Abstract
Small-molecule kinase inhibitors represent a major group of cancer therapeutics, but tumor responses are often incomplete. To identify pathways that modulate kinase inhibitor response, we conducted a genome-wide knockout (KO) screen in glioblastoma cells treated with the pan-ErbB inhibitor neratinib. Loss of general control nonderepressible 2 (GCN2) kinase rendered cells resistant to neratinib, whereas depletion of the GADD34 phosphatase increased neratinib sensitivity. Loss of GCN2 conferred neratinib resistance by preventing binding and activation of GCN2 by neratinib. Several other Food and Drug Administration (FDA)-approved inhibitors, such erlotinib and sunitinib, also bound and activated GCN2. Our results highlight the utility of genome-wide functional screens to uncover novel mechanisms of drug action and document the role of the integrated stress response (ISR) in modulating the response to inhibitors of oncogenic kinases.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The CRISPR screen and RNA-seq data reported in this study have been deposited to the Gene Expression Omnibus with the accession number GSE188958. The LINCS KINOMEscan data is from project ID 20195 (https://lincs.hms.harvard.edu/db/datasets/20195/main). PDB IDs for the crystal structure data used in this study are 3W2Q (ref. 28), 2GS6 (ref. 49) and GCN2 complexed with dovitinib27. Source data are provided with this paper.
Code availability
All scripts for downstream analysis of the CRISPR screen and RNA-seq data are available on GitHub at https://github.com/cot2005/Tang_et_al_2021.
References
Ferguson, F. M. & Gray, N. S. Kinase inhibitors: the road ahead. Nat. Rev. Drug Discov. 17, 353–377 (2018).
Dar, A. C. & Shokat, K. M. The evolution of protein kinase inhibitors from antagonists to agonists of cellular signaling. Annu. Rev. Biochem. 80, 769–795 (2011).
Lake, E. W. et al. Quantitative conformational profiling of kinase inhibitors reveals origins of selectivity for Aurora kinase activation states. Proc. Natl Acad. Sci. USA 115, E11894–E11903 (2018).
Wen, P. Y. et al. Glioblastoma in adults: a Society for Neuro-Oncology (SNO) and European Society of Neuro-Oncology (EANO) consensus review on current management and future directions. Neuro Oncol. 22, 1073–1113 (2020).
Brennan, C. W. et al. The somatic genomic landscape of glioblastoma. Cell 155, 462–477 (2013).
Lee, J. C. et al. Epidermal growth factor receptor activation in glioblastoma through novel missense mutations in the extracellular domain. PLoS Med. 3, e485 (2006).
Reardon, D. A., Wen, P. Y. & Mellinghoff, I. K. Targeted molecular therapies against epidermal growth factor receptor: past experiences and challenges. Neuro Oncol. 16, viii7–viii13 (2014).
An, Z., Aksoy, O., Zheng, T., Fan, Q. W. & Weiss, W. A. Epidermal growth factor receptor and EGFRvIII in glioblastoma: signaling pathways and targeted therapies. Oncogene 37, 1561–1575 (2018).
Rabindran, S. K. et al. Antitumor activity of HKI-272, an orally active, irreversible inhibitor of the HER-2 tyrosine kinase. Cancer Res. 64, 3958–3965 (2004).
Vivanco, I. et al. Differential sensitivity of glioma- versus lung cancer-specific EGFR mutations to EGFR kinase inhibitors. Cancer Discov. 2, 458–471 (2012).
Ji, H. et al. Epidermal growth factor receptor variant III mutations in lung tumorigenesis and sensitivity to tyrosine kinase inhibitors. Proc. Natl Acad. Sci. USA 103, 7817–7822 (2006).
Barkovich, K. J. et al. Kinetics of inhibitor cycling underlie therapeutic disparities between EGFR-driven lung and brain cancers. Cancer Discov. 2, 450–457 (2012).
Hart, T. et al. Evaluation and design of genome-wide CRISPR/SpCas9 knockout screens. G3 (Bethesda) 7, 2719–2727 (2017).
Joung, J. et al. Genome-scale CRISPR–Cas9 knockout and transcriptional activation screening. Nat. Protoc. 12, 828–863 (2017).
Padyana, A. K., Qiu, H., Roll-Mecak, A., Hinnebusch, A. G. & Burley, S. K. Structural basis for autoinhibition and mutational activation of eukaryotic initiation factor 2α protein kinase GCN2. J. Biol. Chem. 280, 29289–29299 (2005).
Dong, J., Qiu, H., Garcia-Barrio, M., Anderson, J. & Hinnebusch, A. G. Uncharged tRNA activates GCN2 by displacing the protein kinase moiety from a bipartite tRNA-binding domain. Mol. Cell 6, 269–279 (2000).
Lageix, S., Zhang, J., Rothenburg, S. & Hinnebusch, A. G. Interaction between the tRNA-binding and C-terminal domains of yeast Gcn2 regulates kinase activity in vivo. PLoS Genet. 11, e1004991 (2015).
Castilho, B. A. et al. Keeping the eIF2 α kinase Gcn2 in check. Biochim. Biophys. Acta 1843, 1948–1968 (2014).
Ye, J. et al. The GCN2–ATF4 pathway is critical for tumour cell survival and proliferation in response to nutrient deprivation. EMBO J. 29, 2082–2096 (2010).
Sattlegger, E. & Hinnebusch, A. G. Separate domains in GCN1 for binding protein kinase GCN2 and ribosomes are required for GCN2 activation in amino acid-starved cells. EMBO J. 19, 6622–6633 (2000).
Marton, M. J., Vazquez de Aldana, C. R., Qiu, H., Chakraburtty, K. & Hinnebusch, A. G. Evidence that GCN1 and GCN20, translational regulators of GCN4, function on elongating ribosomes in activation of eIF2α kinase GCN2. Mol. Cell. Biol. 17, 4474–4489 (1997).
Brush, M. H., Weiser, D. C. & Shenolikar, S. Growth arrest and DNA damage-inducible protein GADD34 targets protein phosphatase 1 α to the endoplasmic reticulum and promotes dephosphorylation of the α subunit of eukaryotic translation initiation factor 2. Mol. Cell. Biol. 23, 1292–1303 (2003).
Elbein, A. D. Inhibitors of the biosynthesis and processing of N-linked oligosaccharide chains. Annu. Rev. Biochem. 56, 497–534 (1987).
Davis, M. I. et al. Comprehensive analysis of kinase inhibitor selectivity. Nat. Biotechnol. 29, 1046–1051 (2011).
Keyvanjah, K. et al. Pharmacokinetics of neratinib during coadministration with lansoprazole in healthy subjects. Br. J. Clin. Pharmacol. 83, 554–561 (2017).
Reichardt, P. et al. Correlation of KIT and PDGFRA mutational status with clinical benefit in patients with gastrointestinal stromal tumor treated with sunitinib in a worldwide treatment-use trial. BMC Cancer 16, 22 (2016).
Maia de Oliveira, T. et al. The structure of human GCN2 reveals a parallel, back-to-back kinase dimer with a plastic DFG activation loop motif. Biochem. J. 477, 275–284 (2020).
Sogabe, S. et al. Structure-based approach for the discovery of pyrrolo[3,2-d]pyrimidine-based EGFR T790M/L858R mutant inhibitors. ACS Med. Chem. Lett. 4, 201–205 (2013).
Arteaga, C. L. & Engelman, J. A. ERBB receptors: from oncogene discovery to basic science to mechanism-based cancer therapeutics. Cancer Cell 25, 282–303 (2014).
Lightfoot, H. L., Goldberg, F. W. & Sedelmeier, J. Evolution of small molecule kinase drugs. ACS Med. Chem. Lett. 10, 153–160 (2019).
Finlay, M. R. et al. Discovery of a potent and selective EGFR inhibitor (AZD9291) of both sensitizing and T790M resistance mutations that spares the wild type form of the receptor. J. Med. Chem. 57, 8249–8267 (2014).
Frye, S. V. & Johnson, G. L. Inhibitors paradoxically prime kinases. Nat. Chem. Biol. 5, 448–449 (2009).
Poulikakos, P. I., Zhang, C., Bollag, G., Shokat, K. M. & Rosen, N. RAF inhibitors transactivate RAF dimers and ERK signalling in cells with wild-type BRAF. Nature 464, 427–430 (2010).
Okuzumi, T. et al. Inhibitor hijacking of Akt activation. Nat. Chem. Biol. 5, 484–493 (2009).
Hu, J. et al. Allosteric activation of functionally asymmetric RAF kinase dimers. Cell 154, 1036–1046 (2013).
Lavoie, H. et al. Inhibitors that stabilize a closed RAF kinase domain conformation induce dimerization. Nat. Chem. Biol. 9, 428–436 (2013).
Goncalves, E. et al. Drug mechanism-of-action discovery through the integration of pharmacological and CRISPR screens. Mol. Syst. Biol. 16, e9405 (2020).
Klaeger, S. et al. The target landscape of clinical kinase drugs. Science 358, eaan4368 (2017).
Wortel, I. M. N., van der Meer, L. T., Kilberg, M. S. & van Leeuwen, F. N. Surviving stress: modulation of ATF4-mediated stress responses in normal and malignant cells. Trends Endocrinol. Metab. 28, 794–806 (2017).
Hidalgo, M. et al. Phase I and pharmacologic study of OSI-774, an epidermal growth factor receptor tyrosine kinase inhibitor, in patients with advanced solid malignancies. J. Clin. Oncol. 19, 3267–3279 (2001).
Demetri, G. D. et al. Efficacy and safety of sunitinib in patients with advanced gastrointestinal stromal tumour after failure of imatinib: a randomised controlled trial. Lancet 368, 1329–1338 (2006).
Angevin, E. et al. Phase I study of dovitinib (TKI258), an oral FGFR, VEGFR, and PDGFR inhibitor, in advanced or metastatic renal cell carcinoma. Clin. Cancer Res. 19, 1257–1268 (2013).
Colic, M. et al. Identifying chemogenetic interactions from CRISPR screens with drugZ. Genome Med. 11, 52 (2019).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Mootha, V. K. et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Malyutina, A. et al. Drug combination sensitivity scoring facilitates the discovery of synergistic and efficacious drug combinations in cancer. PLoS Comput. Biol. 15, e1006752 (2019).
Zhang, X., Gureasko, J., Shen, K., Cole, P. A. & Kuriyan, J. An allosteric mechanism for activation of the kinase domain of epidermal growth factor receptor. Cell 125, 1137–1149 (2006).
Acknowledgements
This research was supported by the National Institutes of Health (F31 5F31CA239401-03 (C.P.T.), 1 R35 NS105109 03 (I.K.M.), P30CA008748 (I.K.M., A.M.I., J.D.C.), UL1TR002384 (O.E.), R01CA194547 (O.E.) and R01GM121505 (J.D.C.)), the National Brain Tumor Society Defeat GBM Initiative (I.K.M.), Cycle of Survival (I.K.M.) and LLS SCOR grants 180078-02, 7021-20 (O.E.). We thank J. Blenis, N. Rosen and C. Sawyers for helpful suggestions.
Author information
Authors and Affiliations
Contributions
C.P.T. and I.K.M. conceptualized and led the overall project, analyzed results and wrote the manuscript with input from all coauthors. O.E. assisted with experimental design and bioinformatic analysis. J.R.F. assisted with CRISPR KO screens. A.M.I. directed and assisted with metabolic assessments and analysis. J.D.C. directed and assisted with structural analysis. A.S.L. assisted in the design of neratinib in vivo experiments. C.C. performed all of the in vivo experiments. O.C. assisted with the orthotopic in vivo experiment. C.P.T. performed all other experiments, bioinformatic analysis and data visualization.
Corresponding author
Ethics declarations
Competing interests
I.K.M. has received research funding from General Electric, Agios and Lilly and has served in advisory roles for Agios, Amgen, Debiopharm, Novartis, Puma Biotechnology, Servier Pharmaceuticals, and Voyager Therapeutics. O.E. is supported by research grants from Janssen, Johnson and Johnson, Volastra, AstraZeneca and Eli Lilly. O.E. is scientific advisor and equity holder in Freenome, Owkin, Volastra Therapeutics and One Three Biotech. A.S.L. is an employee and shareholder of Puma Biotechnology, Inc. J.D.C. has received research funding from the Parker Institute for Cancer Immunotherapy, Relay Therapeutics, Entasis Therapeutics, Silicon Therapeutics, EMD Serono (Merck KGaA), AstraZeneca, Vir Biotechnology, Bayer, XtalPi, Foresite Laboratories, the Molecular Sciences Software Institute, the Starr Cancer Consortium, the Open Force Field Consortium and Cycle for Survival. J.D.C. is a scientific advisor and equity holder in Interline Therapeutics and Redesign Science and a current member of the Scientific Advisory Board of OpenEye Scientific Software. A complete funding history for the Chodera lab can be found at http://choderalab.org/funding. The other authors declare no competing interests.
Additional information
Peer review information Nature Chemical Biology thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 CRISPR screen resistance candidates are significant and consistent.
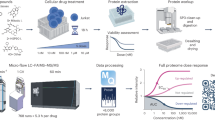
a, Schematic of genome-wide CRISPR screens using the TKOv3 library. b, Representative graph of a CRISPR screen showing the second highest sgRNA (per gene) frequencies of SKMG3 cell line treated with 2 µM neratinib versus vehicle for 14 days. c, Effect of GCN2-KO on neratinib treatment measured by SKMG3 cell growth and assessed by trypan blue exclusion after 6 days. Immunoblot confirming GCN2-KO on right side. Data are presented as mean values ± SD of n = 3 biologically independent samples. d, Spearman correlation matrix of all CRISPR screen experiments showing spearman coefficients. e, false discovery rates of ISR pathway genes from drugZ analyzed CRISPR screens. f, Immunoblot of ATF4 induction by 4 hour 675 nM neratinib in parental and ATF4-KO SF268 cells. c,e,f, Western blots were performed with a minimum of n = 2 biological replicates with similar results obtained each time.
Extended Data Fig. 2 Induction of the ISR by neratinib is GCN2 dependent.
a, Immunoblot of SKMG3 parental and GCN2-KO and SF268 parental and GCN2-KO cells treated with 675 nM neratinib for 4 hours. b, ISR induction time course in TS895 cells EGFP or GCN2-KO cells treated with 675 nM neratinib for indicated times. Western blots were performed with a minimum of n = 2 biological replicates with similar results obtained each time.
Extended Data Fig. 3 Constitutive activation of eIF2α induces cell death.
a, Immunoblot of SF268 cells were transduced with doxycycline (dox) inducible wild-type (WT) or constitutively active (S51D) eIF2α and treated with 5 µg/mL dox for indicated times (upper band, tagged recombinant eIF2α). Western blots were performed with a minimum of n = 2 biological replicates with similar results obtained each time. b, effect of dox induced WT or S51D eIF2α on cell growth. Data are presented as mean values ± SD of n = 3 biologically independent samples.
Extended Data Fig. 4 Induction of the amino acid starvation response by neratinib requires GCN2.
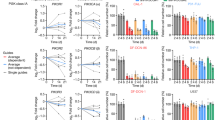
a, Volcano plots and gene set enrichment analysis results showing differential expression of genes in SF268 cells treated with 675 nM neratinib or vehicle for 6 hours and b, 72 hours with gene labels corresponding to the enriched krige amino acid gene set. c, gene set enrichment analysis results showing differential expression of genes in SF268 cells treated with 675 nM neratinib or vehicle for 6 hours and d, 72 hours. e, gene set enrichment plot for the amino acid deprivation gene set for intracranial TS895 tumors treated with neratinib 40 mpk for 3 hours. f, Plot comparing the enrichment of ER stress and amino acid deprivation pathways between SF268 parental and GCN2-KO cell lines treated with 675 nM neratinib for 6 hours. g, Comparison of ISR signaling in the presence of neratinib (1 µM)[lanes 1 and 2 also shown in Fig. 2e], glutamine (Gln) starvation, and ER stress by tunicamycin (5 µg/mL) for 4 hours. Western blots were performed with a minimum of n = 2 biological replicates with similar results obtained each time.
Extended Data Fig. 5 GCN2 is sufficient for neratinib induced eIF2α phosphorylation.
a, Schematic of recombinant GCN2 electroporation rescue. b, immunoblot of eIF2α phosphorylation for SKMG3 cells electroporated with recombinant GCN2 then treated with 675 nM neratinib for 4 hours. Western blots were performed with a minimum of n = 2 biological replicates with similar results obtained each time. c, image quantification of eIF2α phosphorylation, normalized to vinculin.
Extended Data Fig. 6 GCN2 loss confers resistance to GCN2 activating KIT inhibitor.
a, Immunoblot of ISR signaling in GIST-T1 (Left) and SF268 parental/GCN2-KO (Right) treated with the KIT inhibitor sunitinib for 4 hours. Western blots were performed with a minimum of n = 2 biological replicates with similar results obtained each time. b, Bar graph comparing the effect of sunitinib on SF268 parental and GCN2-KO cell growth after 6 days. Data are presented as mean values ± SD of n = 3 biologically independent samples. Significance was determined with a two sided student t-test.
Extended Data Fig. 7 GCN2 activation by dovitinib synergizes with osimertinib.
a, Immunoblot of SF268 cells treated with either neratinib (Ner), osimertinib (Osi), or canertinib (Can) for 4 hours. Western blots were performed with a minimum of n = 2 biological replicates with similar results obtained each time. b, Scatterplot of a CRISPR screen showing the second highest sgRNA (per gene) frequencies of SF268 cell line treated with 3 µM osimertinib versus vehicle for 21 days. ISR pathway members and PTEN are labeled. c, Synergy between GCN2 activation by dovitinib (0 - 1.5 µM) and EGFR inhibition by Osimertinib (0 - 2 µM) in SF268 parental (left) and GCN2-KO (right) cells. Synergy assessed with Bliss synergy scores.
Supplementary information
Supplementary Information
Supplementary Table 1.
Source data
Source Data Fig. 1
Uncropped western blots.
Source Data Fig. 2
Uncropped western blots.
Source Data Fig. 3
Uncropped western blots.
Source Data Fig. 4
Uncropped western blots.
Source Data Fig. 5
Uncropped western blots.
Source Data Fig. 6
Uncropped western blots.
Source Data Extended Data Fig. 1
Uncropped western blots.
Source Data Extended Data Fig. 2
Uncropped western blots.
Source Data Extended Data Fig. 3
Uncropped western blots.
Source Data Extended Data Fig. 4
Uncropped western blots.
Source Data Extended Data Fig. 5
Uncropped western blots.
Source Data Extended Data Fig. 6
Uncropped western blots.
Source Data Extended Data Fig. 7
Uncropped western blots.
Source Data Figures
Statistical source data for Figs. 1–6.
Source Data Extended Data Figures
Statistical source data for Extended Data Figs. 1–7.
Rights and permissions
About this article
Cite this article
Tang, C.P., Clark, O., Ferrarone, J.R. et al. GCN2 kinase activation by ATP-competitive kinase inhibitors. Nat Chem Biol 18, 207–215 (2022). https://doi.org/10.1038/s41589-021-00947-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-021-00947-8
This article is cited by
-
Targeting NEDD8-activating enzyme for cancer therapy: developments, clinical trials, challenges and future research directions
Journal of Hematology & Oncology (2023)
-
Activation of the integrated stress response by inhibitors of its kinases
Nature Communications (2023)
-
Protein tyrosine kinase inhibitor resistance in malignant tumors: molecular mechanisms and future perspective
Signal Transduction and Targeted Therapy (2022)