromatin accessibility dynamics of the Irf4 locus in Ly49H+ NK cells at days 0, 2, 4, and 7 PI as assessed by ATAC-seq. Graphs (bottom) show normalized counts for each peak indicated. d. Tables of enriched known motifs of highly accessible regions (log2FC > 1 & padj < 0.05) on day 4 PI versus day 2 PI from ATAC-seq data of Ly49H+ NK cells after MCMV infection. e. Heatmap of motif score from de novo motif analysis on highly accessible regions (log2FC > 1 & padj < 0.05) during day 0, 2, 4, and 7 transition based on ATAC-seq data of Ly49H+ NK cells after MCMV infection. f. IRF4 expression of sorted naïve splenic NK cells after an overnight stimulation with the indicated stimuli. IRF4 induction is displayed as fold change of IRF4 MFI over unstimulated condition (n = 6 biological replicates per condition). Two-way ANOVA test adjusted for multiple comparisons was used for statistical analysis. g. Representative histogram of IRF4 expression upon indicated stimulation gated on human CD56bright NK cells. Data are presented as paired fold change of IRF4 MFI compared to unstimulated condition (n = 8 donors per condition). Two-way ANOVA test adjusted for multiple comparisons was used for statistical analysis. h. UMAP embedding of scATAC-seq data from in vitro stimulated human NK cells (sorted on CD3−CD14−CD19−CD7+NKG2C+) from HCMV− donors. i. Coverage plot of the IRF4 locus from in vitro stimulated human NK cells as in (H).
Extended Data Fig. 2 IRF4 is dispensable for NK cell development and homeostasis.
a–c. Number of NK cells in blood (A), liver (B), and spleen (C) of WT (n = 8), Irf4+/− (n = 4), or Irf4−/− (n = 6) mice at 8 weeks old. d–f. Maturation status based on the expression of CD27 and CD11b of WT (n = 8), Irf4+/− (n = 4), or Irf4−/− (n = 5 for blood, and n = 6 for liver and spleen) mice in the blood (D), liver (E), and spleen (F) at 8 weeks old. g. Percentage of Ly49H+ NK cells within NK cells (Lineage- NK1.1+ CD49b+) of WT (n = 8), Irf4+/− (n = 4), or Irf4−/− (n = 6) mice at 8 weeks old. h. Experimental schematic of WT:Irf4−/− mixed bone-marrow chimera (mBMC) generation. i–k. Percentage of NK cells of WT or Irf4−/− mice from WT:Irf4−/− mBMC in the blood (n = 60 biological replicates) (I) and spleen (n = 21 biological replicates) (J) after 8 weeks (I-J), 16 weeks (n = 3 biological replicates) or 6 months (n = 5 biological replicates) post-transplant (K). l. Representative of flow plots of CD62L versus CD27 of either WT or Irf4−/− NK cells from WT:Irf4−/− mBMC in the blood and spleen 8 weeks post-transplant. Data are represented as percentage of each subset within total NK cells of each genotype (n = 22 biological replicates in blood, and n = 13 biological replicates in spleen). m. Representative of flow plots of CD11b versus CD27 of either WT or Irf4−/− NK cells from WT:Irf4−/− mBMC in the blood and spleen 8 weeks post-transplant. Data are represented as percentage of each subset within total NK cells of each genotype (n = 22 biological replicates in blood, and n = 16 biological replicates in spleen). n. Histogram and percentage quantification of Ly49H+ NK cells within splenic NK cells of either WT or Irf4−/− NK cells from WT:Irf4−/− mBMC (n = 24 biological replicates). Data are represented as mean ± SEM and are representative of or pooled from at least two independent experiments. Unpaired (A-G) and paired (I-N) two-tailed t-tests were performed.
Extended Data Fig. 3 IRF4 is not required for lymphopenia-driven proliferation and differentiation.
a. Experimental schematic of adoptive transfer of WT and Irf4−/− NK cells into Rag2−/− Il2rg−/− mice. b. Fold change chimerism between WT and Irf4−/− NK cells on day 7 over day 0 (pre-transfer) in blood, liver, and spleen (n = 3 biological replicates in blood, and n = 6 biological replicates in liver and spleen). c. Representative histogram of IRF4 expression between transferred WT and Irf4−/− NK cells on day 7 post-transfer into Rag2−/− Il2rg−/−. d. Representative flow plots of CD27 and CD11b expression between WT and Irf4−/− NK cells on day 7 post-transfer. Data are represented as percentage of each subset within WT or Irf4−/− NK cells in indicated tissues (n = 3 biological replicates). e. Histogram of CD122 and quantification of CD122 MFI between WT and Irf4−/− NK cells on day 7 post-transfer in the spleen (n = 9 biological replicates). f. Histogram of CD132 and quantification of CD132 MFI between WT and Irf4−/− NK cells on day 7 post-transfer in the spleen (n = 9 biological replicates). g. Histogram of pSTAT5 and quantification of pSTAT5 MFI between WT and Irf4−/− NK cells on day 7 post-transfer in the spleen (n = 9 biological replicates). H-J. h–j. Quantification of Ki-67 staining in the blood and spleen between WT and Irf4−/− NK cells (H, n = 5 biological replicates per group), BIM and BCL2 MFI from splenic NK cells (I-J, n = 9 biological replicates). Data are represented as mean ± SEM and are representative of or pooled from at least two independent experiments. Paired two-tailed t-test was performed unless stated otherwise.
Extended Data Fig. 4 IRF4-deficient NK cells have intact effector functions.
a, b Representative histograms and quantification of IFN-γ production (A) and LAMP-1 (also known as CD107a) (B) from either naive WT (gray) or Irf4−/− (orange) NK cells upon in vitro stimulation with nothing, IL-12 + IL-18, PMA + Ionomycin, or PMA + Ionomycin + IL-12 for 3 hours. c–f. Representative histograms (upper panels) and quantifications (bottom panels) of CD25, Gzmb, CD69, and IFN-γ expression gated on Ly49H+ NK cells from MCMV-infected WT:Irf4−/− mBMC on day 2 PI. Data are represented as mean ± SEM and are representative of or pooled from at least two independent experiments. n = 3 biological replicates per group. Paired two-tailed t-test was performed unless stated otherwise.
Extended Data Fig. 5 scRNA-seq Analysis of wild-type versus IRF4-deficient NK cells.
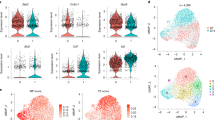
a. UMAP embedding of WT (left) or Irf4−/− (right) of Ly49H+ NK cells colored by genotypes and days post-infection. b. Proportion of each genotype from each time points per Louvain clusters. c. Volcano plot of scRNA-seq as determined by MAST between WT and Irf4−/− Ly49H+ NK cells on day 0, 2, 4 and 7 PI. Blue and yellow points represent significant genes (FDR < 0.05) that are upregulated in WT or Irf4−/− Ly49H+ NK cells, respectively. d. Number of differentially expressed genes between WT or Irf4−/− Ly49H+ NK cells from each time point as identified by MAST. e. Violin plot quantification of normalized reads for MYC transcript by scRNA-seq between WT and Irf4−/− Ly49H+ NK cells. f. Representative histogram of Myc protein by flow cytometry between WT (gray) and Irf4−/− (orange) Ly49H+ NK cells on day 5 PI. g. Violin plot quantification of MAGIC-imputed reads for CD71 (Tfrc), IRP1 (Aco1), and IRP2 (Ireb2) on day 7 PI between WT (blue) and Irf4−/− (yellow) Ly49H+ NK cells by scRNA-seq.
Extended Data Fig. 6 Nutrient requirements for adaptive NK cell response.
a. Experimental schematic of PBS or BCH treatment upon MCMV infection in WT mice. b. Quantification of Ly49H+ NK cell numbers of MCMV-infected mice treated with PBS or BCH on day 5 PI from blood (n = 3 biological replicates per group). c. Percent of CD62L versus CD27 subsets on day 5 PI of PBS- (black) or BCH-treated (red) mice (n = 3 biological replicates per group). d. Representative histogram and gMFI of CD71 expression on Ly49H+ NK cells on day 5 PI in PBS or Deferiprone-treated mice. Upregulation of CD71 was used as a readout for successful iron chelation (n = 5 biological replicates per group). a–g. Percent of KLRG1+ Ly49H+ NK cells (E), and gMFIs of IL18Ra (F) and CD44 (G) on Ly49H+ NK cells on day 5 PI from PBS or Deferiprone-treated mice (n = 5 biological replicates per group). Data are represented as mean ± SEM and are representative of at least two independent experiments. Two-way ANOVA adjusted for multiple comparisons was performed unless stated otherwise.
Extended Data Fig. 7 Gating strategies to identify NK cells.
a. Gating strategy for sorting WT or Irf4−/− Ly49H+ NK cells. b. Gating strategy for phenotypic analysis of Ly49H+ NK cells after lymphocytes, singlets and live cells cleanup (as depicted in Extended Data Fig. 7a). c. Gating strategy for human NK cells.
Supplementary information
Supplementary Table 1
A list of antibodies used for flow cytometric analysis and scRNA-seq HTO-tag.
Source data
Source Data main figures
Statistical source data.
Source Data extended data
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Santosa, E.K., Kim, H., Rückert, T. et al. Control of nutrient uptake by IRF4 orchestrates innate immune memory. Nat Immunol 24, 1685–1697 (2023). https://doi.org/10.1038/s41590-023-01620-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-023-01620-z
This article is cited by
-
Cardinal features of immune memory in innate lymphocytes
Nature Immunology (2023)
-
Ironman training for NK cells
Nature Immunology (2023)